Quantitative bioanalytical methods development by LC-MS/MS is complicated by the interference of the matrix, as the analyte response can differ significantly within a matrix.1 It is necessary to develop robust analytical methods to ensure the long-term integrity of the results. This application note describes the use of a simple algorithm to estimate the matrix factor automatically.
The TargetLynx Matrix Calculator provides a convenient way to evaluate the matrix factor during the development of bioanalytical methods. This is a helpful technique to identify methods that may be susceptible to matrix effects that could potentially affect long-term method robustness.
In quantitative bioanalysis, the analytical technique of choice is LC-MS/MS due to the high sensitivity and selectivity that it affords. Quantitative bioanalytical methods development by LC-MS/MS is complicated by the interference of the matrix, as the analyte response can differ significantly within a matrix.1 It is necessary to develop robust analytical methods to ensure the long-term integrity of the results.
Matrix effects, resulting from coeluting matrix components that compete for charge in the ionization process, manifest themselves as suppression or enhancement of the analyte signal. Matrix effects are caused by numerous factors:
All of the above can cause significant errors in the accuracy and precision of bioanalytical assays.2
As part of the method validation process, it is necessary to measure the ion suppression due to the matrix and calculate a matrix factor. This is often a time-consuming process – often requiring the transfer of the MS data to external software programs. This application note describes the use of a simple algorithm to estimate the matrix factor automatically.
