This application note demonstrates the ability of Progenesis QI for proteomics to achieve qualitative and quantitative results for both labeled and label free quantitation workflows.
Data acquired by means of data-dependent acquisition and data-independent acquisition strategies show consistency and precision in their reported quantitation values, independent of the quantitation methodology adopted.
LC-MS is routinely applied for the qualitative and quantitative analysis of complex proteomes, to characterize biological processes, and understand disease states. Experiments can readily generate large, complex data sets in effect making the analysis and interpretation of results rate-determining steps. This lag factor has created a demand for improved data-analysis systems including efficient and accurate data-compression routines, intuitive software interfaces with menu-guided workflows, flexible experimental designs without sample-number restrictions, consistent peak detection for improved accuracy and precision, complete data matrices without missing values for reliable statistics, and the ability to analyze fractionated samples. These features, core to the Progenesis QI for proteomics solution, will be illustrated for both the identification and quantification of isotopically-labeled and label-free proteomic data sets.
|
LC system: |
nanoACQUITY UPLC |
|
Column: |
5 μm Symmetry C18 , 180 μm x 20 mm 2G Trap Column and a 1.8 μm HSS T3 C18 , 75 μm x 150 mm NanoEase Analytical Column |
|
Column temperature: |
35 °C |
|
Flow rate: |
300 nL/min |
|
Mobile phase: |
water (0.1% formic acid) (A) and acetonitrile (0.1% formic acid) (B) |
|
Gradient: |
3% to 40% B in 90 min |
|
Injection volume: |
1 μL |
|
MS system: |
SYNAPT G2-Si |
|
Ionization mode: |
ESI (+) at 3.2 kV |
|
Cone voltage: |
30 V |
|
Acquisition range: |
50 m/z to 2000 m/z (DDA and HDMSE ) |
|
Acquisition rate: |
0.2 s (DDA) or 0.5 s (HDMSE ) |
|
Collision energy (HDMSE): |
5 eV (low energy) and from 19 eV to 45 eV (elevated energy) |
|
Collision energy (DDA): |
m/z-dependent ramp |
Progenesis QI for proteomics v2.0 (Nonlinear Dynamics, Newcastle Upon Tyne, UK)
ProteinLynx Global SERVER v3.0.2 (Waters Corporation, Milford, MA)
ProteoLabels v1.0 (University of Liverpool, UK)
Mascot v2.5 (Matrix Science, London, UK)
Spotfire v9.1 (Tibco, Palo Alto, CA)
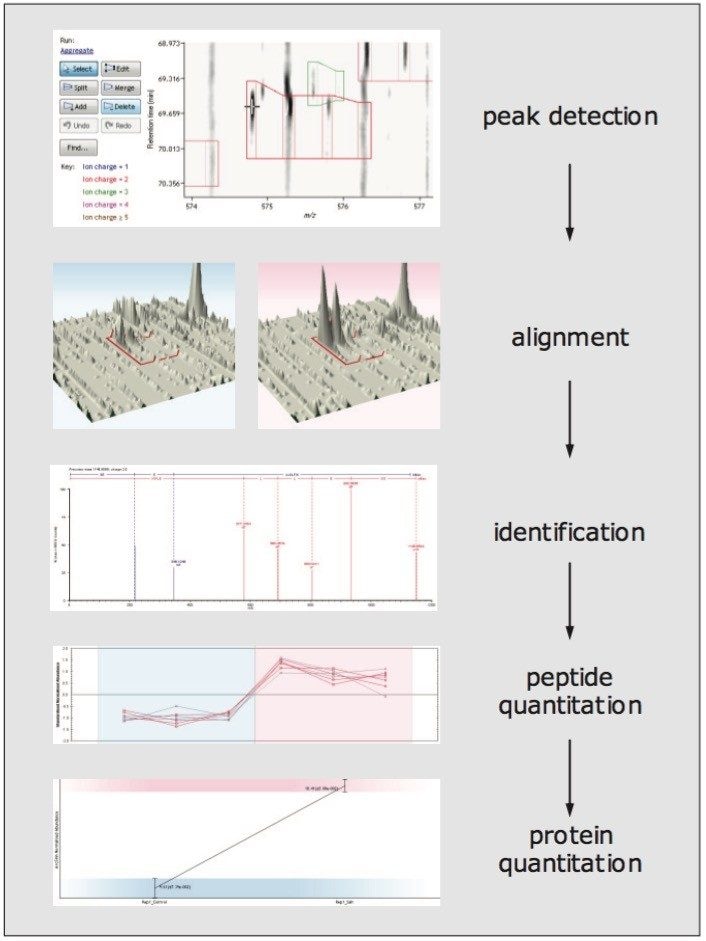
As illustrated in Figure 1, peak detection is conducted first [1]. To assess the precision of peak detection, the separate data and detected peaks and features from six technical LC-IM-DIA-MS replicates of an E. coli digest were compared, providing an average of 28,793 ± 458 detected features.
