In this application note, we demonstrate the use of REIMS with chemometric modeling performed in real time with LiveID Software to accurately determine the species level identification of five commercially popular, visually and genetically similar white sea fish species.
REIMS with LiveID Technology requires no sample preparation and provides accurate and nearly instantaneous results. The reduced amount of time required for analysis and data interpretation that REIMS offers in contrast to the current PCR-based methods represents a significant improvement in operational efficiency. REIMS with LiveID has been demonstrated as a complementary technique for the detection of commercial fish fraud.
Economically motivated adulteration (EMA) of seafood products is a global issue occurring at alarmingly high rates, and it is estimated that on average 30% of commercial fish products sold are either misrepresented or mislabeled.1 This equates to fraud of almost $120B within the global seafood industry, as the Food and Agriculture Organization of the United Nations (FAO) estimate the global seafood industry to be worth $400B annually, with global industry analysts expecting this value to rise to $430B by 2018.2
Genomics, proteomics, metabolomics and lipidomics are four alternative and in some cases complimentary systems biology approaches often employed for food fraud detection studies.3 The majority of fish fraud detection studies utilize genomic profiling as DNA is found in all cells and organisms and can be analyzed in all types of tissue ranging from freshly caught fish to processed and cooked samples.4 While very accurate qualitative and quantitative results are achievable using polymerase chain reaction (PCR), it comes at the expense of long and often complex sample preparation coupled with long assay running times which sometimes extend to more than a working day. In terms of managing fraud in fast moving supply chains, this is a substantial disadvantage.
Rapid Evaporative Ionization Mass Spectrometry (REIMS) is a form of ambient ionization mass spectrometry that, as is the case with many analytical innovations, was created for medical research purposes. It operates using an electrosurgical knife or bipolar forceps which create an aerosol (smoke) when cutting into a tissue sample. The aerosol is evacuated from the sample through a transfer line into the ionization source of a mass spectrometer where a heated collision surface is situated and the ionization process occurs. Although the majority of publications utilizing REIMS have centered on medical and bacterial identification applications,5,6 there are early indications that it may also find applications in the detection of food fraud.7 Results are obtained nearly instantaneously (2–3 seconds) and the technique can achieve results for solid samples without the need for any form of sample preparation.
In this application note, we demonstrate the use of REIMS with chemometric modeling performed in real time with LiveID Software to accurately determine the species level identification of five commercially popular, visually and genetically similar white sea fish species: Gadus morhua (cod), Pollachius virens (coley), Melanogrammus aeglefinus (haddock), Pollachius pollachius (pollock), and Merlangius merlangus (whiting). Unlike most other analytical systems currently employed for species level identification in food, Waters REIMS Research System with iknife sampling device and LiveID has the capability to determine results in real time. This combination of mass spectrometric data and chemometric modeling is extremely beneficial to the food industry for the rapid identification of fish fraud including species level identification, capture method, geographical origin, and the potential for point-of-control testing.
|
Sampling device: |
iKnife (monopolar electrosurgical knife) |
|
Diathermy generator: |
Erbe VIO 50 C |
|
Diathermy mode: |
Autocut |
|
Power setting: |
30 W |
The REIMS source was connected to a monopolar electrosurgical knife (Model PS01-63H, Hangzhou Medstar Technology Co, Ltd, Jiaxing City, China) through a 3 m long, 1 cm diameter ultra-flexible tubing (evacuation/vent line).
|
MS system: |
Xevo G2-XS QTof, sensitivity mode |
|
Source: |
REIMS |
|
Acquisition mode: |
TOF MS |
|
Ionization mode: |
ESI- |
|
Mass range: |
200 to 1200 m/z continuum |
|
Scan speed: |
0.5 s/scan |
|
Cone voltage: |
30 V |
|
Heater bias: |
40 V |
Prior to analysis, the Xevo G2-XS QTof Mass Spectrometer was calibrated using a 5 mM sodium formate solution (in 90% IPA) at a flow rate of 0.2 mL/min for 2 min. A lock mass solution of Leucine Enkephalin (Leu Enk) (m/z 554.2615) (2 ng/μL) in isopropanol (IPA) was infused using at a continuous flow rate of 0.1 mL/min to be used as a lock mass for accurate mass correction.
The model was trained using five commercially popular white fish species. All tissue samples (fillets, tails, and unspecified areas) of cod, coley, haddock, pollock, and whiting were sourced from trusted suppliers and stored at -80 °C. Prior to REIMS analysis, the samples were thawed at room temperature for 2 hours in the fume hood where the REIMS sampling took place.