Using the TargetLynx Application Manager, three sample types are defined in the sample list:
The solvent and matrix blank experiments (Figure 1) are run multiple times and the coefficient of variation (CV) is calculated when the data is processed.
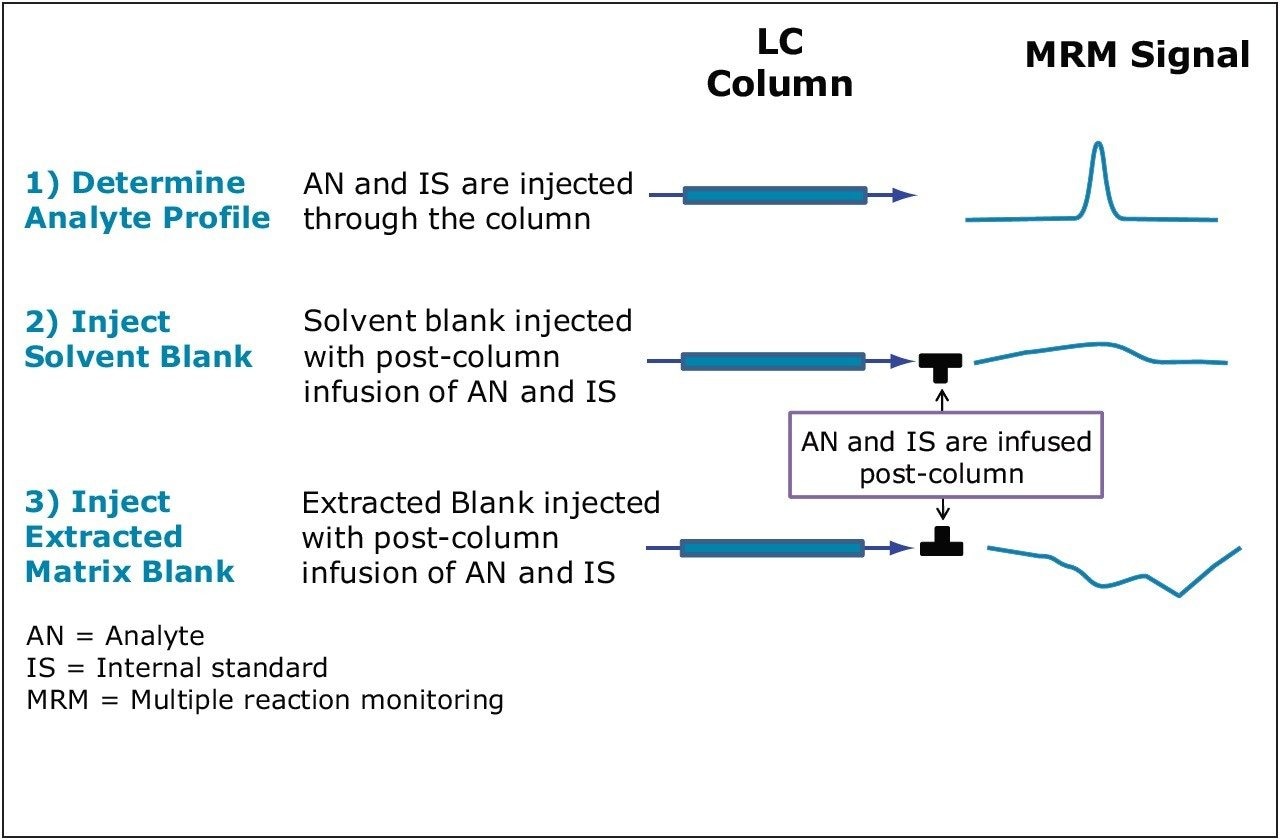
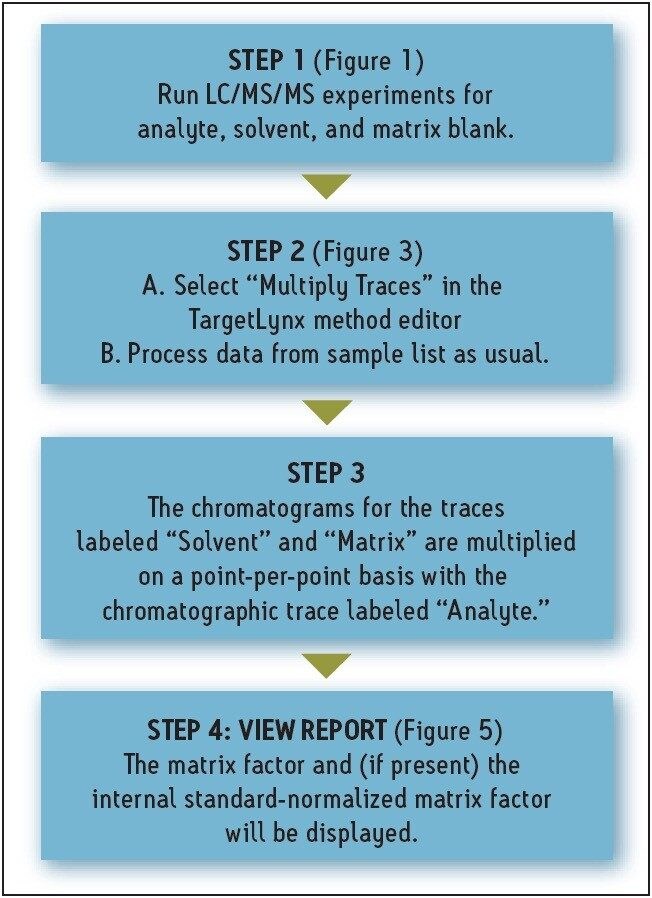
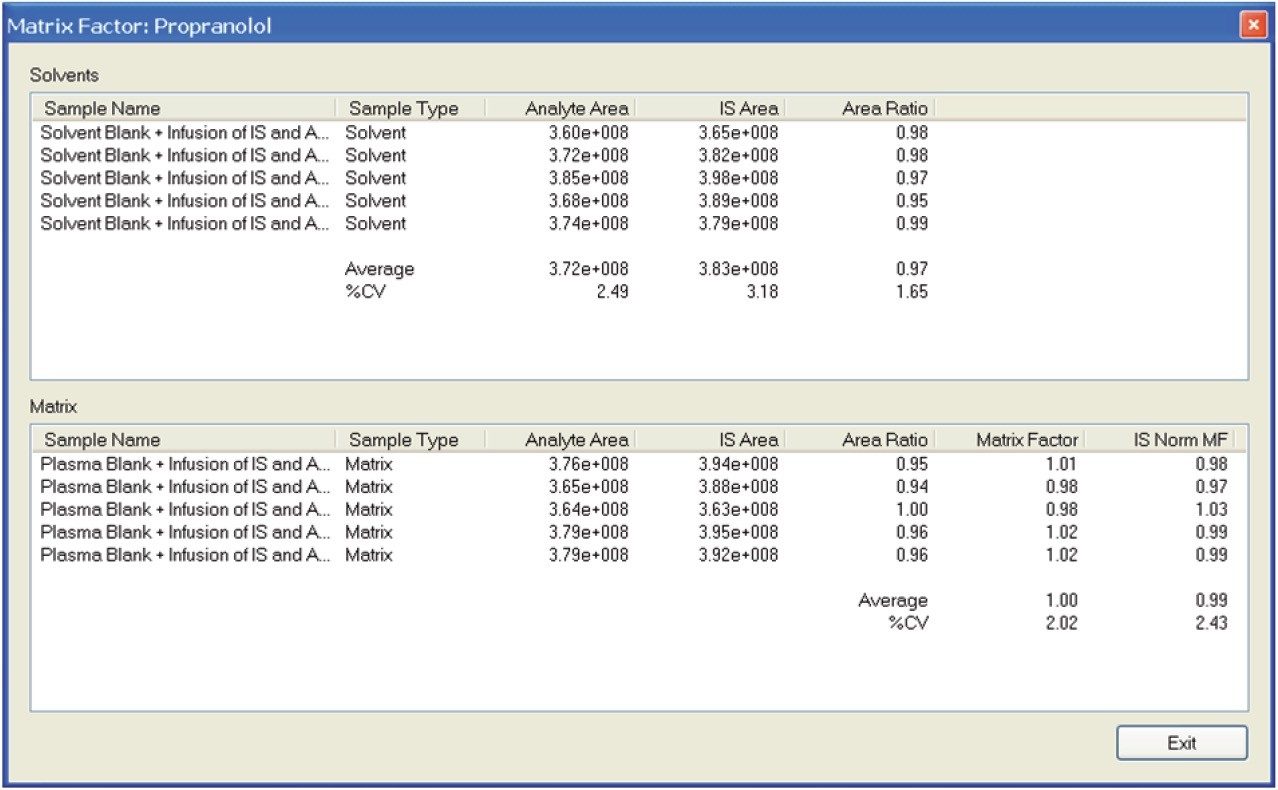
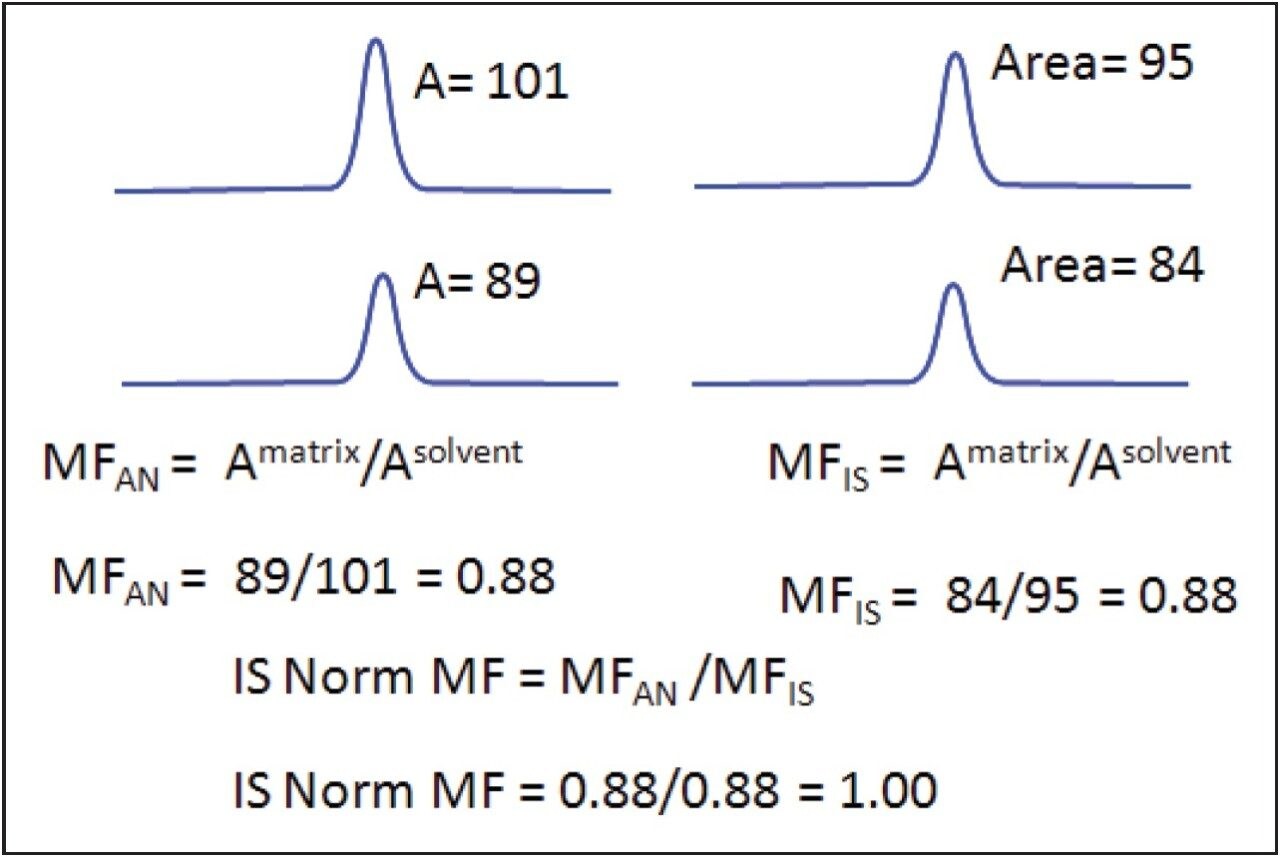
Figure 2 illustrates the workflow for calculating the matrix factor. A sample matrix factor calculation is shown in Figure 4. (MF= Matrix Factor, A = Area). This formula is incorporated into the matrix calculator, and the MF and IS normalized MF are calculated automatically when the data from the three LC/MRM experiments are processed.