Most of the data were identified in all samples using tolerances for matching peaks of m/z ± 5 ppm, tr ± 0.5 min, and td ± 5% units, as shown in the left-hand pane of Figure 2. The top 95% raw-abundance percentile of the complete data set was considered. To improve detection across injections and samples, alignment and co-detection of peaks were conducted and an aggregate constructed. The detection boundaries of the latter are passed back to individual samples, affording a complete data matrix that improves downstream analysis via multivariate statistics.
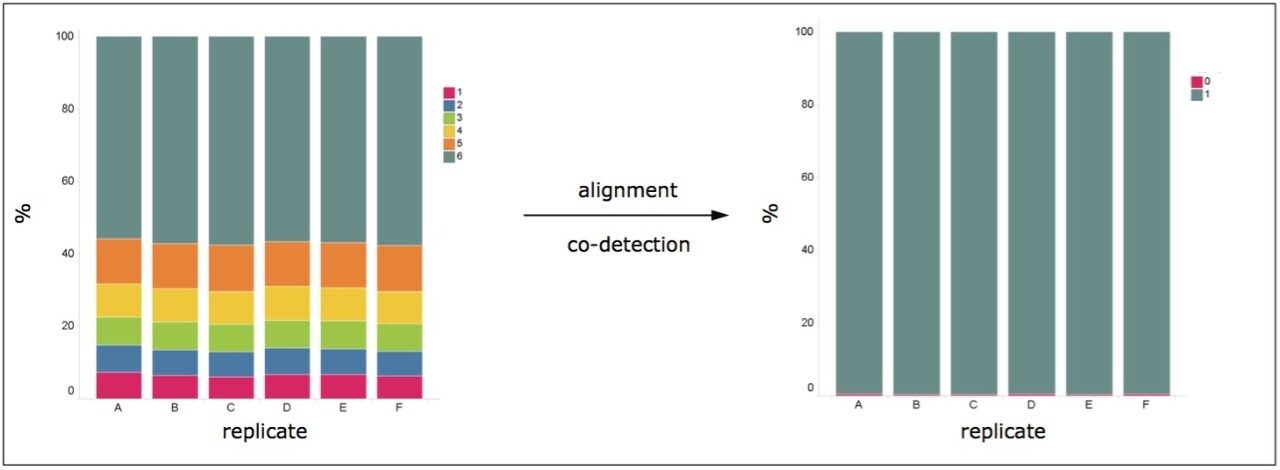
Applying this principle and the same match criteria as that used for the previously described one-to-one replicate comparisons, the vast majority of the detected features in the individual runs could be identified in the aggregate (right-hand pane of Figure 2). Comparing individual runs, approximately 55% of the features could be detected in all runs, illustrated by the light-blue sections of the Venn intersection shown at the left-hand side of Figure 2. In contrast, nearly 100% of the features could be identified in the aggregate run, as illustrated by the Venn diagrams as illustrated at the right-hand side of Figure 2. An average increase of 98.3% in co-detected features was observed.
