
For research use only. Not for use in diagnostic procedures.
The MassTrak AAA Solution is a total system for the analysis of physiological amino acids for research use only. It provides a robust, reliable, less time consuming tool allowing higher sample throughput for physiological amino acid analysis. The analysis is complete in less than 32 minutes and allows for the identification and quantification of 42 common amino acids and related compounds in a single run.
The analysis of free amino acids in physiological fluids is an important tool in the study of physiological processes. Changes in the free amino acid concentrations can reflect altered states of metabolic pathways. These changes may be due to the influence of nutrition, environmental factors or genetic disturbances of metabolism. Expanding research in this field creates a demand for more amino acid analysis. A more robust method can offer the higher throughput that is required with this increasing sample load.
Improved throughput is a function of both reducing run time and improving method robustness. Current ion exchange methods are time consuming and frequently require lengthy method adjustment. Waters has developed a new solution for research use only. The Waters MassTrak AAA Solution combines a well-characterized amino acid derivatization, QC tested columns and eluents, a pre-configured UltraPerfomance LC method, standards, and compliance-ready software.
Following the benchtop pre-column derivatization of the analytes, separation and detection are achieved with a reversed-phase UPLC column and TUV detector, respectively. The analysis is complete in less than 32 minutes and allows for the identification and quantification of 42 common amino acids and related compounds in a single run. Full profile amino acid analyses are conducted using a defined analytical method. Electronic reports are generated using pre-defined software templates. These may be customized to suit specific reporting needs.
Sample preparation utilizes the MassTrak AAA derivatization kit and Total Recovery vials. The amino acids are derivatized with 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate (AQC) (US Patent 5,296,599 and European Patent EP 0533 200 B1).
Amino acid standard mixtures are prepared as described in the users guide.
• Acidics and Neutrals Standard Mixture
• Basics Mixture
• Glutamine
• Allo-Isoleucine
• Trytophan
• Norvaline (Internal Standard)
Derivatization followed the protocol as described in the System Guide.
• MassTrak AAA Reagent
• MassTrak AAA Borate Buffer
• MassTrak AAA Reagent Diluent
|
LC System: |
Waters ACQUITY UPLC System with TUV Detector |
|
Column: |
MassTrak AAA Column, 2.1 x 150 mm, 1.7 μm |
|
Column Temp: |
43 ˚C |
|
Flow Rate: |
400 μL/min. |
|
Mobile Phase A: |
MassTrak AAA Eluent A Concentrate, diluted 1:10 |
|
Mobile Phase B: |
MassTrak AAA Eluent B |
|
Weak Needle Wash: |
5/95 Acetonitrile/Water |
|
Strong Needle Wash: |
95/5 Acetonitrile/Water |
|
Gradient: |
MassTrak AAA Standard Gradient (provided in solution) |
|
Detection: |
UV @ 260 nm |
|
Injection Volume: |
1 μL |
|
Injection Mode: |
Partial Loop with Needle Overfill (PLNO) |
The sample preparation for the biological samples is dependent on the matrix. Plasma samples require deproteinization prior to analysis. Typical example:
A reliable amino acid analysis method must meet specific requirements. The accurate identification of the individual amino acids is essential. Additionally, the amino acids need to be accurately quantified. This quantification must be linear over the range commonly observed in physiological samples.