|
Solvent delivery: |
Waters ACQUITY UPLC Binary Solvent Manager |
|
Sample delivery: |
Waters ACQUITY UPLC Sample Manager |
|
Column: |
ACQUITY UPLC BEH C18 1.7 μm, 2.1 x 50 mm |
|
Column temp.: |
45 °C |
|
Sample temp.: |
4 °C |
|
Injection vol.: |
5 μL |
|
Flow rate: |
600 μL/min |
|
Mobile phase A: |
0.1% ammonium hydroxide in water |
|
Mobile phase B: |
Methanol |
|
Gradient 1: |
5% to 95% B in 0.70 min |
|
Gradient 2: |
5% to 95% B in 2.00 min |
|
MS system: |
Waters Xevo TQ MS |
|
Ionization mode: |
ESI positive |
|
Capillary voltage (ESI): |
1.0 kV |
|
Compound: |
Fluticasone propionate 501 > 293 |
|
Cone voltage: |
18 V |
|
Collision energy: |
20 eV |
|
Compound: |
Fluticasone propionate– D3 504 > 293 |
|
Cone voltage: |
18 V |
|
Collision energy: |
20 eV |
|
Compound: |
Phospholipids 184 > 184 |
|
Cone voltage: |
75 V |
|
Collision: |
4 eV |
|
RADAR: |
Qualitative Scan100 to 900 amu |
|
Scan speed: |
10,000 amu/s |
|
Source temp.: |
150 °C |
|
Desolvation temp.: |
450 °C |
|
Desolvation gas: |
1000 L/hr |
Plasma proteins were precipitated using acetonitrile (2:1 ratio).
Fluticasone propionate was analyzed in plasma. Acetonitrile (2:1) was used to precipitate plasma proteins. Two sets of experimental conditions were used, a short gradient of 0.70 min and a longer gradient of 2.00 min. The elution profile of fluticasone was determined for each chromatographic method. Fluticasone and its D3 internal standard were infused into the LC stream post-column. A solvent blank was injected (n=5) followed by blank-extracted plasma (n=5). TargetLynx was used to process the data and to automatically calculate the matrix factor for both sets of experimental conditions.
