In this application note, we will demonstrate the use of chemical intelligence to broaden our approach to metabolite identification.
Use of chemical intelligence provides confident answers in a more efficient workflow, enabling the drug metabolism scientist to output results quickly and make better business decisions.
A large number of routes of metabolism have been characterized by the scientific community and this has yielded a diverse knowledge base of commonly occurring metabolic transformations. The first line of metabolite identification is to proactively search for these known pathways. This knowledge combined with the exact mass measurement instrumentation has facilitated the positive identification of these metabolites in a routine fashion. This is a powerful tool in the arsenal of the drug metabolism chemist and many metabolites have been positively identified in this manner. However, for new drug entities, drug metabolism is driven by reactions that occur between the drug structures and the enzymes responsible for drug metabolism. These very specific interactions often spawn metabolites that are cleavage products of the starting structure and therefore cannot be fully understood by mining a database containing metabolites derived from a set of known compounds. If these unique biotransformations are not taken into consideration, many novel metabolites may not be positively identified. The combination of structural knowledge and metabolite identification tools is typically not used to direct metabolism identification studies up front.
In this application note, we will demonstrate the use of chemical intelligence to broaden our approach to metabolite identification. This results in an analysis that gives more confident answers in a more efficient workflow, allowing the drug metabolism scientist to output results more quickly and effectively to drive key program decisions.
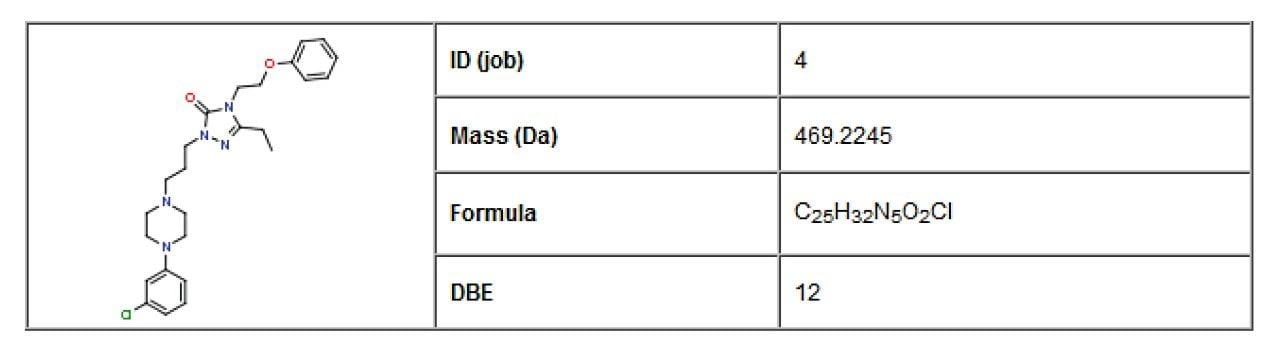
Nefazodone was chosen as a case study for this application note. An MSE data set was collected for a human liver micosomal incubation of nefazodone. MSE is a patented method of data acquisition that allows simultaneous collection of precursor and product ion information for virtually every detectable species in a mixture in a single injection. The structure of nefazodone was included in the sample list as a standard .mol file and is the starting point for the informatics workflow, as shown in Figure 1.
The structure of a drug is significantly modified through the process of xenobiotic metabolism. There are many pathways for these modifications but the process can be roughly defined as Phase I and Phase II metabolism. The distinction is made due to the chemical nature of the metabolism. Phase I is dominated by CYP450 mediated metabolism and consists of reduction, oxidation, and hydrolysis reactions. These activate the drug and prepare it for Phase II. Phase II metabolism is typically a conjugation reaction. Of specific interest are oxidative dealkylations and other complex mechanisms resulting in cleavages. These routes of metabolism provide a challenge for the drug metabolism scientist as they can drastically alter the chemical properties of the resulting metabolites, making them difficult to detect. It would therefore be advantageous to predict such metabolites in order to drive drug metabolism studies.
The use of in silico tools to predict these dealkylations is a first step in the processing of the MSE data. The drug structure is supplied and the products of dealkylation are predicted using straightforward, customizable rules. These dealkylations are then displayed in the resulting report, which provides information regarding the potential metabolic fragments, as shown in Figure 2.
