This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates the ability of DESI-MSI to determine the gender of an individual from their fingerprint.
Analysis of fingerprints using non-imaging DESI-MS coupled to a QTof instrument (Xevo G2-XS) for gender prediction of donors.
Over the past decade, mass spectrometry imaging (MSI) has been used increasingly by researchers to investigate the distribution of metabolites, drugs, peptides, and proteins at tissue surfaces. The chemical composition of fingerprints is dependent on glandular secretions, excretion of ingested substances, and exposure to environmental chemicals. Analysis of fingerprints may thus provide native and environmental information about a donor, which is of particular interest to forensic scientists.1,2
Matrix-assisted laser desorption ionization (MALDI) mass spectrometry has been widely applied to the analysis of fingerprints. Bradshaw et al. were able to assess the presence of blood by detecting the ions from haeme and haemoglobin.3 Gender was predicted with 85% accuracy by comparing protein and peptidic profiles.4 Caffeine, fatty acids, and amino acid detection has also been reported.1 MALDI used in imaging mode (MALDI-MSI) allowed investigation of spatial variations in the chemical composition of finger marks; additionally, localization of sweat pores has been achieved.1
To a lesser extent, desorption electrospray ionization mass spectrometry (DESI-MS) has been used, and lipidic endogenous compounds – as well as spiked exogenous drugs and explosives – have been detected in fingerprints.5
Recently, there has been a significant increase in the application of DESI because this soft ionization technique can be performed under ambient environmental conditions. Furthermore, it requires little to no sample preparation and is minimally invasive, which makes it suitable for direct tissue analysis. DESI-MSI is compatible with both the Waters SYNAPT G2-Si and Xevo G2-XS Mass Spectrometers.

Two men and two women provided each six fingerprints. These were collected at the same time, from different fingers, on Polysine glass slides (Thermo Fisher Scientific). DESI