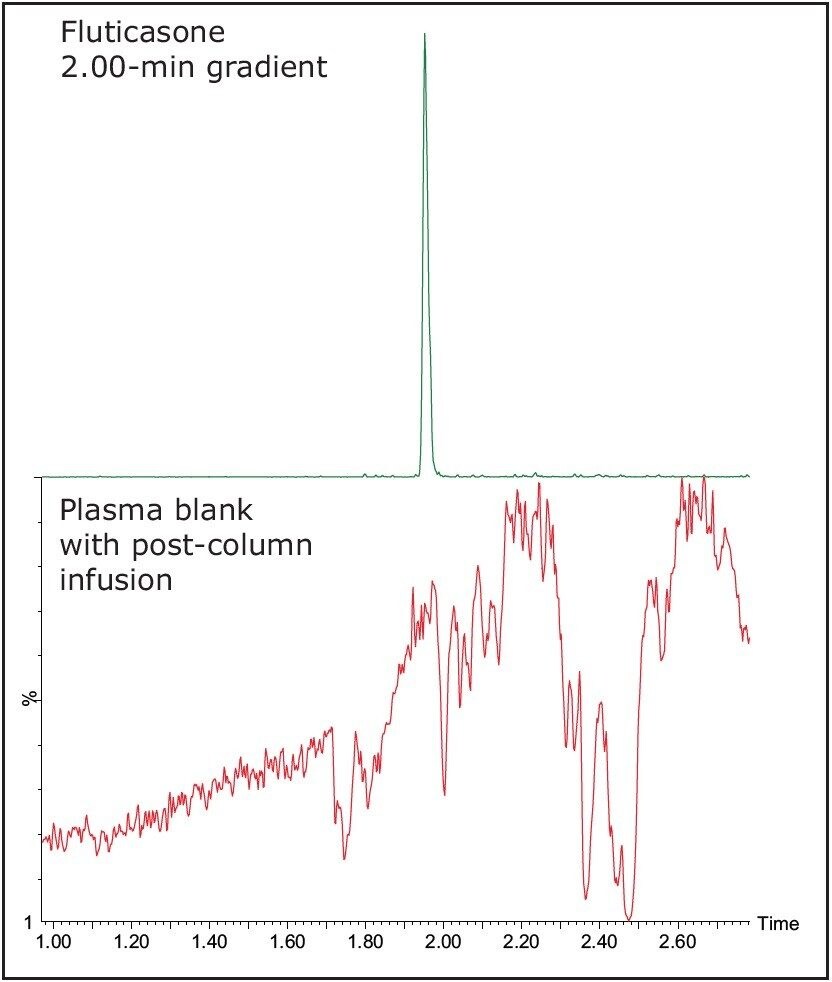
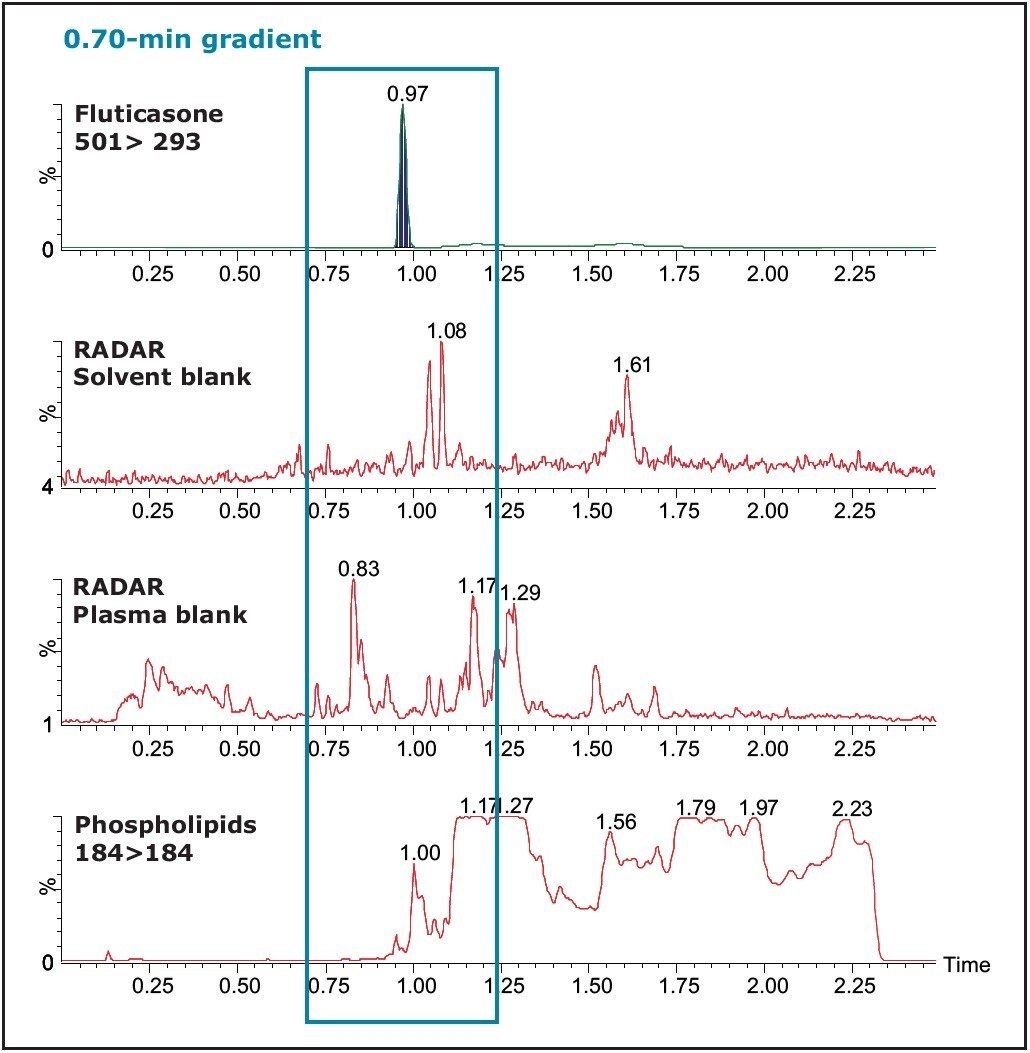
It can be seen in Figure 7 from the LC-MS/MS chromatogram that the fluticasone has a retention time of 0.97 min. The analyte chromatogram is superimposed on the plasma-blank injection where there is a post-column infusion of the analyte and the internal standard. The fluticasone elutes in a region of the chromatogram where there appears to be a suppression event occurring.