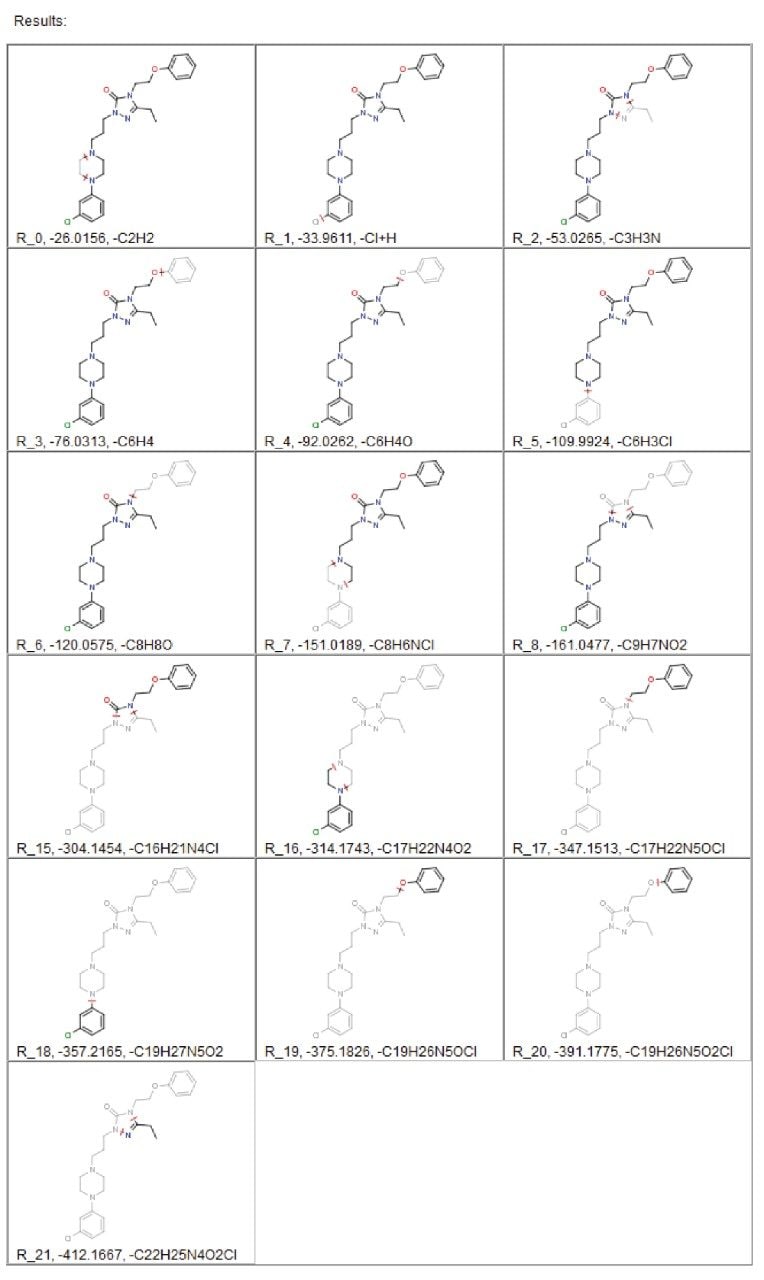
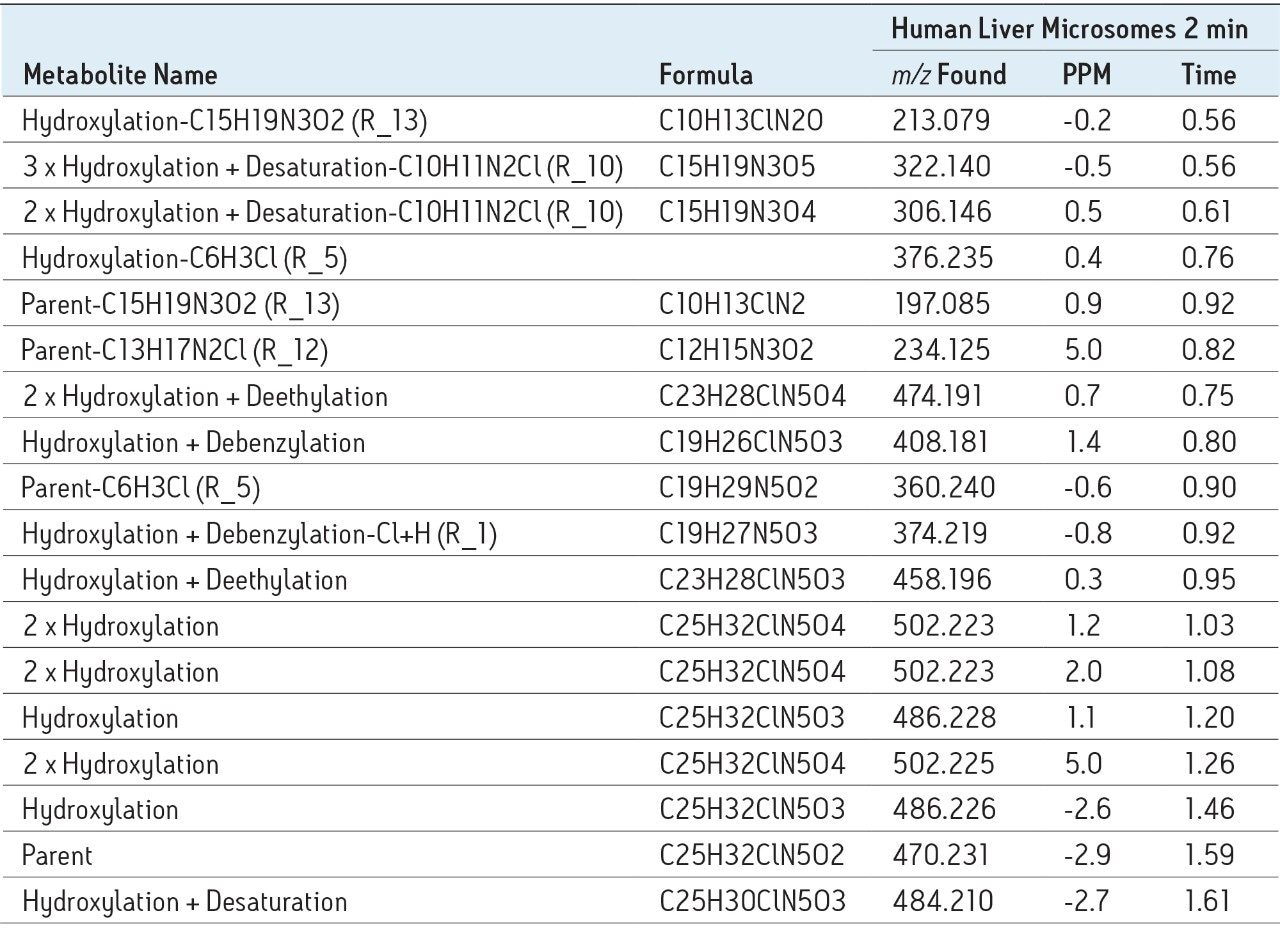
The information generated in this report includes structures, formula, and exact masses for each postulated product. This information is passed to a metabolite identification table where it is combined with a database of commonly occurring metabolic processes. This generates an “expected metabolite list,” which the software uses in targeting the specific exact mass in order to search for metabolites. In the example of nefazodone, 44% of all metabolites found were the result of a direct dealkylation or a combination of dealkylation and another metabolic route, as shown in Table 1.
The use of Mass Defect Filters (MDF) is widespread and is proven to target specific metabolites due to the chemical nature of the change in the structure of a compound. MDFs have been broadly applied to exact mass measurement analyses as a means to remove false positives and simplify a dataset reducing the complexity of the metabolite identification.1,2 A common implementation of a MDF is to target a specific mass defect using knowledge of the parent m/z only; more advanced methods work by calculating a linear change of mass defect with m/z. Both these methods have been shown to be effective in a small number of substrates; however, as the chemical structure becomes more complex these implementations have limitations. In the workflow presented here, the predicted dealkylations are utilized to drive the MDF. Figure 3 shows the mass defect of dealkylated nefazodone and a typical linear mass defect tolerance. In this graphic, there are potential metabolites that fall outside of the linear window. A much broader filter must be used in this case (shown in blue shading), reducing the usefulness significantly. When a MDF is applied using chemical intelligence it can be much more selective, eliminating more false positives hence reducing the scientist’s workload.
