This is an Application Brief and does not contain a detailed Experimental section.
LC-MS analytical method development requires flexible and reproducible sample preparation for accurate and robust analyte quantification. This work highlights automated calibration curve generation using the Andrew+ Pipetting Robot, with cloud-native OneLab Software, for accurate and reproducible LC-MS quantification of nitrosamine impurities, using an ACQUITY UPLC I-Class PLUS System and Xevo TQ-XS Mass Spectrometer.
Sample preparation is often the most time consuming, but critical, step for the laboratory analyst. Any errors made in the sample preparation step will follow throughout the analysis, potentially causing high variability in performance and in extreme cases failed analysis. In addition, development, optimization, and execution of these assays can prove to be time-consuming and difficult to transfer between scientists and laboratories.1
Creation of calibration standards and quality control (QC) samples is a requirement in all quantitative analysis. These samples are generated using the basic technique of sequential (or serial) dilution of the analyte(s) from a concentrated solution(s).2 While often not difficult, preparation of these samples is repetitive and time-consuming, requiring consistent preparation to ensure the reliable performance of the analytical method. This makes it ideal for incorporating lab automation for this task. It allows the analyst to do other tasks, streamlines the sample preparation process, reduces the potential of human error, and ensures consistent analytical method performance.
The objective of this application brief was to develop an automated sample preparation using the Andrew+ Pipetting Robot for the LC-MS quantification of N-nitrosamine impurities and demonstrate its performance. Due to their carcinogenicity and incidence in a variety of commodities and pharmaceuticals, robust and reliable sample preparation and analysis for their routine quantification during and after drug development and manufacturing is required.
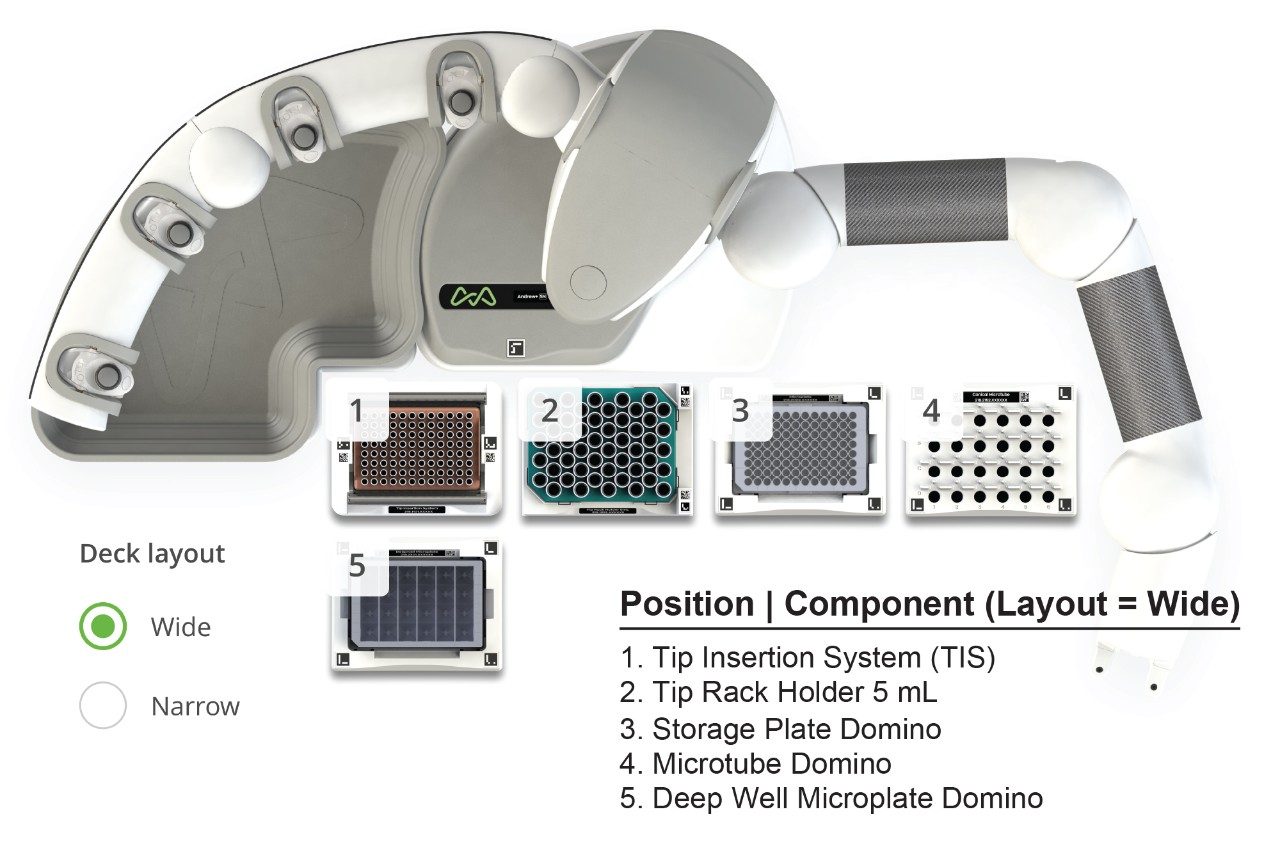
The calibration sample preparation protocol was created for the Andrew+ Pipetting Robot based on the sample preparation described in application note p/n: 720006899EN. Briefly, three individual calibration curves containing the six nitrosamine impurities NDMA, NDEA, NEIPA, NDIPA, NDBA, and NMBA were prepared in a water/methanol solution (80/20), water from a concentrated 1 µg/mL stock solution in a 96-well sample collection plate (p/n: 186005837). The reagent preparation protocol and calibration line protocol were generated in OneLab, the cloud-native software used for both designing and executing protocols on the Andrew+ Pipetting Robot. This developed protocol not only executes all steps for the calibration line preparation, ranging from 0.025–100 ng/mL, but also provides all the information for consumables and reagents used during the preparation. Figure 1 highlights the OneLab serial dilution/calibration curve generation (0.025–100 ng/mL) used in this analysis. Following nitrosamine calibration curve generation, the analysis plate was sealed with a Silicone/PTFE 96-well cap mat (p/n: 186006332). LC-MS/MS analysis of the prepared sample was performed using a Waters Xevo TQ-XS Tandem Mass Spectrometer coupled to an ACQUITY UPLC I-Class PLUS System under MassLynx Software control. Full details of the LC-MS method are also described in application note p/n: 720006899EN.