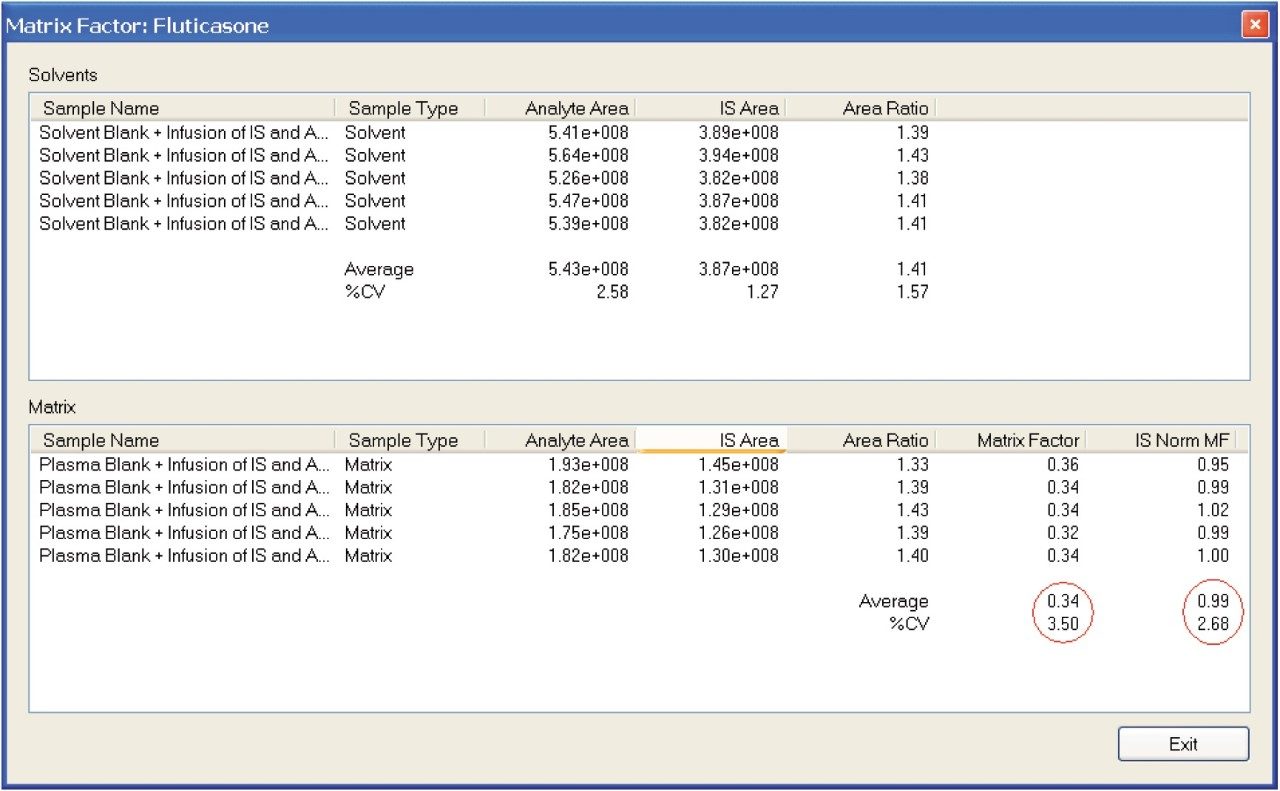
The calculated matrix factor of 0.34 shown in Figure 8 reflects this significant matrix effect. The deuterated internal standard ensures that the results are normalized to 0.99. This matrix factor is not acceptable for a bioanalytical assay. We can see from the infusion chromatogram that the peak of interest elutes at the same retention time as significant matrix material.