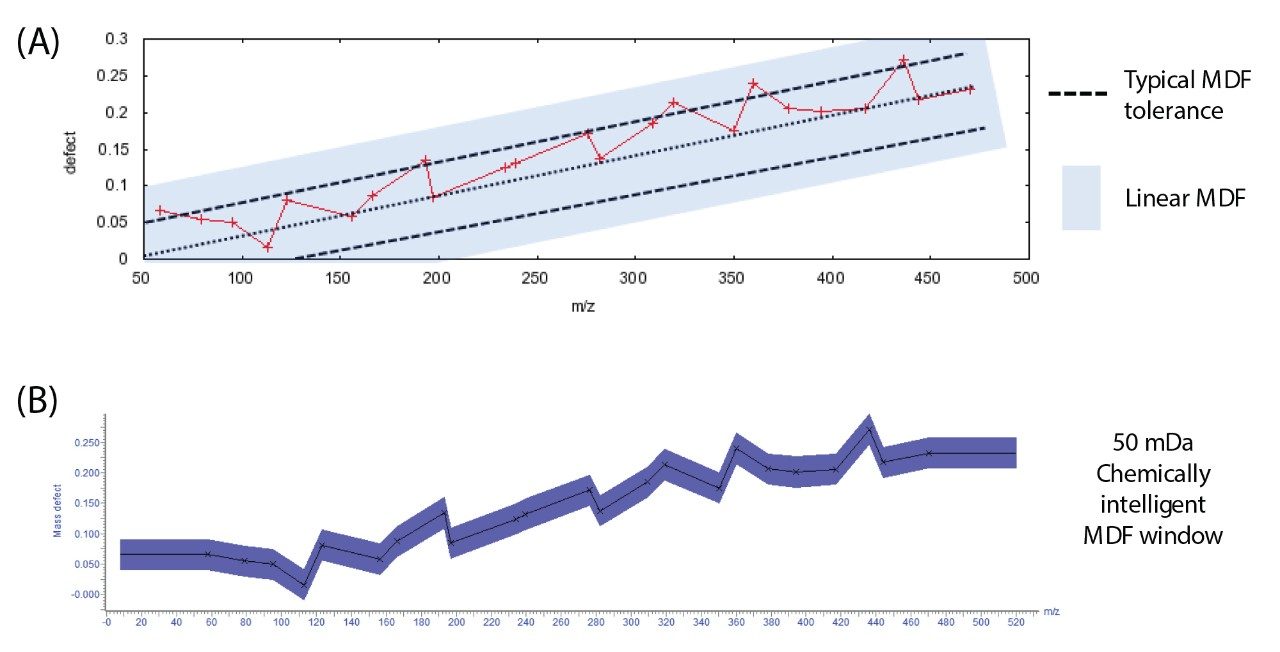
After this post-acquisition processing step the data now contain only those ions that are potentially related to the parent. Chromatographically viable peaks are then automatically extracted and searched against the comprehensive list of predicted biotransformations described previously.
A fully annotated chromatogram, shown in Figure 4, is then presented to the user in a clear and easily interpreted format.
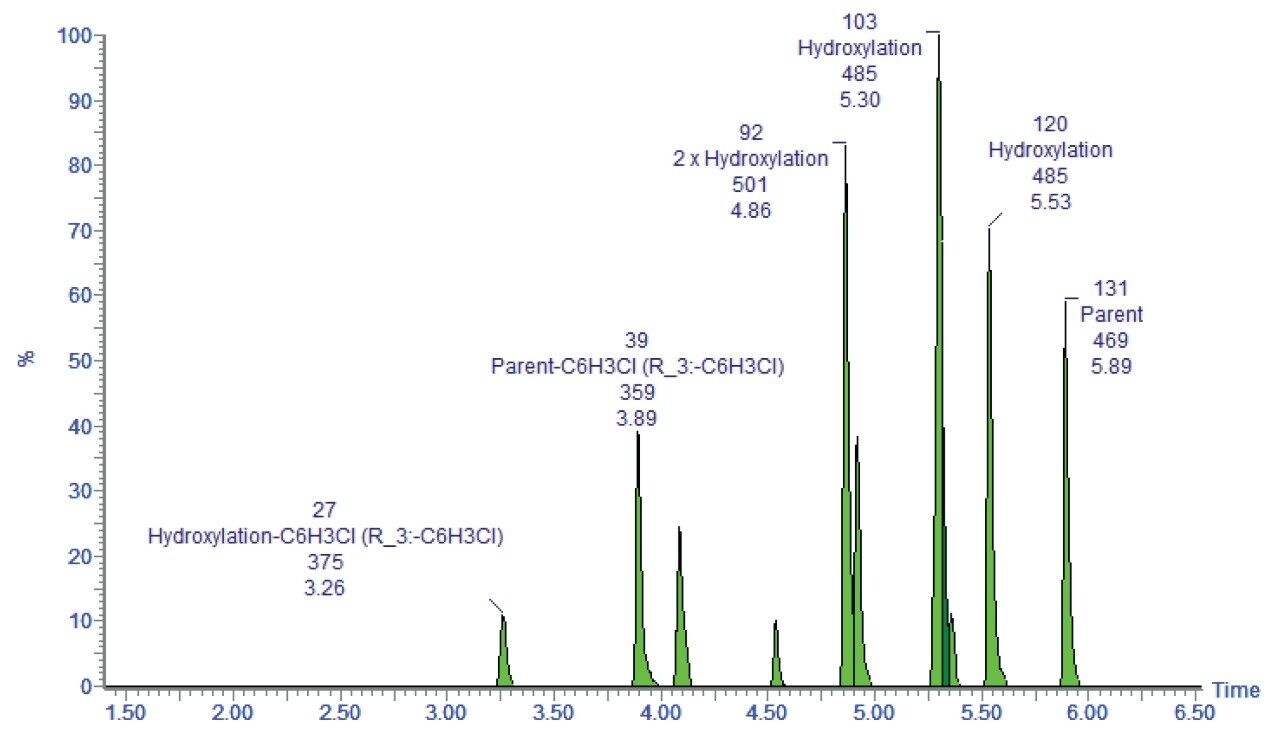
Identifying the metabolites by exact mass precursor ion gives the assignments of possible metabolic route; however, additional information is usually required to a.) Confirm the biotransformation, b.) Identify the site of biotransformation, and c.) Measure the rate of formation. MSE allows all of these measurements to be performed in a single analysis.