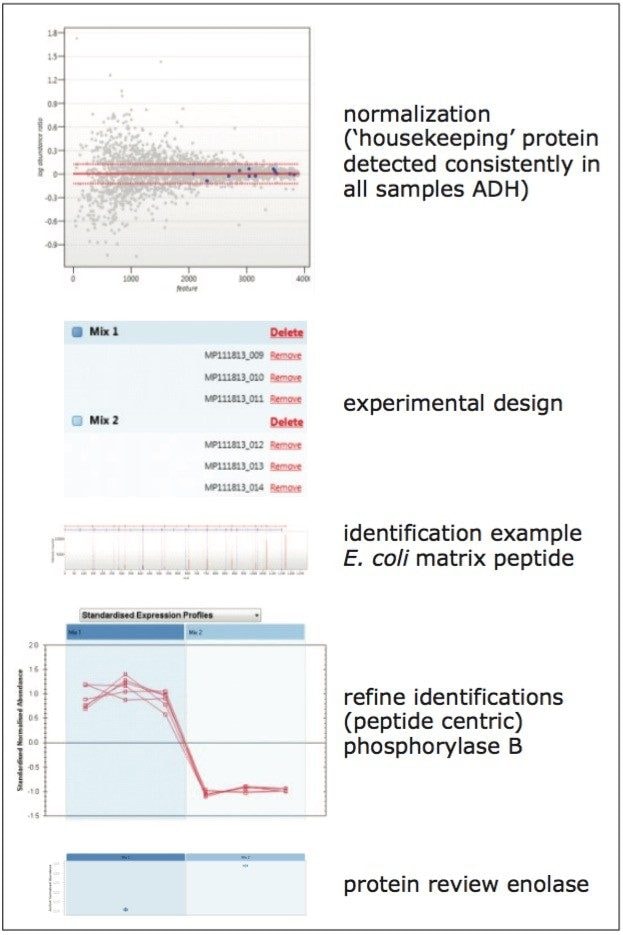
Three replicates of each E. coli sample, differentially spiked with BSA, ADH, enolase, and glycogen phosphorylase B, were also analyzed by mobility assisted, data-independent LC-MS. Figure 3 shows part of the quantitative analysis of the data, including a results summary for the protein spikes, using ADH as the internal standard. All spikes were confidently quantified with expected ratios, as specified by the manufacturer.
In quantitative workflows for which stable-isotope labeling is used, such as SILAC or dimethyl-labeling experiments, peptide pairs are expected to have similar retention and ion-mobility drift times. The results in Figure 4 illustrate the detection of a dimethyl-labeled peptide pair showing a massspectrum detail (a), a section of the chromatographic separation with a partially resolved pair highlighted in red (b), and the ion-mobility separation (c) from a human cell-line sample. As expected for dimethyl-labeled peptides, the chromatographic apices are offset, but cross sections and drift times are very similar. Peptide and protein quantification was conducted with ProteoLabels.
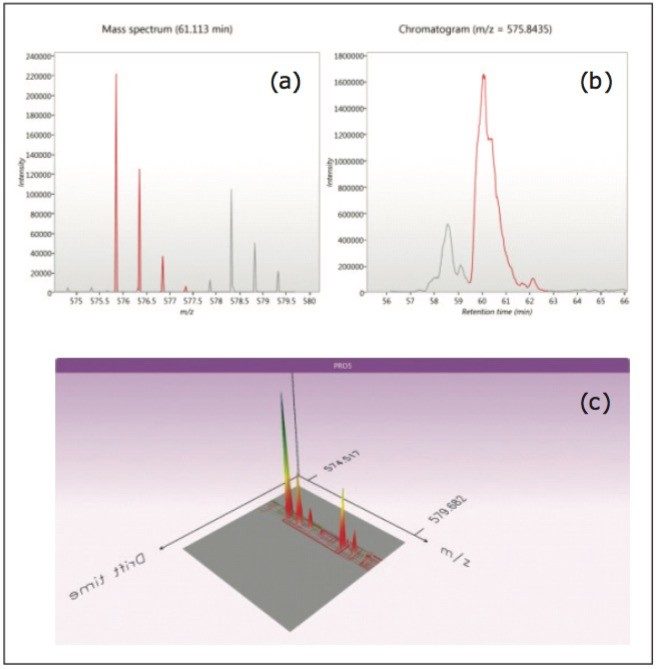
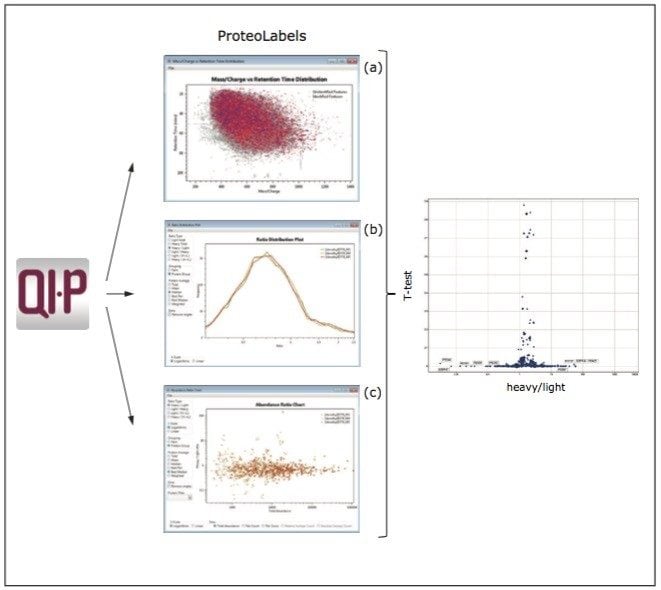
Figure 5 shows three ProteoLabels visuals including dimethyl-pair identification mapped onto a m/z versus tr contour plot (a), three technical replicate overlaid ratio distributions (b), and an abundance ratio chart highlighting the precision of the replicate experiments. A quantitative, protein-centric graphical summary of the experiment is shown on the right, with Student’s t-test p-value versus log2 fold change and the number of quantifiable peptides represented by size.