Electrosurgical dissection in all experiments was performed using an Erbe VIO 50C generator (Erbe Medical UK Ltd, Leeds, UK). The generator was operated in Autocut mode with a power setting of 30 W. All samples were cut on the return electrode plate and a venturi gas jet pump driven by nitrogen (1 bar) evacuated the aerosol produced at the sample site towards a heated kanthal coil that was operated at 6.4 W (2.8 A at 2.3 V).
Depending on the size, each tissue sample was sampled between 8 and 12 times for repeatability with each cut lasting approximately 3 to 5 s. This enabled multiple locations on each tissue sample to be analyzed. The delay between sampling and appearance of a signal was approximately 2 s, with no carryover effects visible between each burn and/or sample.
Multivariate statistical software package LiveID (v.1.1) was used as a model builder and recognition tool. To generate models from the untargeted profiling REIMS ToF MS data acquired in MassLynx MS Software (v.4.1) the following data pre-treatment steps were performed: lock mass correction applied using the Leu Enk ion at m/z 554.2615; all spectra contained within each “burn event” termed the region of interest (ROI) were combined to form a single continuum spectrum; Adaptive Background Subtraction (ABS) algorithm was applied to reduce the chemical background in the combined spectra; data resampling (binning to 0.5 Da) was performed to reduce the data dimensionality; the resulting spectrum was normalized using the Total Ion Chromatogram (TIC). All chemometric models were calculated using the mass region of 600–950 m/z. The peak detection threshold was automatically set within LiveID from file to file based on the minimum spectral intensity value plus 10% of the difference between the maximum and minimum intensities.
T = IntensityMin + 0.1*(Intensity Max – Intensity Min)
Following data pre-treatment steps, a Principal Component Analysis (PCA)/Linear Discriminate Analysis (LDA) model was generated. First, an unsupervised PCA (Singular Value Decomposition algorithm) transform was applied to the spectral data calculating the scores and loadings plots; a supervised LDA transform was then applied to the scores calculated by the PCA transform. LDA is a transform that maximizes the inter-class variance, while minimizing the intra-class variance, resulting in a projection where examples from the same class are projected close to each other and, at the same time, the class centers (means) are as far apart as possible. Although it is not a true regularization technique, PCA-LDA is found to reduce the chance of over-fitting that may occur with a pure LDA model.
During the recognition step, the model transformed spectra acquired from test samples with an unknown classification into the associated model-space, after which, a classifier determined into which class (if any) the spectra belonged. The model classifier uses a multivariate normal distribution (MVN) for each model class. During the model building phase, these distributions are constructed by transforming the training spectra to generate scores for the n principal components/ linear discriminants selected for the model. The number of dimensions in the MVNs is also equal to n. The MVNs produced a likelihood measure for each class, and Bayes' rule was then applied to derive posterior probabilities.
In silico 5-fold stratified validation was performed to determine the predictive accuracy of the fish speciation model. The model building dataset was divided in five partitions (5-fold), each of which contains a representative proportion of each class within it (stratified). Four partitions (80%) of the dataset were used to build a model under the same conditions as the original model. This model was used to predict the classifications of the one partition (20%) of the training set that was left out. The cycle was repeated iteratively five times and each partition was predicted once by a model trained from the other four. The output of the validation details the total number of correct and incorrect classifications, as well as the number of outliers. Outliers were calculated according to the Mahalanobis distance8 to the nearest class center. If this distance was greater than the outlier threshold, the sample was considered an outlier.
Additional and complementary statistical analyses were performed using Progenesis QI (NonLinear Dynamics, Newcastle, UK), EZInfo, and SIMCA-P (Umetrics Sartorius Stedim Biotech, Sweden) to determine the chemical identifications of candidate biomarkers and potential involvement of discrete biochemical pathways.