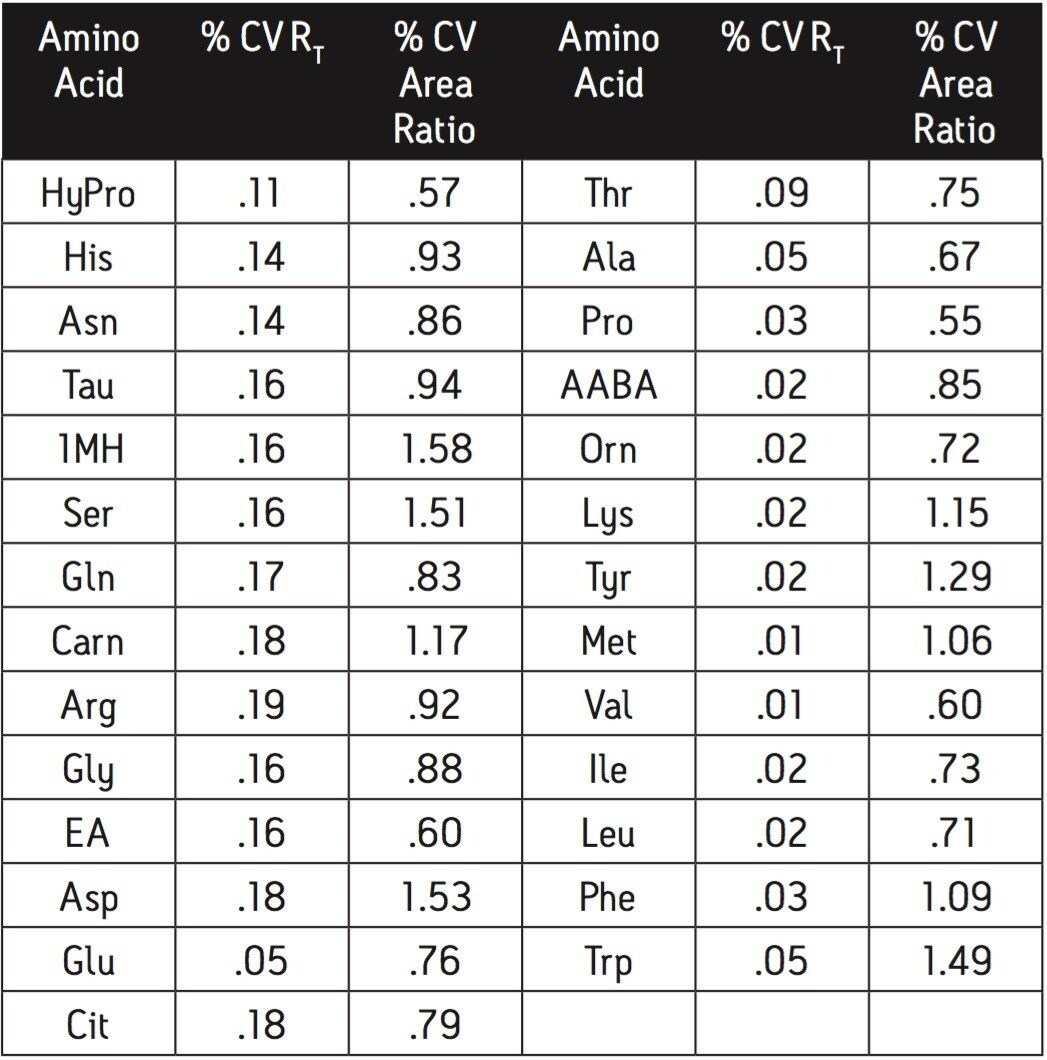
The reliability and robustness of the analysis was measured to ensure accurate identification and quantification of all the amino acids included in the assay. The amino acids are identified by retention time relative to the supplied standard. Reliability of the identification of each amino acid is based on retention time and its the coefficient of variation expressed as a percentage (% CV) (Table 1).
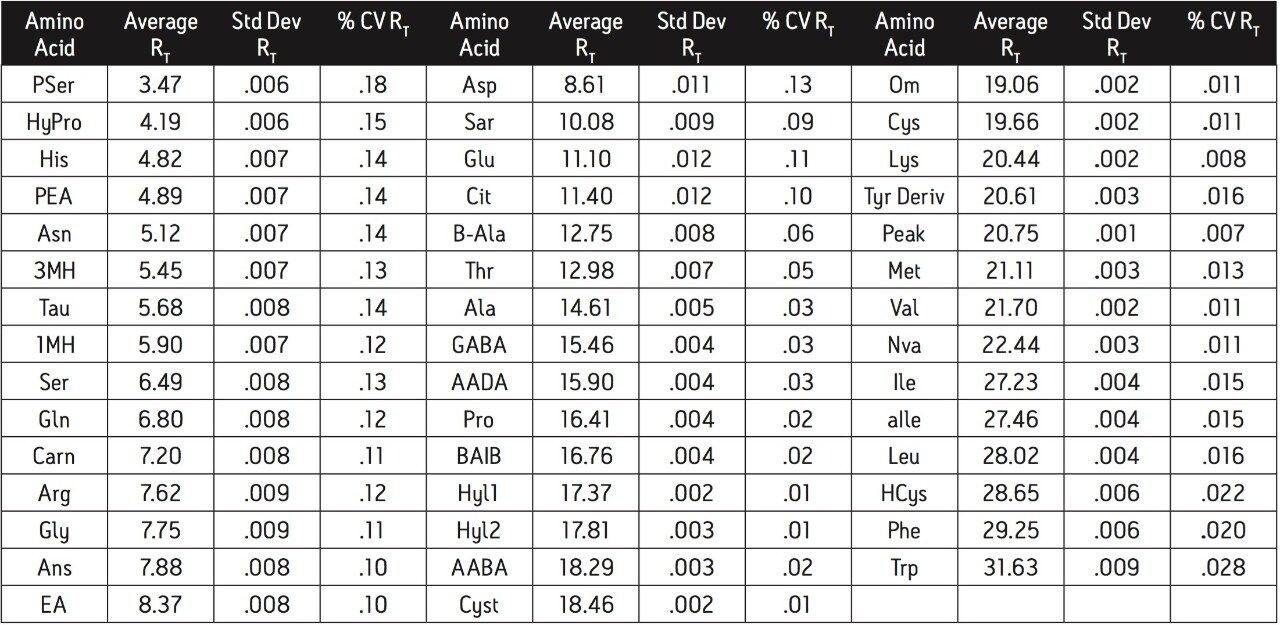
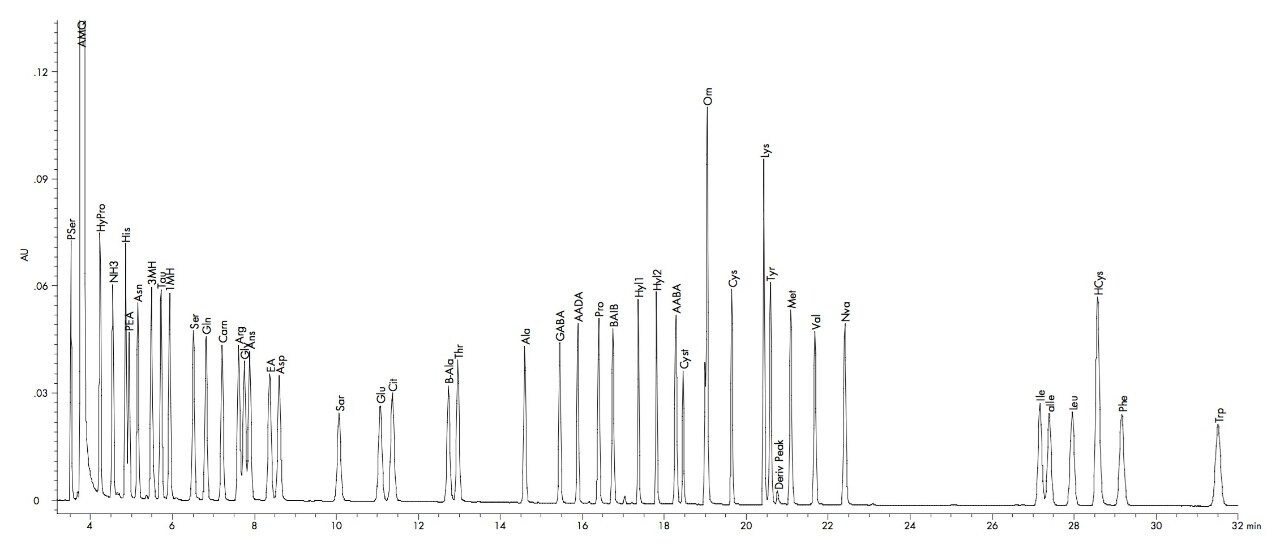
Retention time precision for a standard is typically less than 0.2% CV over a series of runs. This inter-run variability of retention time is much less than the retention difference between adjacent peaks, thereby ensuring accurate and reliable identification. For example, His and PEA are the most closely spaced peaks. The retention time standard deviation for both His and PEA is .001. The difference in retention time between these peaks is .007, approximately three times the sum of the standard deviations. This low variability in retention time allows for increased confidence in the identification of each amino acid. To further ensure that each peak is correctly identified the retention time is referenced to a well resolved peak in each sample.