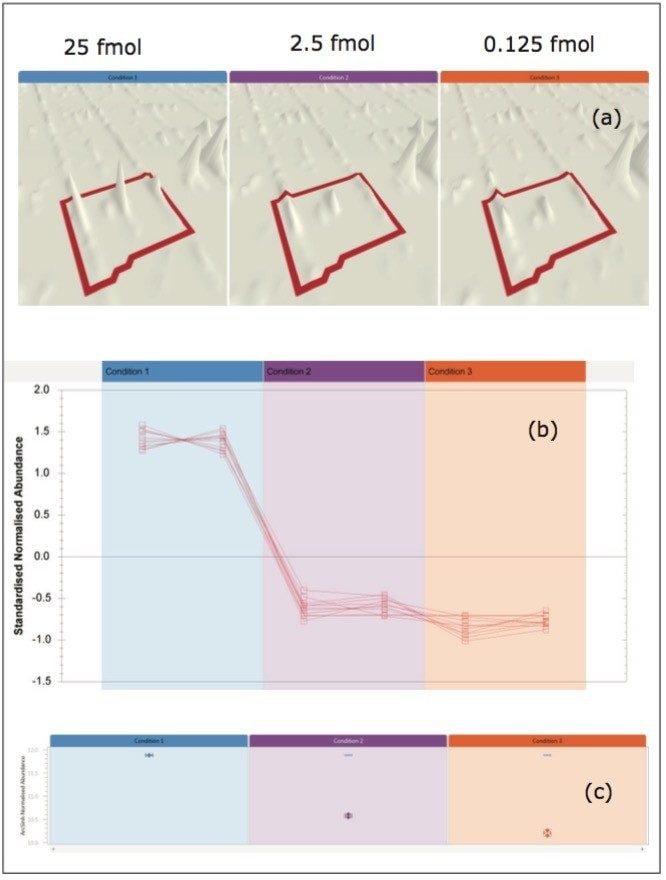
Progenesis QI for proteomics also affords the label-free quantitation of DDA data. Figure 6 shows the detection and subsequent results for the labelfree quantification of one of the UPS1 standards (SYUG_HUMAN) that was differentially spiked in a tryptic digest of yeast, with on-column amounts of 25, 2.5, and 0.125 fmol, respectively, and analyzed by DDA. The isotopic clusters with peptide and protein distribution profiles are presented. Pane (a) of Figure 6 illustrates the detection of one peptide precursor, with averages for the duplicates of the three measured samples. Pane (b) represents the intensity distribution profiles of all peptides identified to SYUG_HUMAN, with a protein-level summary, including variances, shown in pane (c).
Progenesis QI for proteomics was successfully applied for a number of “bottom-up” proteomic applications, including the analysis of labeled and nonlabeled data acquired in DDA or DIA mode of acquisition. Moreover, consistent peak detection and the formation of an aggregate allowed for enhanced differential and statistical analysis. Lastly, the precision and accuracy of DIA and DDA quantitation were significantly improved using a co detection based, label-free, quantitation approach.
Dimethyl-labeled HL60 human B cells were kindly donated by Bobby Florea, Bio organic Synthesis, Faculty of Science, Leiden Institute of Chemistry, the Netherlands.
ProteoLabels software was provided by Andrew Collins and Andy Jones from Institute of Integrative Biology, University of Liverpool, United Kingdom.
720005239, December 2014