For research use only. Not for use in diagnostic procedures.
This application note presents a simplified approach for matrix factor determination for varying levels of hemolysis using the spiked-experiment approach in UNIFI.
The capabilities of UNIFI Software allowed for easy matrix factor determination using a spiked experimental approach.
The reliability of analytical data, the basis for critical toxicological and efficacy findings, is an essential part of bioanalysis. LC-MS/MS is the technique of choice in quantitative bioanalysis due to the high selectivity and sensitivity it offers, as well as the time savings afforded by significantly reduced chromatographic separation and minimal sample preparation. LC-MS/MS quantitative analysis is influenced by a phenomenon called ion suppression or matrix effects, wherein matrix components present in the biological sample influence the response of the analyte under investigation. The need to adequately address matrix effects data during the method development and validation process has been clearly identified.1-3 This information is reported as matrix factor (MF), defined as the analyte response in the presence of matrix components divided by the analyte response in pure solution. As drug compounds under investigation become increasingly potent, they require lower doses for efficacy and toxicology assessment. This translates to lower limits of quantitation (LLOQ) during bioanalysis, wherein the matrix components in the sample can be present in levels that are much higher than the target analyte.
In addition, over the course of pre-clinical and clinical trials, very often a number of samples to be analyzed will contain varying degrees of hemolysis arising from erroneous processing of the blood to plasma. Therefore, it is suggested that hemolyzed samples also be considered during method development and validation to assess any potential effects arising from the matrix. For example, the current EMEA guidelines require that, in addition to six unique lots of plasma, hemolyzed plasma should also be tested for matrix effects.
UNIFI Software enables the user to easily quantify matrix factor via two methods: post-column infusion and using a spiked experiment. The software is designed to do all necessary calculations and data summaries that a user requires, removing the need for other software packages such as Excel. In this application note, we present a simplified approach for matrix factor determination for varying levels of hemolysis using the spiked-experiment approach in UNIFI.
|
System: |
ACQUITY UPLC |
|
Column: |
ACQUITY UPLC BEH C18, 2.1 x 50 mm, 1.7 μm |
|
Flow rate: |
600 μL/min |
|
Column temp.: |
45 °C |
|
Mobile phase A: |
0.1% Formic acid |
|
Mobile phase B: |
Acetonitrile |
|
Gradient: |
5% B to 95% B over 2 min |
|
Mass spectrometer: |
Xevo TQ-S |
|
MS/MS parameters: |
|
|
Transitions: |
clopidogrel 322.1 > 212.1 d4-clopidogrl 326.1 > 216.1 |
|
Ionization mode: |
Positive ESI |
|
Capillary voltage: |
1.00 kV |
|
Collision energies: |
16 V |
|
Cone voltage: |
35 V |
Three lots of hemolyzed plasma were prepared by adding the appropriate volume of hemolyzed whole blood (human, K2EDTA) to plasma (human, K2EDTA) resulting in 5%, 10%, and 15% hemolysis (for example, 50 μL of hemolyzed blood was combined with 950 μL of plasma to yield 5% hemolyzed plasma). In addition to these three lots, non-hemolyzed blank plasma was also used in the matrix factor evaluation. Each lot of matrix was extracted in replicates of six using a protein precipitation extraction technique where 100 μL of the appropriate matrix was precipitated with 300 μL of methanol, vortex, mixed, then centrifuged. For spiked QCs, supernatant was combined with clopidogrel/d4-clopidogrel solution to yield final concentrations of 550 pg/mL, 85 pg/mL, and 8.5 pg/mL. Solutions at the same three concentrations were prepared in blank diluent (75% methanol in water).