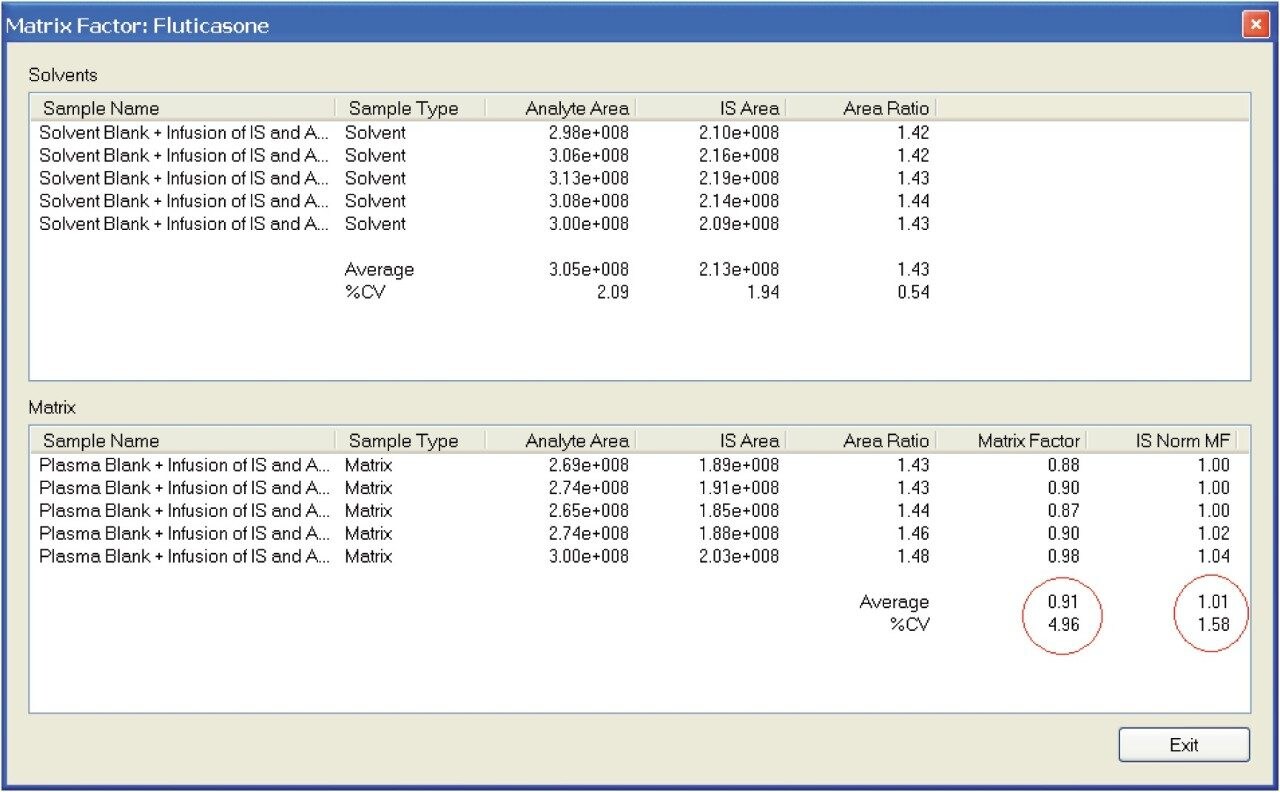
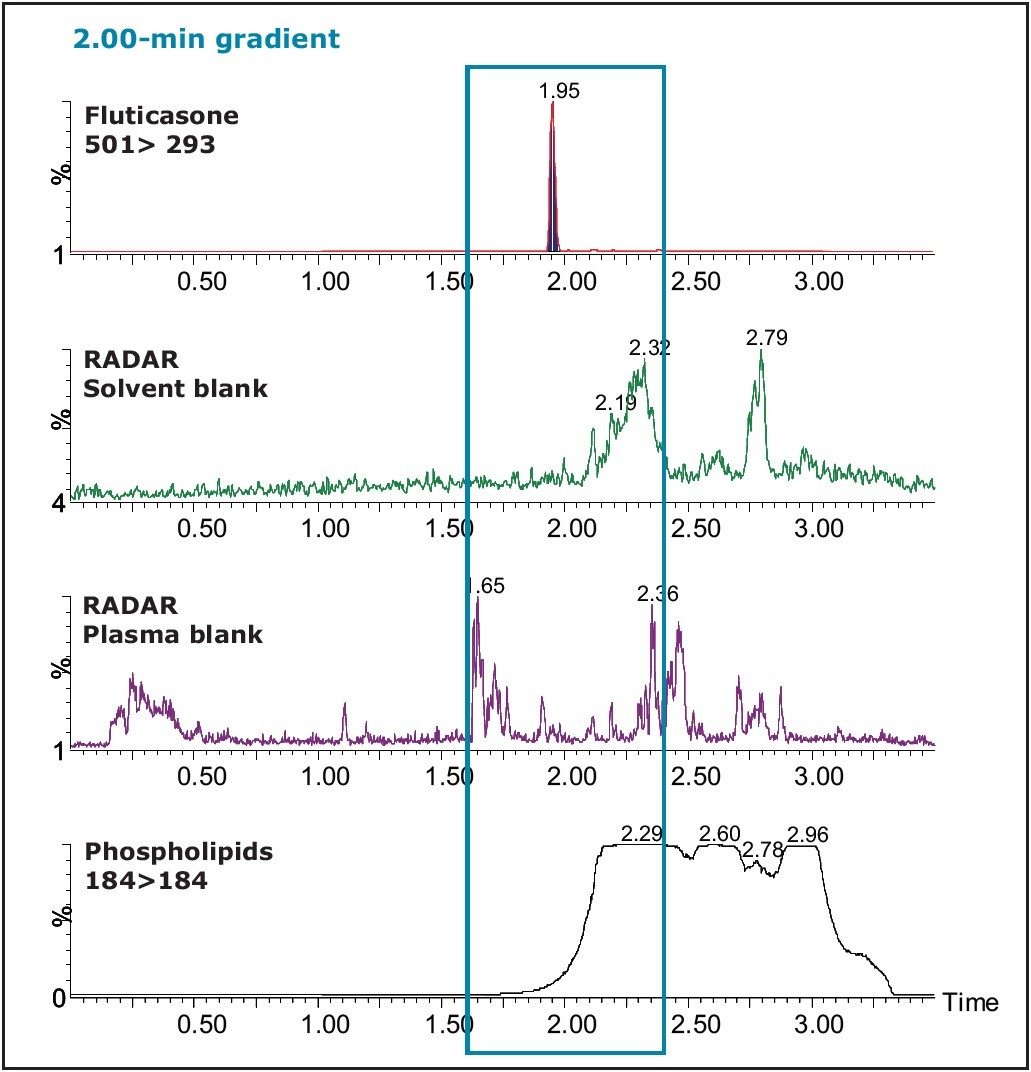
When a 2.00-min gradient was used, the retention time of the fluticasone peak was 1.95 min (Figure 9). The matrix factor report shown in Figure 10 was calculated to be 0.91, which normalizes to 1.01, due to the presence of the internal standard. It can be noted that the proximity of the analyte peak to the region of suppression has changed to a region of less coelution.

The Xevo TQ MS employs RADAR Scanning Technology to acquire both MS and MS/MS data simultaneously (formerly referred to as dual scan-MRM). This technology provides a convenient way to monitor the background while collecting MRM data for quantitation.3
Depending on the chromatographic conditions, analytes can coelute with endogenous matrix components and/ or metabolites, leading to potential matrix effects and possible reduced-assay robustness. When MRM is used exclusively, only the specified precursor > product ions are seen. Qualitative RADAR scans show all masses that are defined in the MS method scan.
In Figure 11, the proximity of the analyte peak to the phospholipids can readily be visualized. It has been reported that the concentration of the phospholipids at the retention time of the analyte can greatly influence the existence of matrix effects.
It can be seen from the RADAR qualitative scan of the plasma blank in Figure 12 that there is more chromatographic resolution between the matrix peaks and the analyte when it elutes at 1.95 min. This is supported by the data from the matrix calculator.
720003580, July 2010