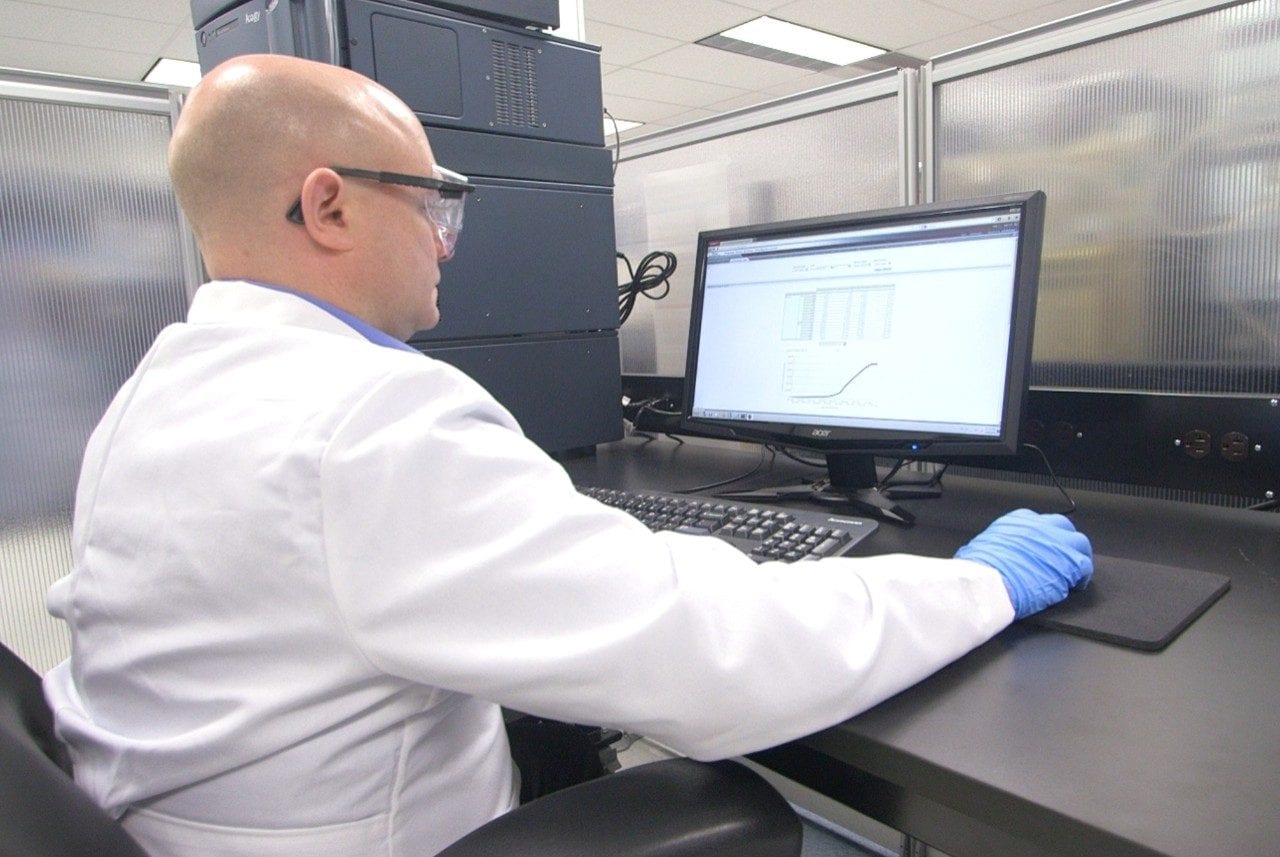

This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates to how the export functionality within the Empower CDS can be leveraged in the interpretation of complex mass spectra data.
Empower 3 supports the ability to directly export MS spectral data for increased flexibility in data interpretation.