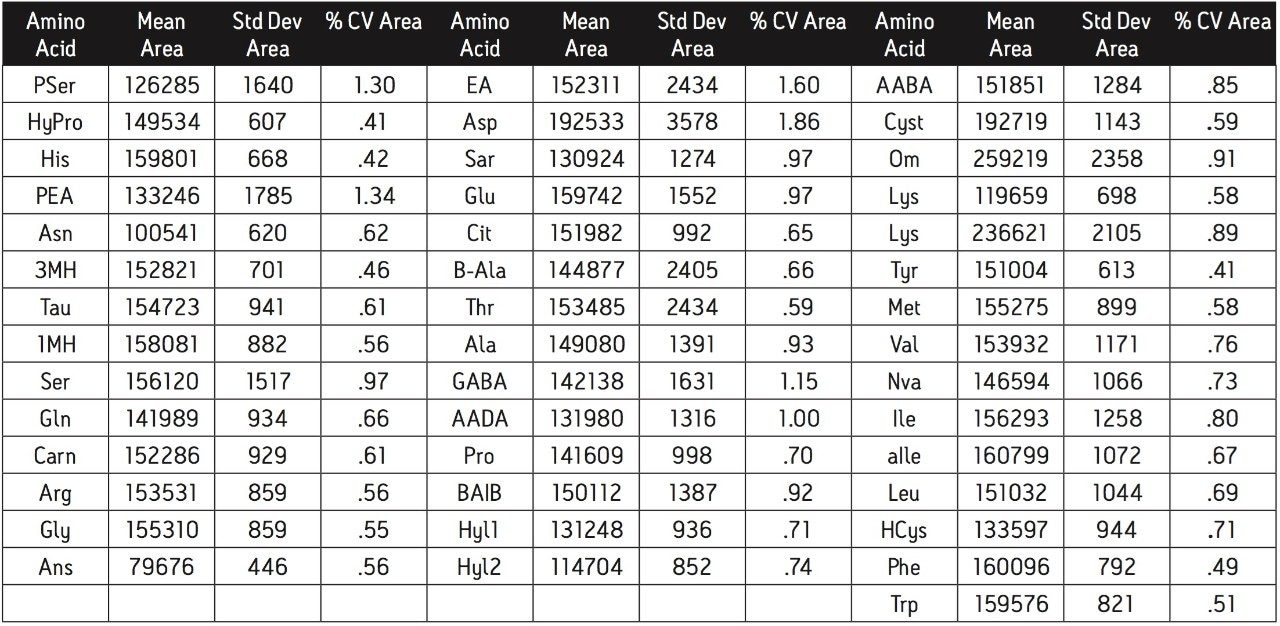
Reliable quantification of the amino acids is dependent upon reproducibility, sensitivity and linearity. With the MassTrak AAA Solution, the inter-run precision, as measured by the CV of peak area counts, is within 2 % for all the amino acids (Table 2). The variability in quantification is comparable, if not superior, to existing methods. The quantification can be more precise when an internal standard is incorporated.
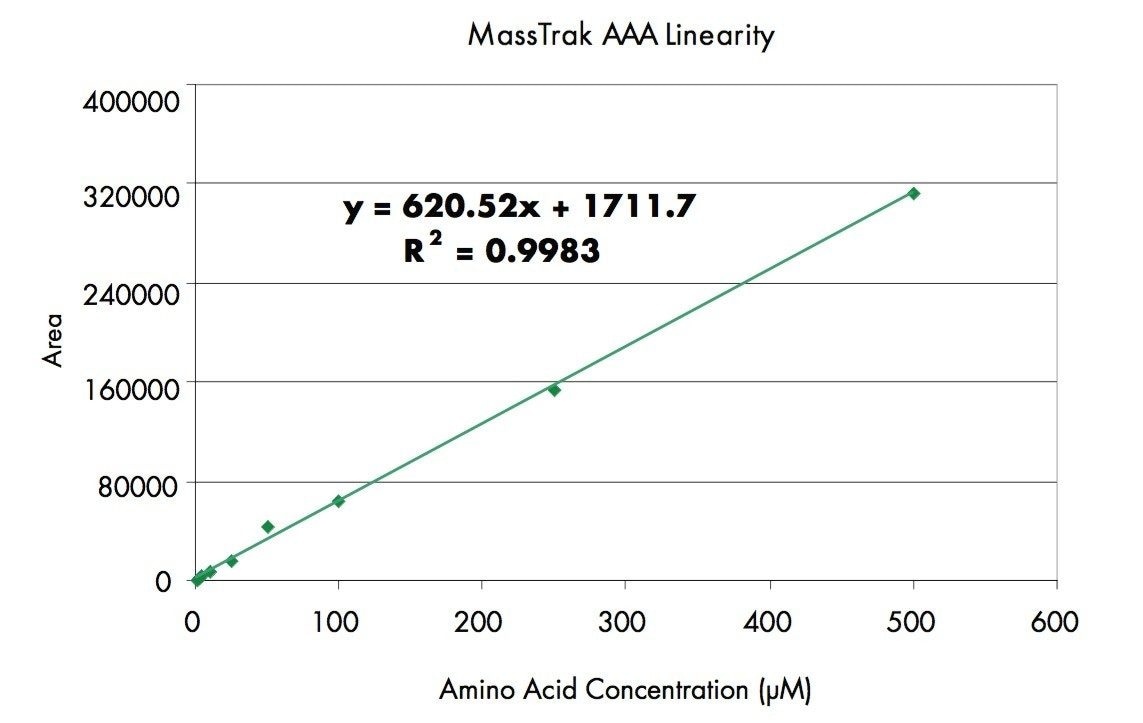
Another important feature of any amino acid analysis is linearity of response. The MassTrak AAA solution exhibits a the linear response over the range 1-2μM to 500 μM with the complete amino acid standard. Each of the amino acids in the mixture has a linear response from 1-2 μM to 500 μM with a R2 ≥ 0.995 (e.g. Figure 2). For points at the low end of the calibration curve or less than 5μM, the deviation from calculated values is <20 %. Points at higher levels are within 10 % from the calculated concentrations. The % deviation for both the low and high end of the calibration curve is within generally accepted limits.
