Amino acid analysis is used in the protein structure laboratory to provide two kinds of information. First, the total quantity of amino acids is a direct measure of the amount of protein in a sample. Second, the measurement of the proportions of amino acids provides information to confirm the identity of the protein and to detect modifications. Both applications require robust, accurate, and sensitive measurements that both identify and quantitate the amino acids. There is increasing need for these labs to provide these correct results faster and more economically.
Amino acid analysis is used in the protein structure laboratory to provide two kinds of information. First, the total quantity of amino acids is a direct measure of the amount of protein in a sample. Second, the measurement of the proportions of amino acids provides information to confirm the identity of the protein and to detect modifications. Both applications require robust, accurate, and sensitive measurements that both identify and quantitate the amino acids. There is increasing need for these labs to provide these correct results faster and more economically.
The Waters UPLC Amino Acid Analysis Application is a turnkey solution to address these needs. This total system solution includes a well established and understood sample derivatiza-tion kit, eluents, chromatographic column, and a separation system based on the ACQUITY UPLC using UV detection under Empower software control.
In this experiment, this system solution is used to measure the composition and concentration of a known protein. The accuracy of the determination is compared to the known correct results.
Acid-hydrolyzed bovine serum albumin (BSA) samples were prepared in an independent laboratory as part of a collaborative study. The samples were supplied at an estimated concentration of 1.0 mg/mL in 0.1 M HCl sealed under argon in sealed ampoules. Samples were stored at -80 °C until analysis.
The sample was diluted 1:10 with 0.1 M HCl prior to derivatiza-tion. The standard derivatization protocol was modified to include neutralization of excess acid with 0.1 M NaOH. Conditions for derivatization and analysis are described in detail in the Waters UPLC Amino Acid Analysis Application System Guide (P/N 71500129702).
|
LC system: |
ACQUITY UPLC System with TUV detection at 260 |
|
Column: |
AccQ•Tag Ultra 2.1 x 100 mm, 1.7 μm |
|
Column Temp: |
55 °C |
|
Flow Rate: |
700 μL/min |
|
Mobile Phase A: |
1:20 Dilution of AccQ •Tag Ultra Eluent A concentrate Part Number: 186003838 |
|
Mobile Phase B: |
AccQ•Tag Ultra Eluent B |
|
Gradient: |
AccQ•Tag Ultra Hydrolysate Method |
|
Injection Volume: |
1 μL |
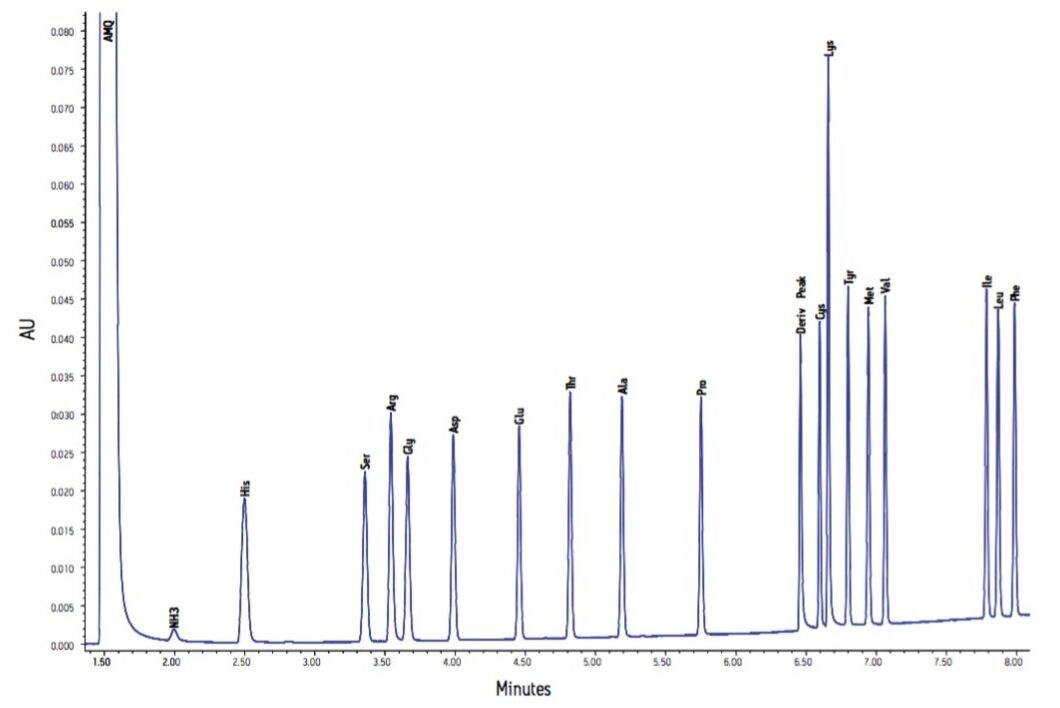
Figure 2 shows the chromatogram of a standard that contains amino acids typically found in protein hydrolysate samples. Each amino acid is 10 pmoles on column. For quantitative analyses, a three point calibration at 0.5, 10, and 25 pmoles on column was applied.