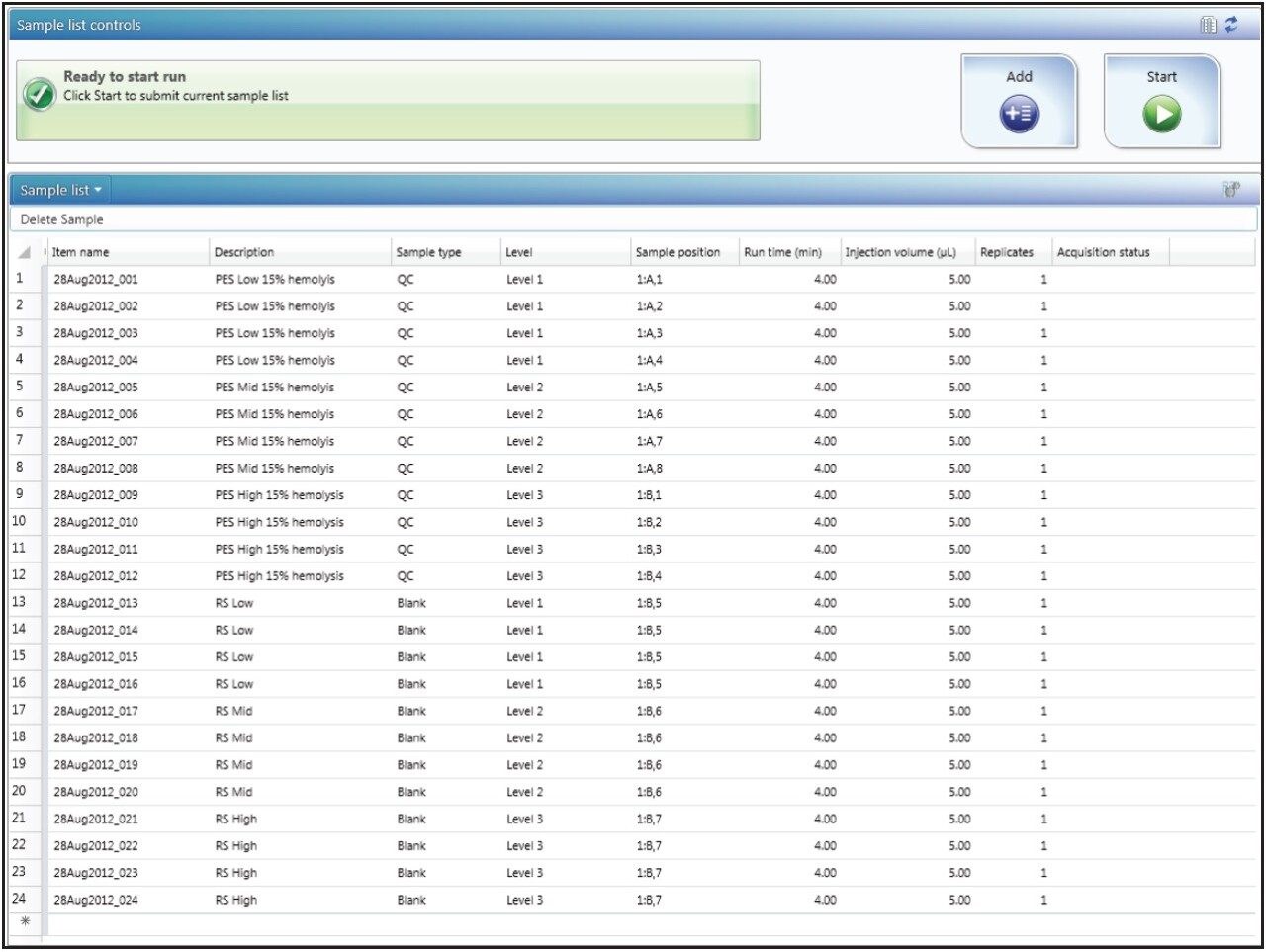
UNIFI Software architecture allows for specific analysis types to be defined, whereby the method automatically selects specific settings, parameters, and calculations that characterize the analysis type. For example, the spiked experiment MF analysis type is set up so that the software will calculate matrix factor based on the specific sample types entered by the user. In the sample list, the user inputs the solution standard as a blank, and the spiked matrix extracts as QCs, as shown in Figure 1. If there are multiple concentrations to be used for MF determination (such as low, mid, and high concentrations), these are defined as levels for both the blank and QC sample types.