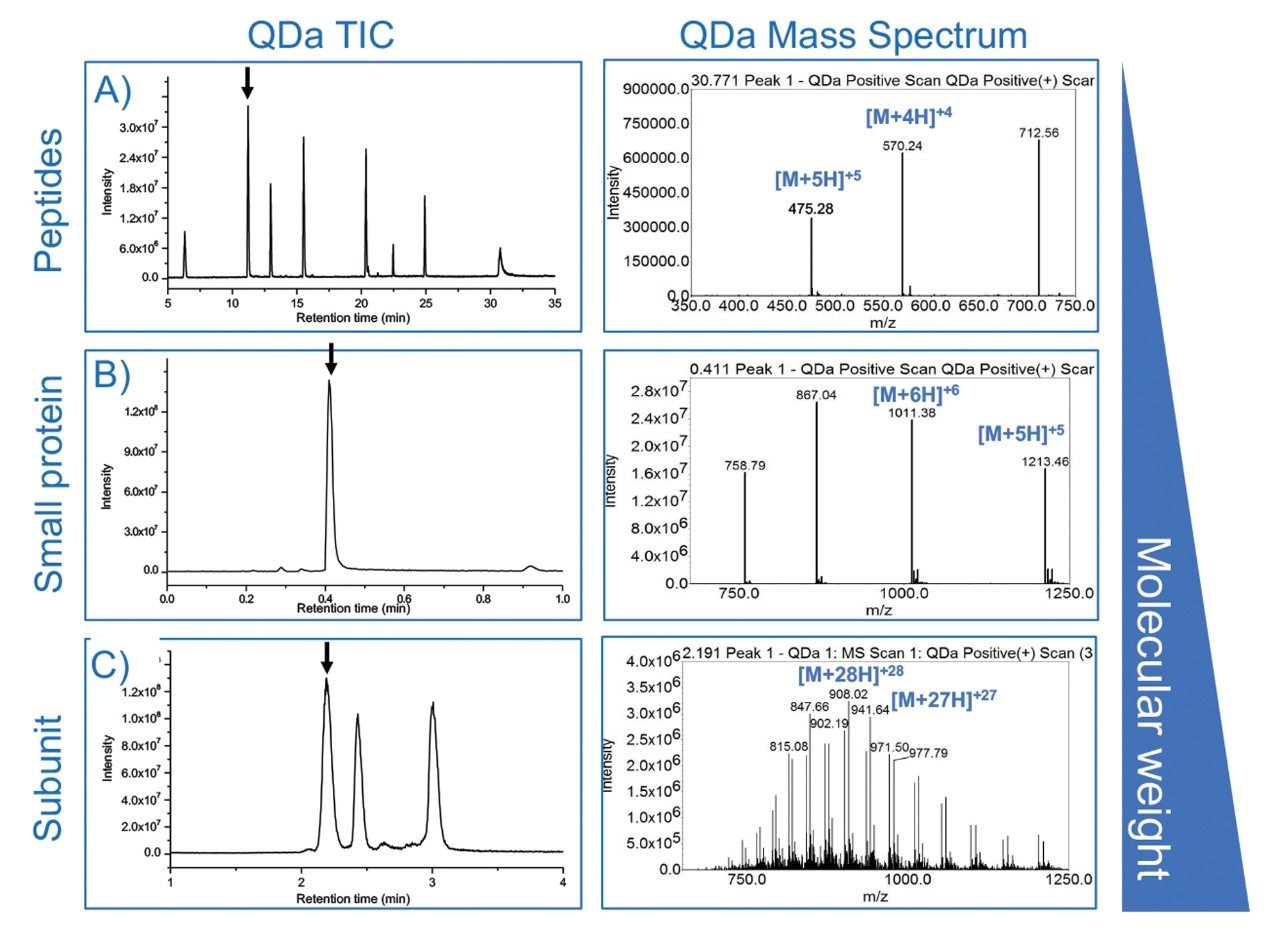
Interpretation of mass spectral data acquired within the Empower CDS can be straightforward when analyzing small analytes that produce relatively “simple” mass spectrum with a limited number of charge states as shown in Figures 1A and 1B. Albeit a manual process, nominal mass of the analyte can be determined from the Empower MS spectra using the formula for mass per charge (m/z), or m/z = (M + zA)/z, which can be applied to the individual m/z charge states in the spectrum. For routine monitoring or samples that consistently produce the same charge states, custom fields can be programmed to automate this calculation to determine nominal mass. While this approach works for simple or moderately simple molecules, more complex samples such as the one shown Figure 1C would benefit from deconvolution algorithms to efficiently process MS spectra. Conveniently, Empower 3 supports the ability to directly export spectral data for increased flexibility in the interpretation of MS data for improved productivity in the lab.