The analysis of a typical BSA hydrolysate is shown in Figure 3. The estimated starting concentration is consistent with this chro-matogram representing a total of 9 ng of protein on column. This analysis was repeated a total of 75 times, over five separate days, with two columns and a total of five mobile phase preparations. The 75 injections represent five independent sample dilutions, each dilution derivatized five separate times. Each derivatized sample was injected in triplicate.
The amino acid composition, expressed as residues/mole of protein, was compared to the value expected from the known sequence. Table 1 shows the mean and standard deviation for each amino acid over all 75 analytical injections. Tryptophan and cysteine/cystine are excluded from the calculation because they are destroyed by acid hydrolysis. The measured molar composi-tion agrees well with the expected values from the sequence. The reliability of the UPLC Amino Acid Analysis Application is confirmed by the reproducibility of results over the large number of determinations that intentionally includes the variability that could arise from multiple columns, eluents, and derivatizations.
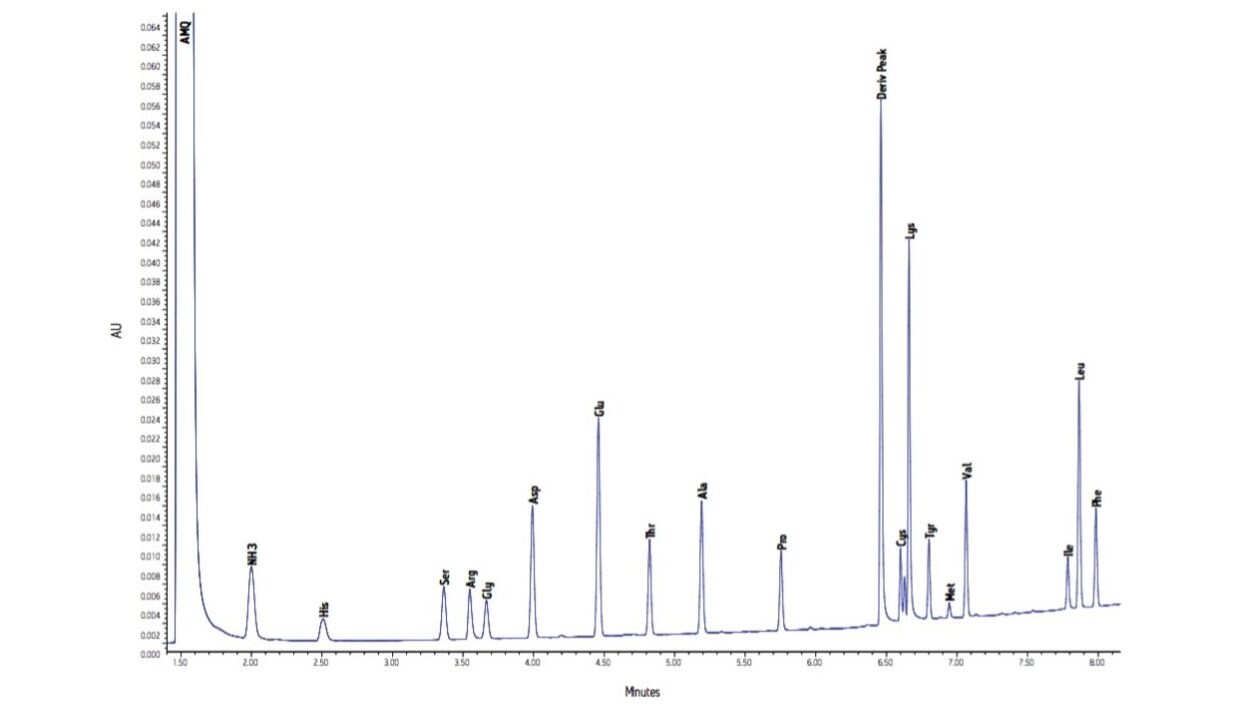
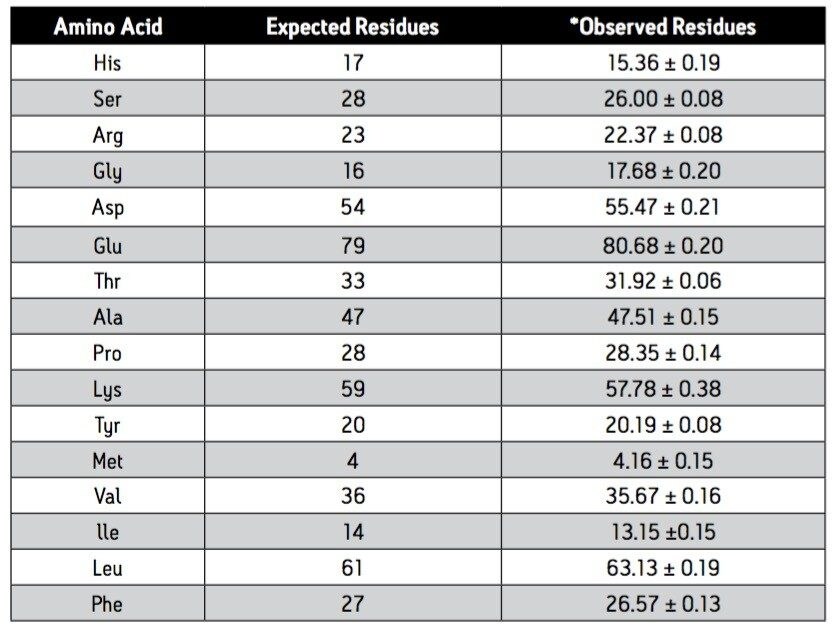
Table 1. Comparison of Observed with Expected Composition Derived from Known Sequence of BSA
*Average of 75 data points (25 derivatizations, each injected in triplicate; mean value ± standard deviation)
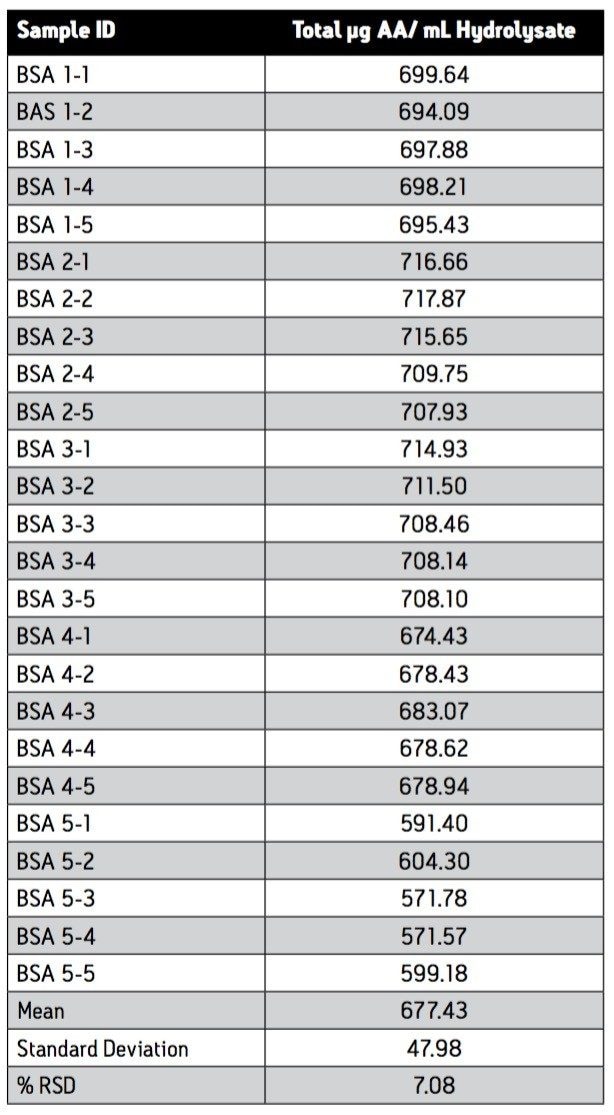