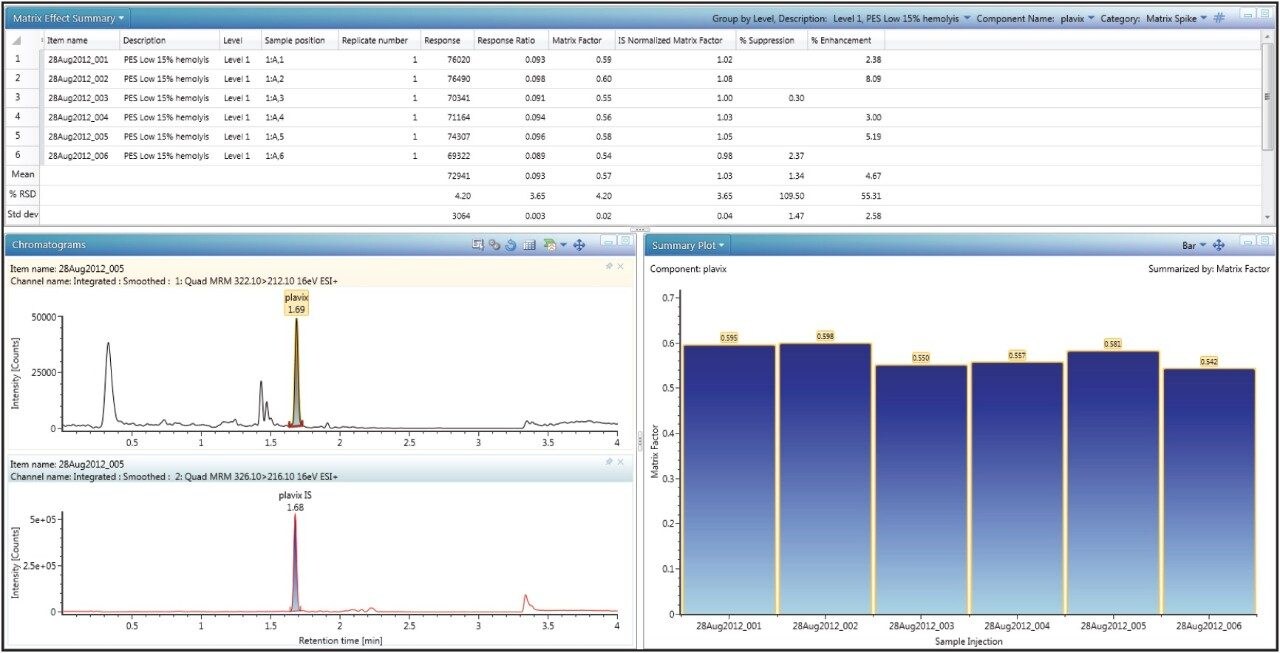
Once the data has been acquired and processed, the MFs will automatically be calculated based on the summary calculations built into that particular analysis type, therefore eliminating the need to use additional software such as Excel to calculate and summarize the MF values. By simply choosing ‘matrix factor results’ on the review tab, the calculated matrix factor data is displayed on a per component basis, as shown in Figure 2, with calculated statistics such as mean, standard deviation, and relative standard deviation (or coefficient of variation). In addition, the user can view chromatograms and summary plots within the same window.
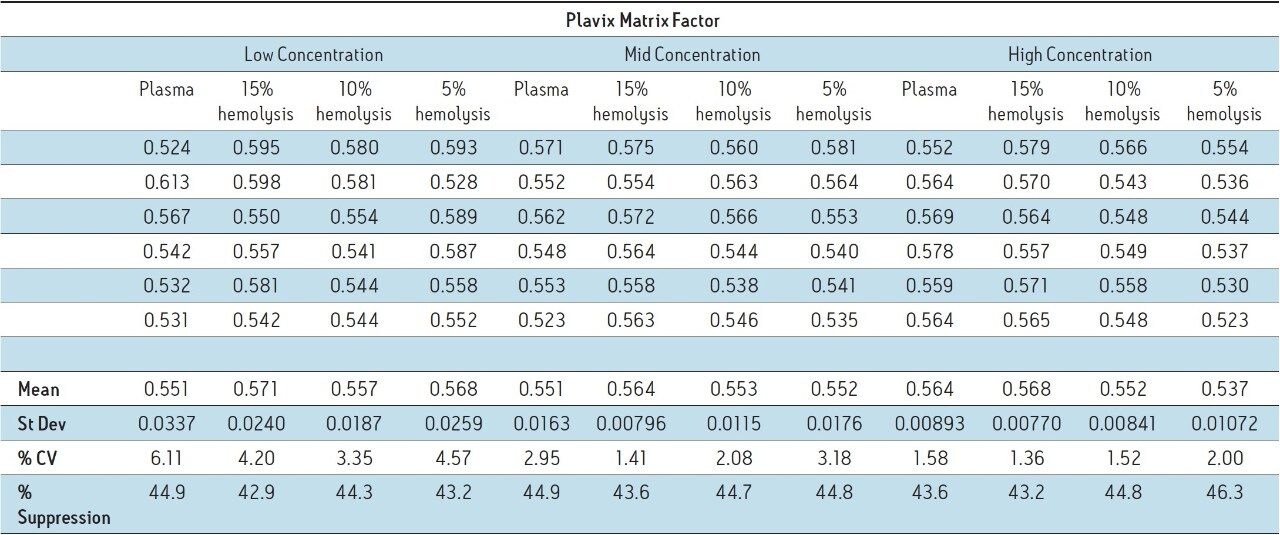
The resulting MFs for all lots of hemolyzed and non-hemolyzed matrix prepared at each concentration level are displayed in Table 1. A matrix factor with a value of less than 1 indicates suppression, a value greater than 1 indicates enhancement, and a value of 1 indicates there is no effect of the matrix on the analyte signal. The data indicates that there is no discernible variation between the different lots of matrix at each concentration level. Therefore, the varying degrees of hemolysis are not impacting the produced signal. In addition, all three concentration levels assessed showed similar matrix factor values. In fact, the mean of all 72 injections