Empower 3 allows for the direct export of MS spectral data via the integrated spectrum review window within Empower for increased flexibility in data interpretation. As shown in Figure 2, once mass spectra data is acquired, spectral information such as individual m/z and associated intensities can be accessed via the “Spectrum Points” tab. Using the Window’s “copy” keyboard shortcut, “Ctrl+C,” the spectral data can be exported and saved as text or in a spreadsheet for further processing and interpretation.
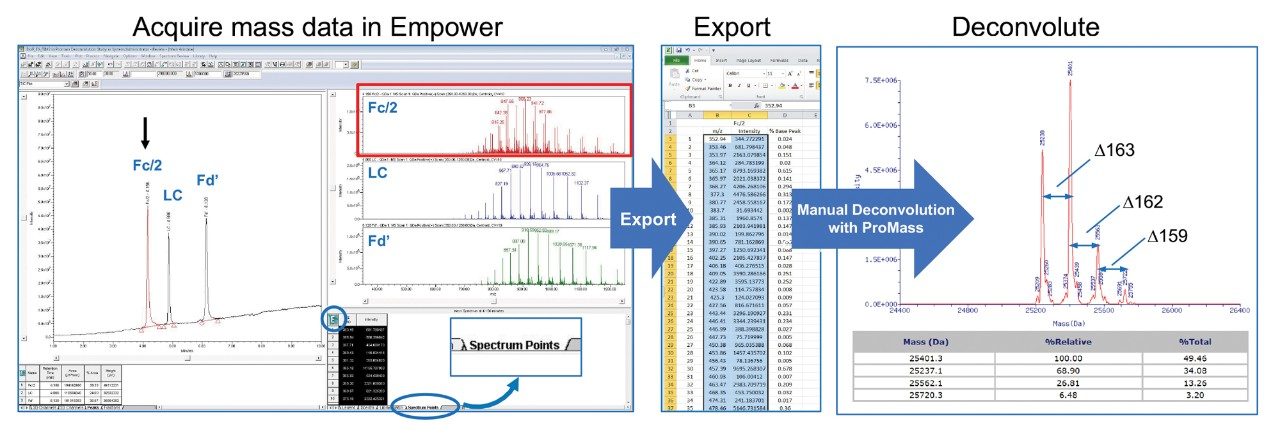
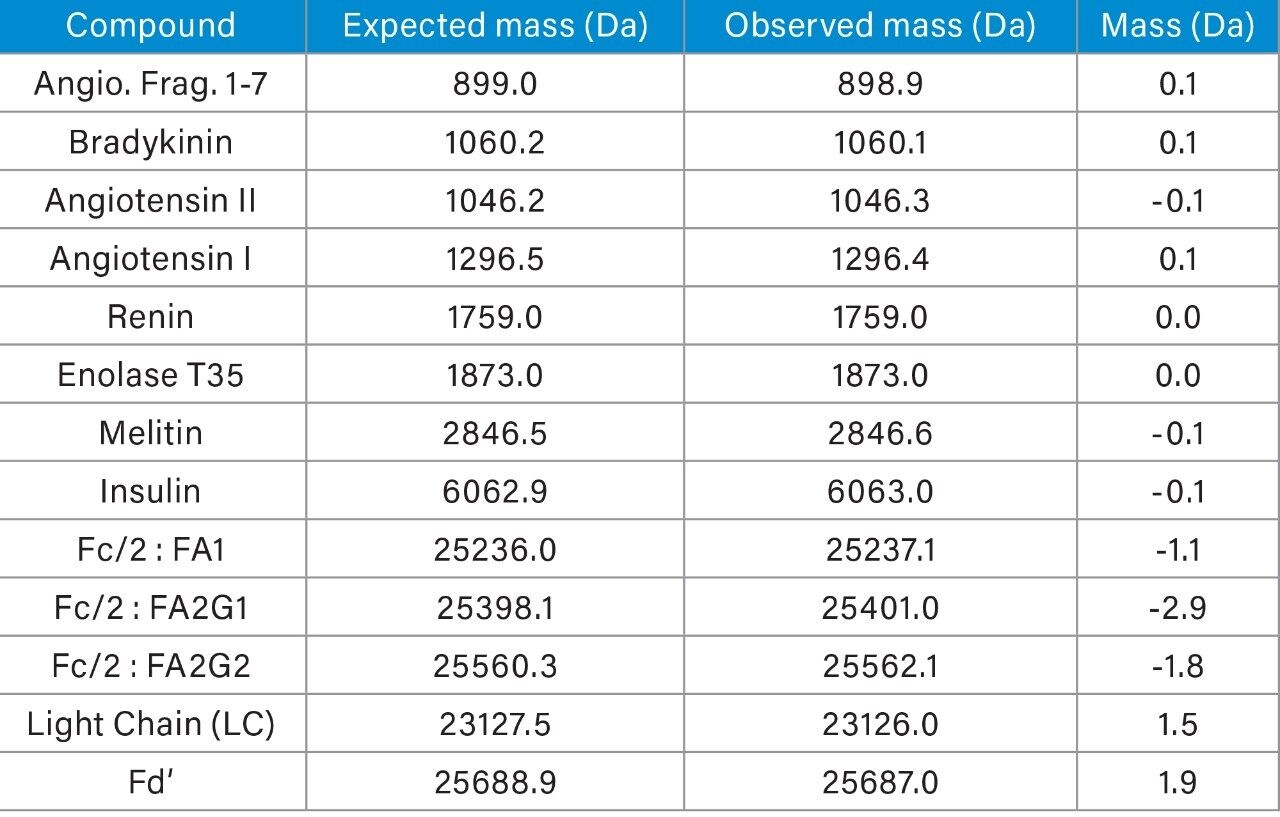
As shown in the far-right panel of Figure 2, MS data acquired using the ACQUITY QDa was deconvoluted using the ProMass software by Novatia to determine the relative abundance and nominal mass of the main glycoforms of the Fc/2 form of a reduced mAb subunit sample. In addition to being able to process complex spectra, ProMass is capable of deconvoluting relatively simple spectra as well as the ones generated from peptides or small proteins such as insulin. Given this, the Waters MassPREP Peptide Standard, insulin glargine, and IDES digested NIST mAb (Figure 1) were chosen to assess the applicability and accuracy of the method over a wide range of molecular weights with MS spectra of varying complexity. As shown in Table 1, the deconvoluted mass data produced by manually exporting spectra to ProMass was within instrument specification of +/- 0.2Da × z, with the larger NIST mAb subunits exhibiting higher mass difference due to ionizing at higher charge states (z).
Empower 3 allows for the direct export of MS spectral data via the integrated spectrum review window within Empower for increased flexibility in data interpretation. This functionality is useful in LC-MS workflows, particularly for applications involving critical peaks with complex mass spectra, where information about the presence and relative quantity of a peak’s glycoforms is not discernable from mass spectra alone.
720006606, July 2019