The analytical data was used to calculate absolute amount of protein in the sample. The amount of each amino acid were expressed as the residue molecular weight. The sum of weights of the amino acids is equal to the weight of the protein. Table 2 sum-marizes the result of the 75 determinations with the mean for the triplicate injections shown for each derivatization. The measured amount corresponds to 0.7 mg/mL in the starting material. The estimated amount used to prepare the hydrolysate has not been independently verified. It should be noted in addition that this measurement of protein amount does not include the contribution of cysteine/cystine and tryptophan, as they are mostly destroyed by the hydrolysis of the protein.
The reproducibility of determination of the 75 analyses gives a RSD of 7%. Detailed examination shows that much of this vari-ance is due to the difference of experiment 5 from the other four analyses. Since all of the replicates in experiment 5 are lower than the others, this variance is consistent with a difference in pipetting in the initial sample preparation. The addition of an internal standard to the sample to be hydrolyzed will improve the reliability of the final analytical result. Norvaline is the preferred internal standard for this purpose
Protein structure and biopharmaceutical laboratories rely on accurate quantitation of amino acids to confirm the identity and amount of protein in their samples. The analyses shown here demonstrate that Waters UPLC Amino Acid Analysis Application can provide assured results for these laboratories.
The molar ratios of amino acids are reproducible over multiple derivatizations and replicate injections. The measured composi-tion agrees with that expected from the sequence of BSA.
Determination of the absolute amount of protein in the samples is determined by summing the residue weights of amino acids. The reproducibility of this measurement is on the order of ±7%. The largest contribution to this variance is the initial sample dilution. Incorporation of an internal standard in hydrolysis will improve the precision of the determination. Norvaline is recommended as an internal standard for the method.
The Waters UPLC Amino Acid Analysis Application provides a complete turnkey analytical method for analyzing protein hydrolysate samples. The ACQUITY UPLC system gives very high resolution for certain peak identification and ease of integration. With standard UV detection, all the derivatized amino acids have similar extinction coefficients to facilitate quantitative analysis. Sensitivity levels corresponding to nanograms of protein can be achieved routinely. The ruggedness of this turnkey system solu-tion ensures rapid and unequivocal identification of proteins, with no interference or ambiguity. The pre-tested column, eluents and reagents ensure that the user will not spend time adjusting the method. The small amount of sample required for good analyses contributes to long column life and minimizes the chance of failure during a series of runs. The high resolution ensures reliable peak identification and quantitation so that runs need not be repeated. The pre-defined methods and reports simplify reporting of results. These analytical benefits are obtained with a short analysis time for the high throughput required for the routine determination of protein composition and concentration.
720002404, October 2007