was 0.557 with a CV of less than 4.0% indicating there is no effect of concentration for this compound. However, the calculated MFs indicate suppression 42.9% to 46.3%, indicating almost half of the signal is being suppressed, which is undesirable for assays where a very low LLOQ is required. This result is not surprising given that the extraction technique used was a protein precipitation that requires relatively minimal clean up to the samples.
In a bioanalytical assay, it is preferable for a deuterated version of the analyte to be used as the internal standard (IS), since it will behave in the same manner chromatographically and spectroscopically as the analyte of interest. This includes ion suppression/enhancement since the analyte signal and the IS signal should be impacted in the same way and to the same extent. To account for this, matrix factor is often reported as IS normalized matrix factor. IS normalized matrix factor is defined as the matrix factor of the analyte divided by the matrix factor of the internal standard. Table 2 shows the IS normalized matrix factor for the four lots of matrix.
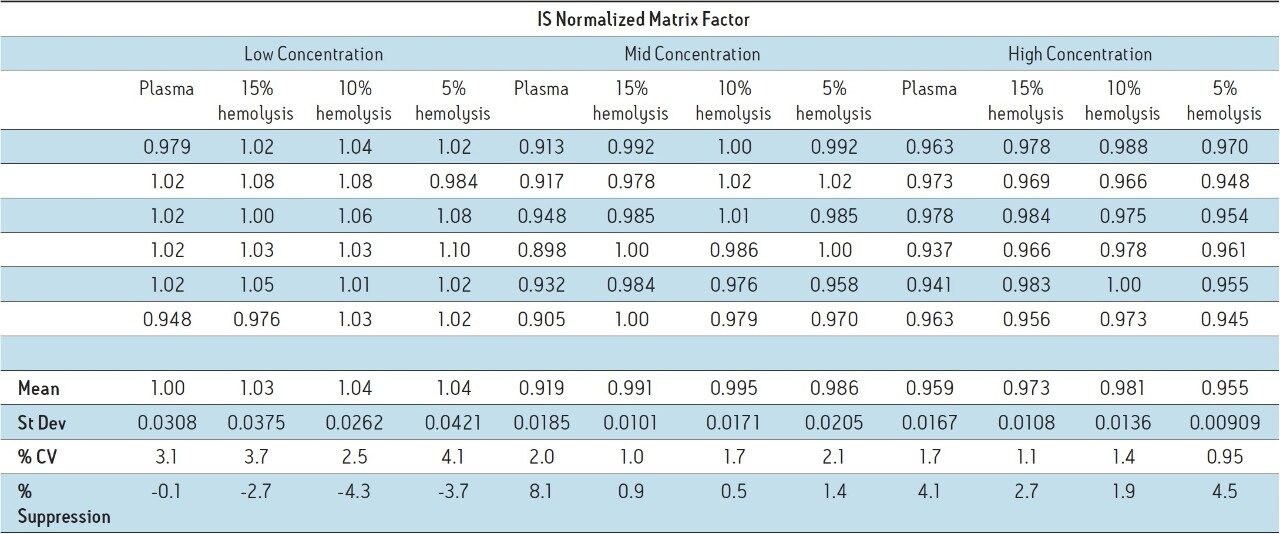
The IS normalized values are all relatively close to 1, indicating that the internal standard was suppressed to the same degree as the analyte, resulting in normalized negligible suppression/enhancement.
720004556, January 2013