analyses were performed using methanol–water (95/5 v/v) at a flow rate of 0.7 µL/min, with 1 µg/mL raffinose as a lockmass compound and 4 bars of N2 as nebulizing gas. The samples were analyzed in profiling mode, following a line drawn crossing the fingerprint (Figure 1), or in imaging mode, with a pixel definition of 100 µm. Combined spectra were lockmass-corrected, background subtracted, and aligned for multivariate statistical analysis.

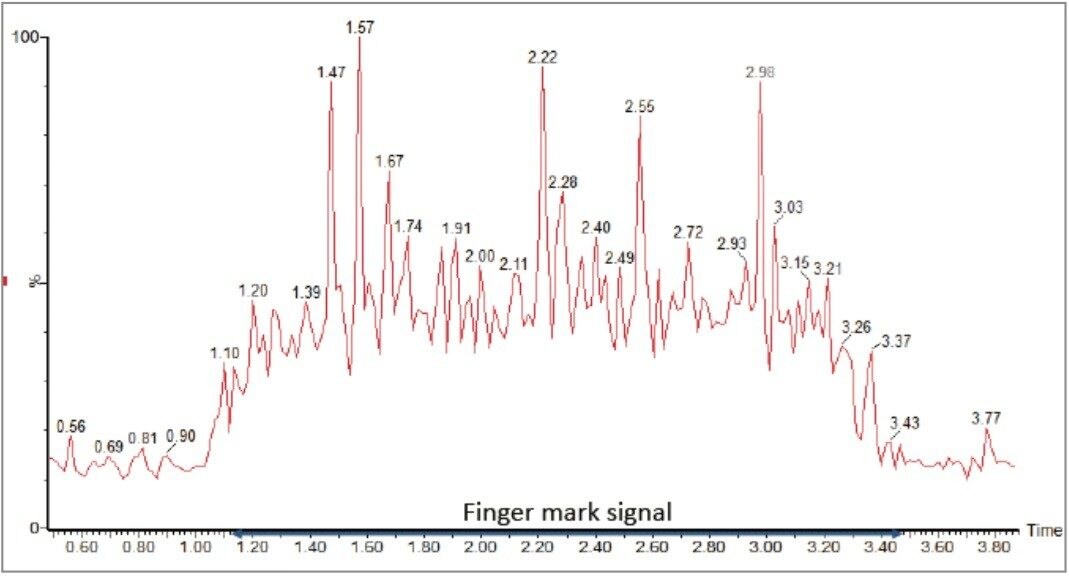
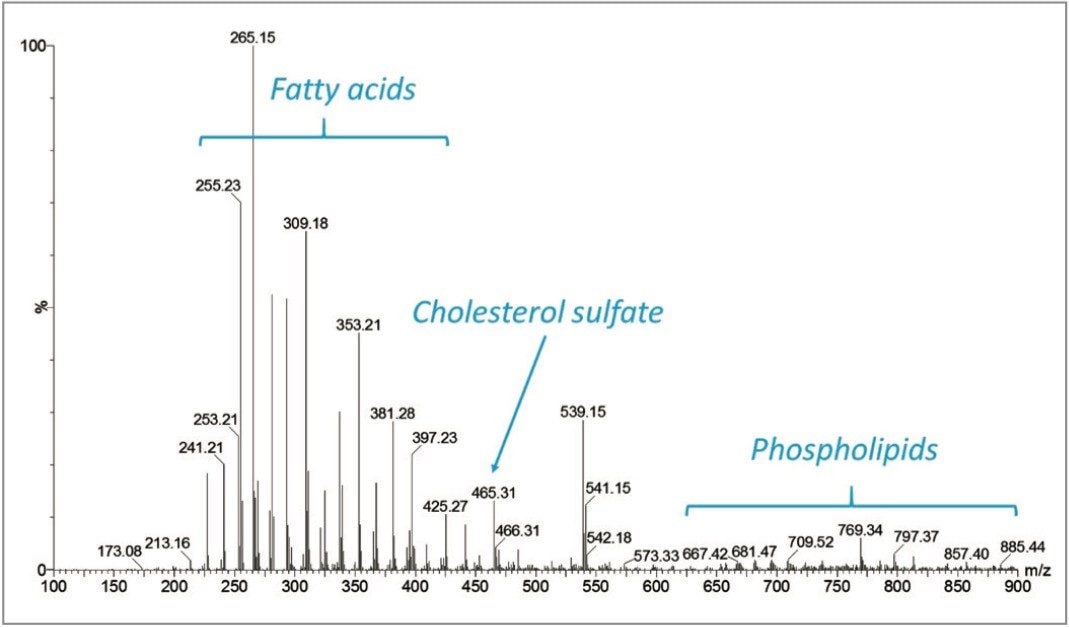
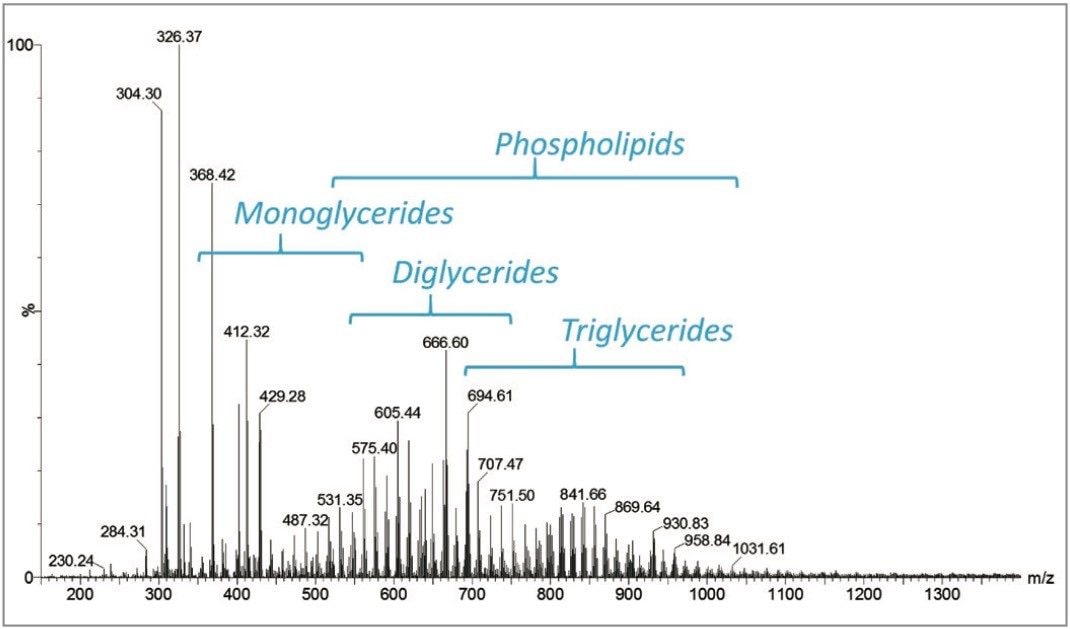
Profiling experiments by DESI on multiple fingerprints were carried out in positive and negative ionization mode. Figure 2 shows the total ion count (TIC) of the line of acquisition across one of the fingerprints in positive mode. The corresponding MS spectrum is displayed in Figure 3 showing the presence of mono, di, and triglycerides, as well as phosopholipids. In negative ionization mode, fatty acids, cholesterol sulfate, and phospholipids were detected (Figure 4).