The Lower Limits of Quantification (LLOQ) and the Limits of Detection (LOD) are dependent upon the response seen for each amino acid and the related background contamination. The LLOQ is defined as the lowest level that can be reproducibly and accurately quantified within 10 % of a known value. With the MassTrak AAA solution the LLOQ is 1-2 μM for amino acids (Figure 3). The limit of detection (LOD) is defined as a signal-to-noise of 4 or greater. All of the amino acids have a LOD less than 0.5 μM.
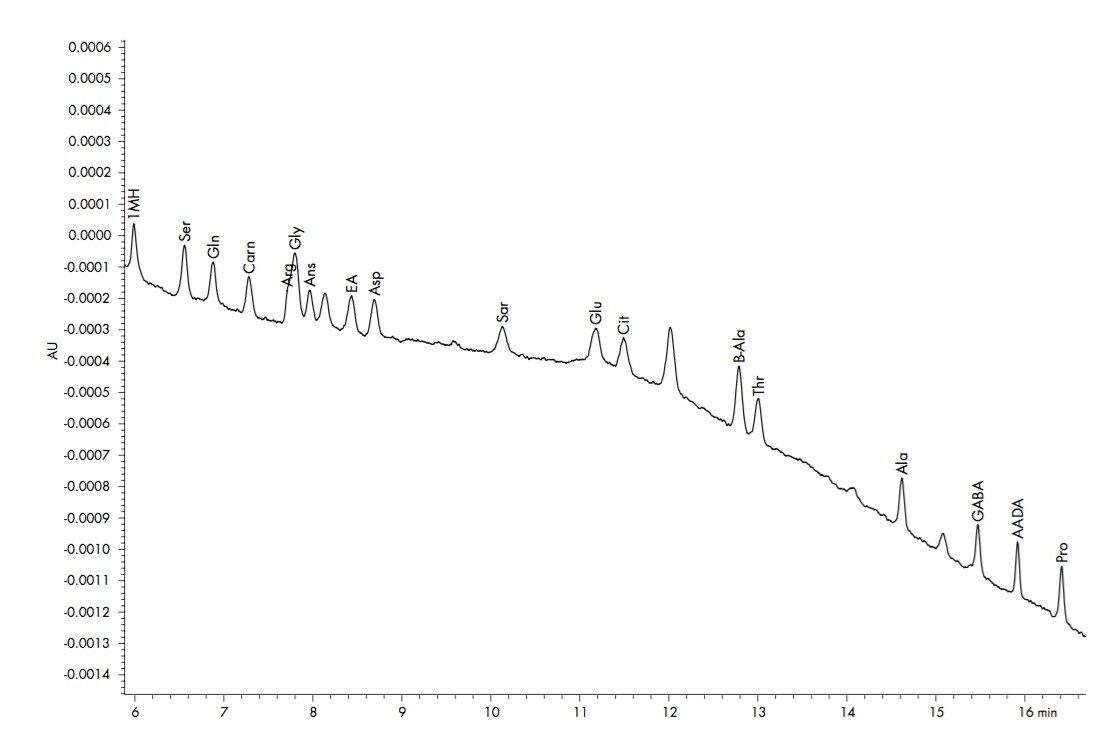
The constraint in the upper limit of quantification (ULOQ) is the amount of excess reagent that is required for the complete derivatization of all of the amino acids in the sample. The derivatization reagent excess should be greater than four times the total amount of amino acids in a sample. Since a known amount of reagent is added to each sample, complete derivatization requires the total amino acid amount in a raw sample not exceed 21 mM. If the sum of all the amino acids exceeds this amount, the more slowly reacting amino acids are not completely derivatized. However, biological samples rarely exceed 21 mM total amino acid content. Single amino acids have been shown to provide linear response up to 10 mM.