To answer the first of these additional requirements the chemically intelligent engine3 is employed to interrogate the product ion spectra produced for the ion corresponding to the parent compound, which automatically assigns fragment structures to the ions produced in the product ion spectrum, as shown in Figure 5.
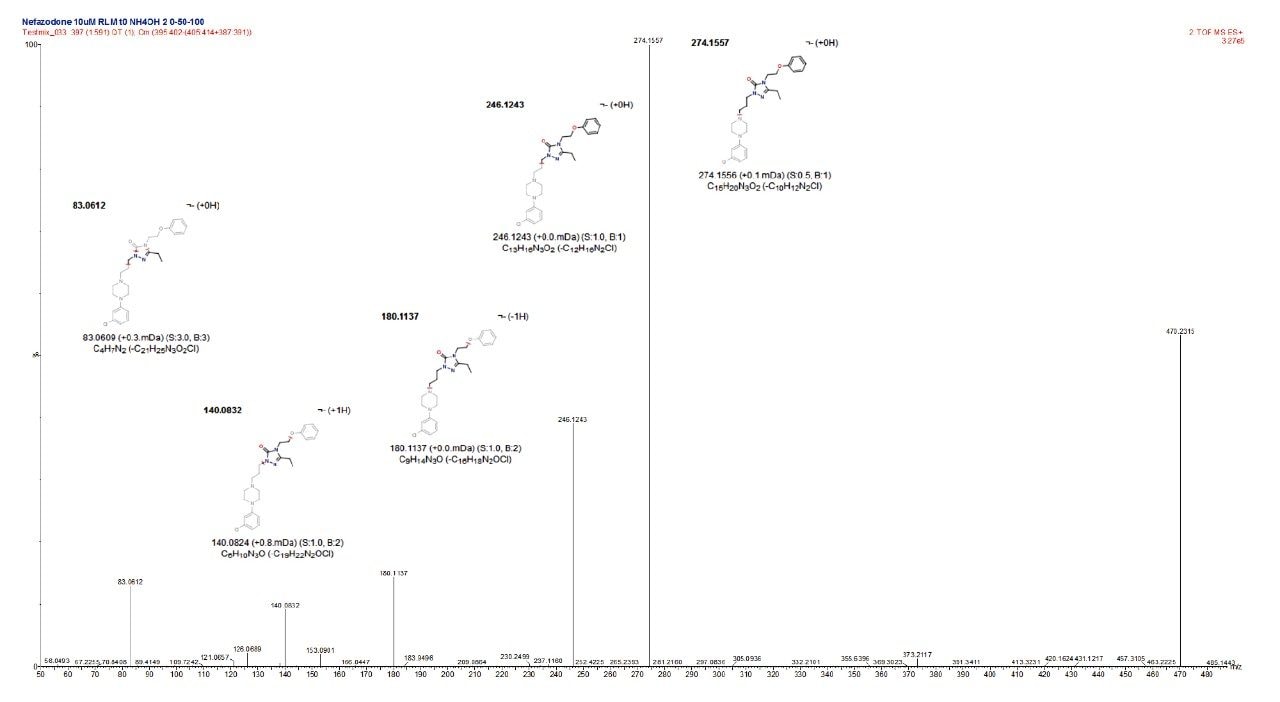
Building on this information, metabolite fragments spectra can then be automatically compared with that of the parent. This triggers an automatic interpretation of chemical shifts between the two product ion spectra to localize the region in the molecule that has changed and identify the site of metabolism, as shown in Figure 6.