Principal Component Analysis (PCA) allowed a good separation of male and female subjects, using only the two first principal components (Figure 5). The prediction of the donors’ gender taking 20% of the dataset out using a PCA-LDA (Linear Discriminant Analysis) model was 96% accurate in negative mode (Table 1) and 100% in positive mode (Table 2).
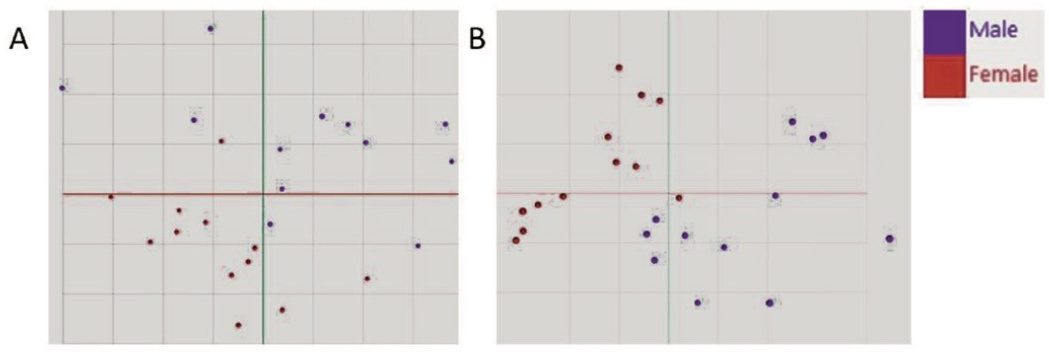
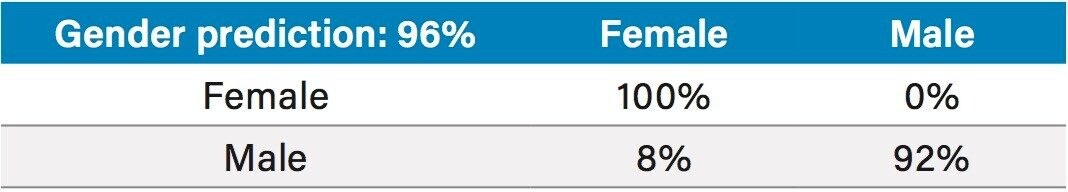
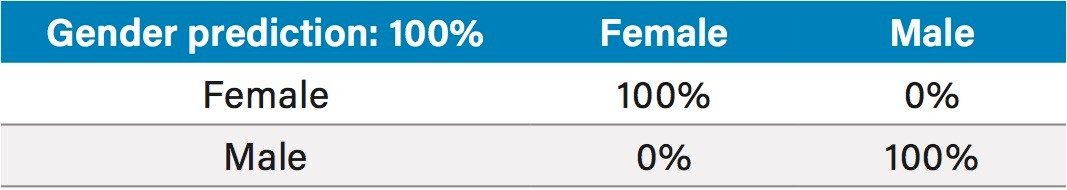
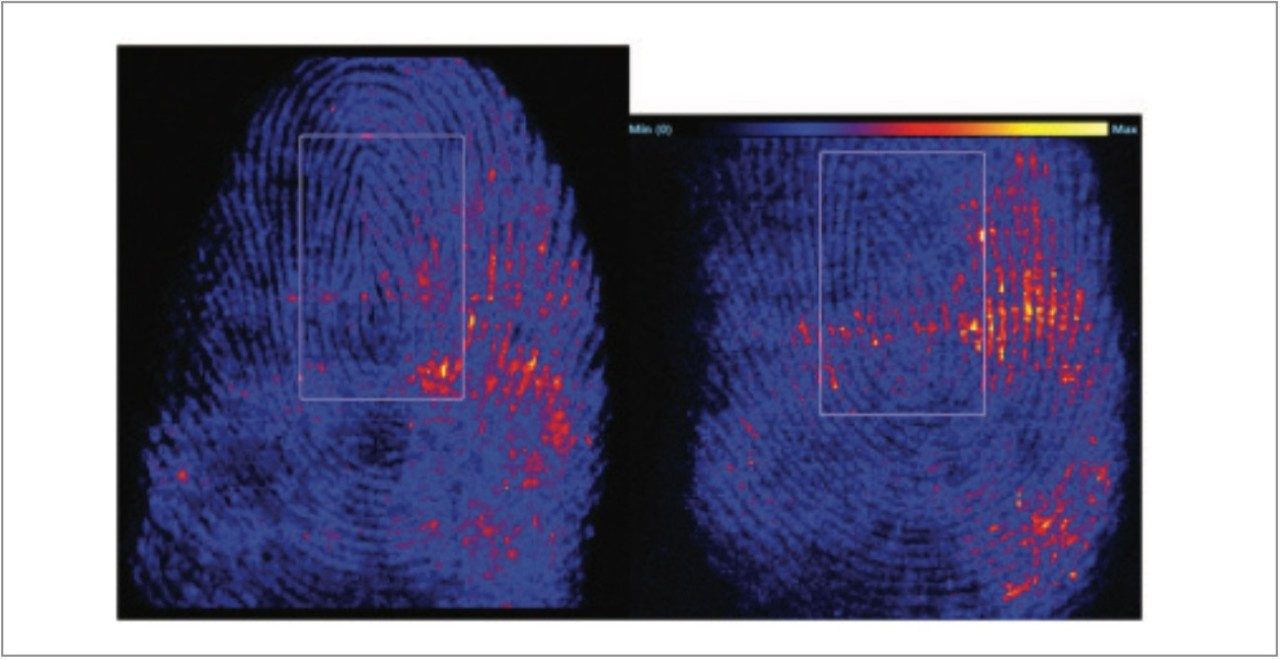
Figure 6 displays ion images of molecules detected in either ionization mode, demonstrating the structural features of the fingerprint, which can be used for the identification of the donor.
DESI-MS using the Waters Xevo G2-XS QTof together with a Prosolia DESI 2D source allows the ambient, direct, and precise analysis of fingerprints. The detection of a great number of compounds belonging to various classes of lipid enables lipid-based investigations. Lipidomic profiles obtained here allowed the prediction of the donors’ gender.
The advantages of this DESI-MSI include:
720005884, July 2018