The MassTrak AAA Solution is designed for the analysis of physiological amino acids in a variety of matrices, including deproteinized plasma (Figure 4). Replicate analyses of a plasma sample demonstrate the retention time precision within run 0.2% CV inter-run (Table 3). The area precision for the deproteinized samples is typically within 2% CV inter-run. The low level of variability in a complex matrix demonstrates the robustness of the method even in the presence of possible interferences and provides added confidence, accurate identification and quantification.
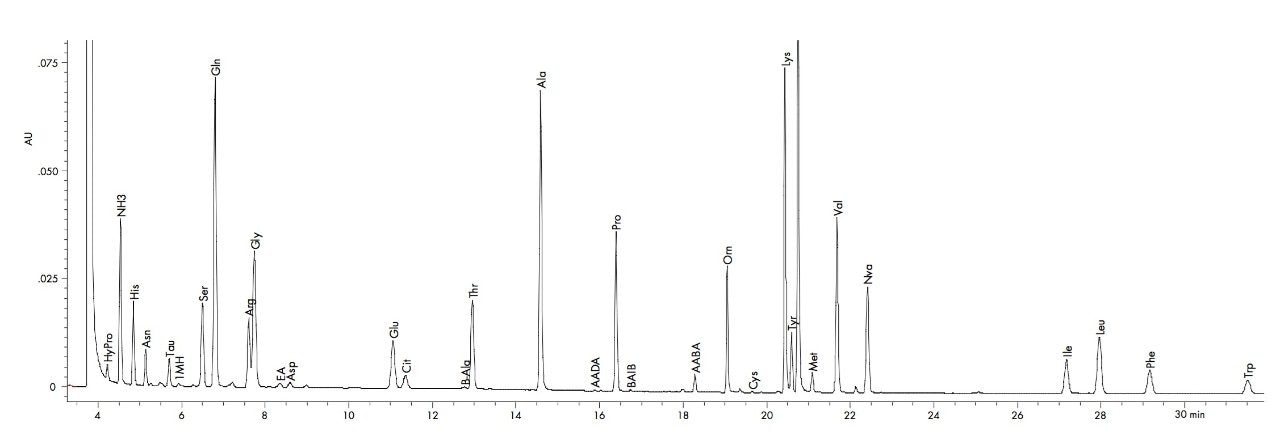
The MassTrak AAA Solution is a total system for the analysis of physiological amino acids for research use only. This solution includes columns, eluents, standards, derivatization reagents, instrumentation, software and support, all of the necessary components to provide continuous and reliable operation.
The MassTrak AAA Solution is based on a well-studied derivatization chemistry. The quantitative analysis is therefore robust and easy to troubleshoot should any problems should arise. The derivatives are separated using UltraPerfomance LC using an ACQUITY UPLC system with TUV detection. The separation chemistry uses columns and eluents that are tested during manufacture to assure consistent and successful analyses. The software includes system suitability tests that monitor the performance while running the series of samples. It also incorporates custom reports that are adapted to the analysts requirements.
This solution provides accurate and precise qualitative and quantitative amino acid analysis. Quantitative precision is better than 2 % CV and the demonstrated linearity exceeds the levels that are commonly observed in physiological samples. The LLOQ of 1 μM meets the generally desired quantifiable levels in physiological samples. In addition, the LOD of 0.5 μM provides greater sensitivity than comparable methods.
The MassTrak AAA Solution provides a robust, reliable, less time consuming tool allowing higher sample throughput for physiological amino acid analysis.
720002903, April 2009