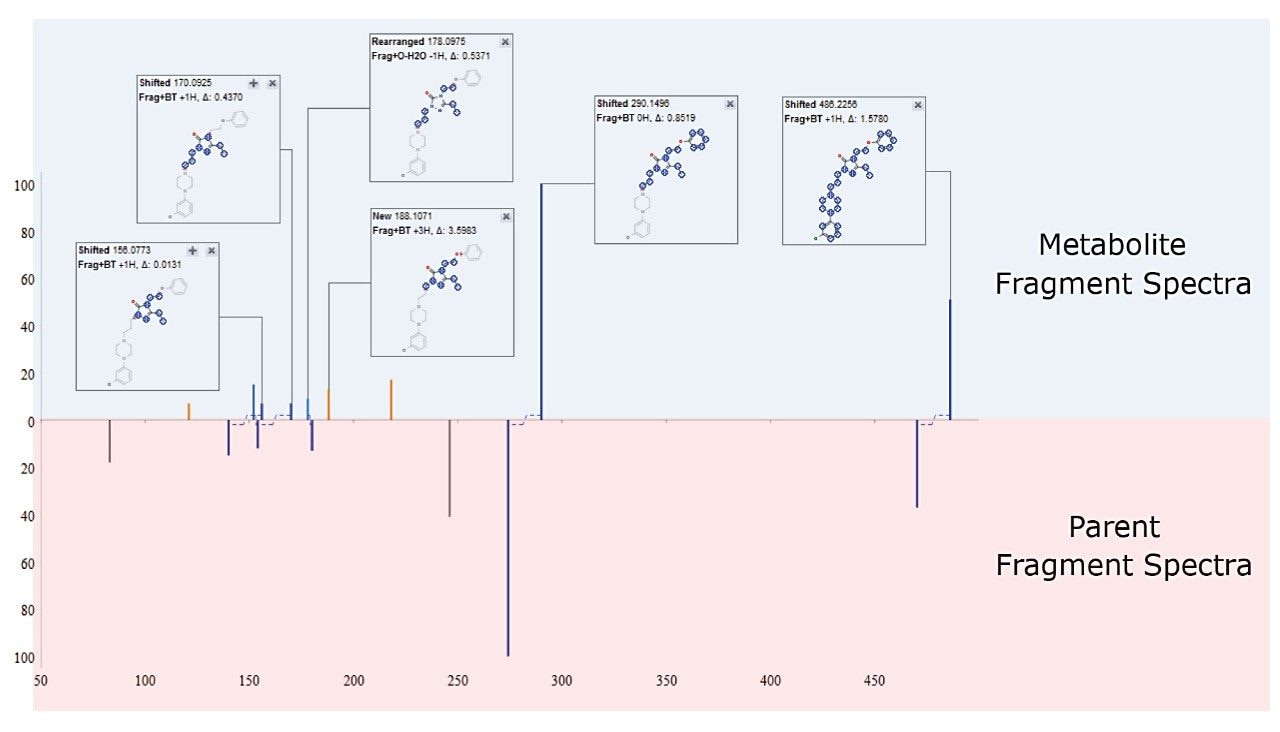
Leveraging structural knowledge of a compound is key to understanding its metabolism. In addition to the prediction of Phase I dealkylation for the metabolite identification step, the ability to leverage this chemical intelligence during in silico processing inititates processes, such as filtering and identification of biotransformations. This results in a significantly lower false positive rate for metabolite candidates, reducing the need to eliminate unrelated peaks and allowing the users time and expertise to be spent in the characterization of true metabolites. Utilizing these techniques vastly reduces the time taken to perform metabolite identification and allows critical decisions to be made earlier, driving the drug discovery and development process.
720003943, April 2011