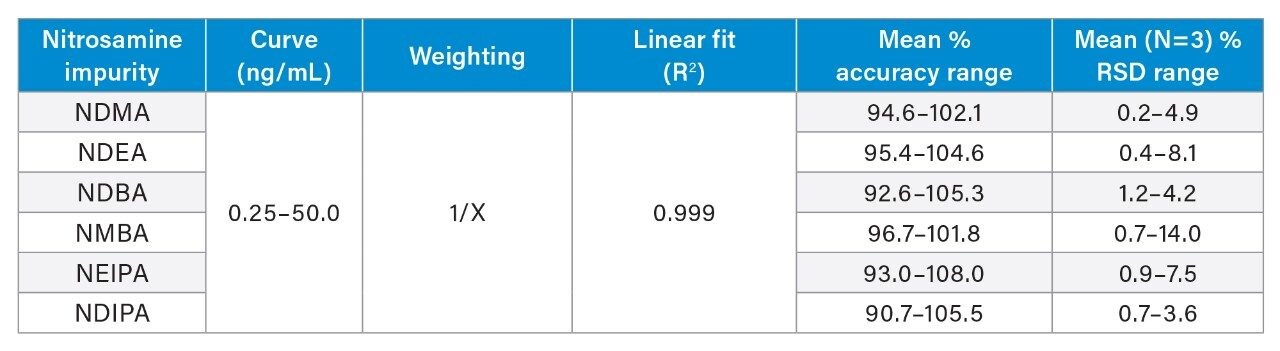
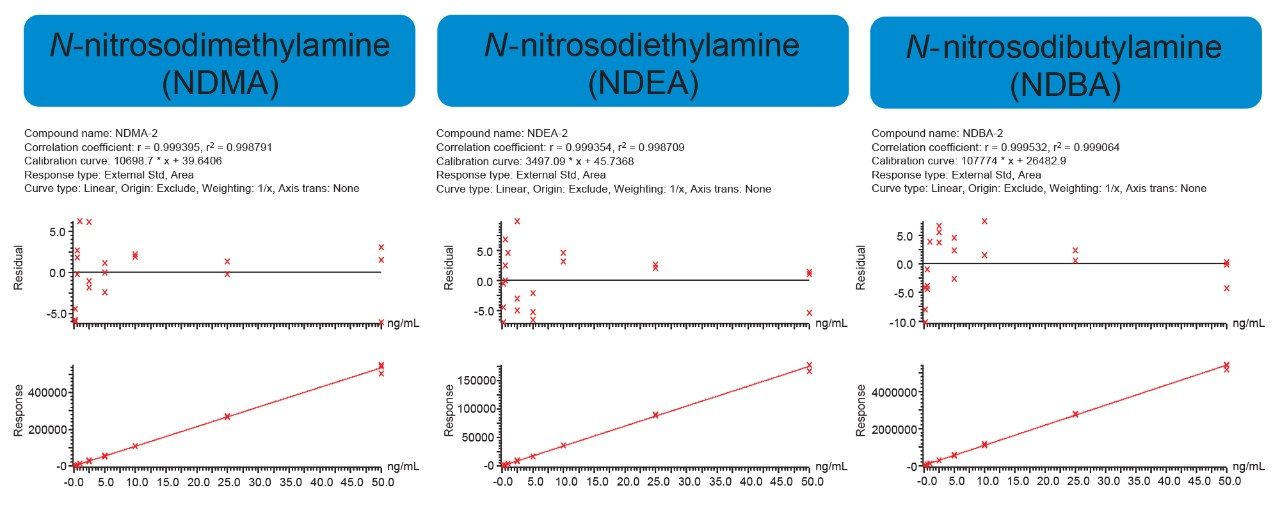
In accordance with small molecule analytical method development guidance,3,4 a developed assay must be able to demonstrate linearity (correlation coefficient or R2 ≥0.98), accuracy (±15%), and precision (±15%). These criteria were easily achieved for all six (6) N-nitrosamine impurities using the OneLab sample preparation method executed on the Andrew+ Pipetting Robot and subsequent LC-MS analysis. Mean (N=3) calibration performance is demonstrated in Table 1. For all six N-nitrosamine impurities, linear dynamic ranges were 0.025–50 ng/mL, with linear fits ≥0.999 using simple 1/x weighting. Additionally, mean accuracy values ranged from 90.7–108.0% with RSDs from 0.2–14.0%, indicating a highly accurate and reproducible method. Representative calibration curves for three of the six nitrosamine impurities are highlighted in Figure 2.
This automated method created with OneLab, and performed on the Andrew+ Pipetting Robot, developed herein, was able to perform simple serial dilution to generate calibration curves for analysis of nitrosamine impurities with no manual intervention by the analyst. This broadly applicable method can be use independently, or as part of a more complex sample workflow for robust LC-MS quantification analysis. In addition, OneLab methods are traceable and easily transferable, ensuring method reproducibility performance: day-to-day, user-to-user, system-to-system, and lab-to-lab.
720007134, January 2021