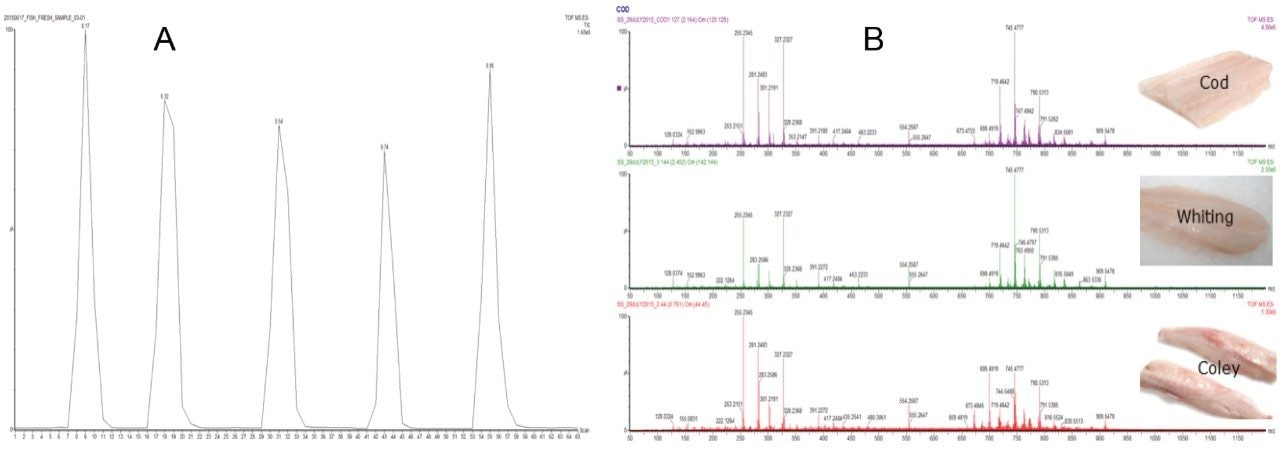
Raw spectrometric data (Figure 1) obtained from authenticated samples of cod (n= 194), coley (n=51), haddock (n=133), pollock (n=50), and whiting (n=50) giving a total of 478 samples were pre-processed and subjected to multivariate analysis where PCA followed by supervised LDA were applied using LiveID.
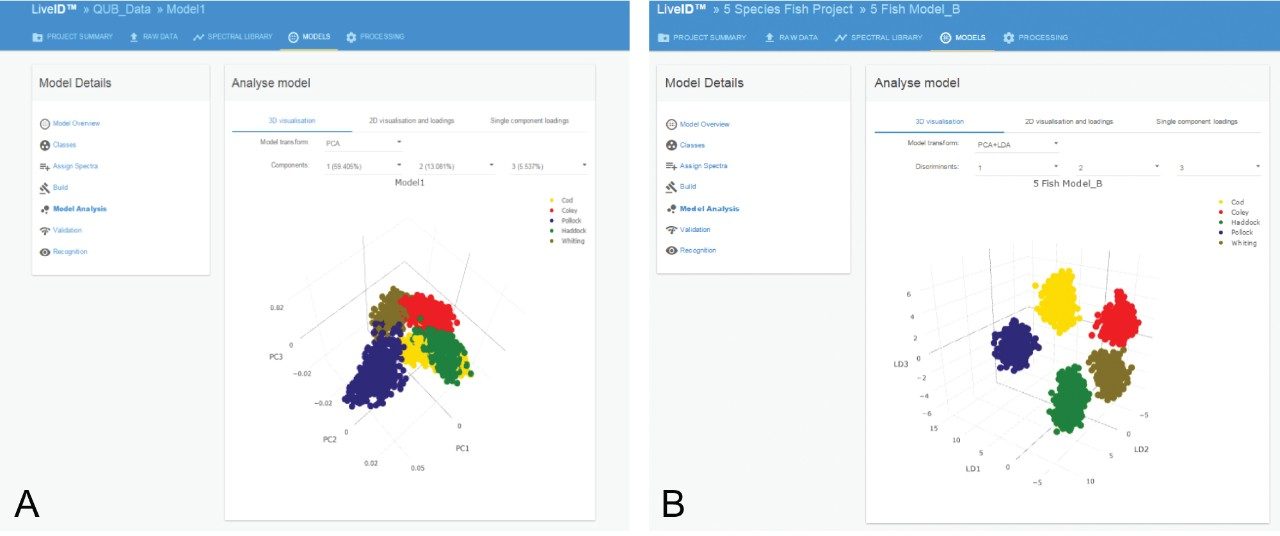
80 PCA components and 4 LDA components were used to generate the chemometric models. Clustering was apparent within the three-dimensional (3-D) PCA scores plot using components 1, 2, and 3 which explained approximately 78% of the variance (Figure 2a). However, clear separation between the five species of fish was obtained within the 3-D PCA/LDA score plot using components 1, 2, and 3 (Figure 2b).
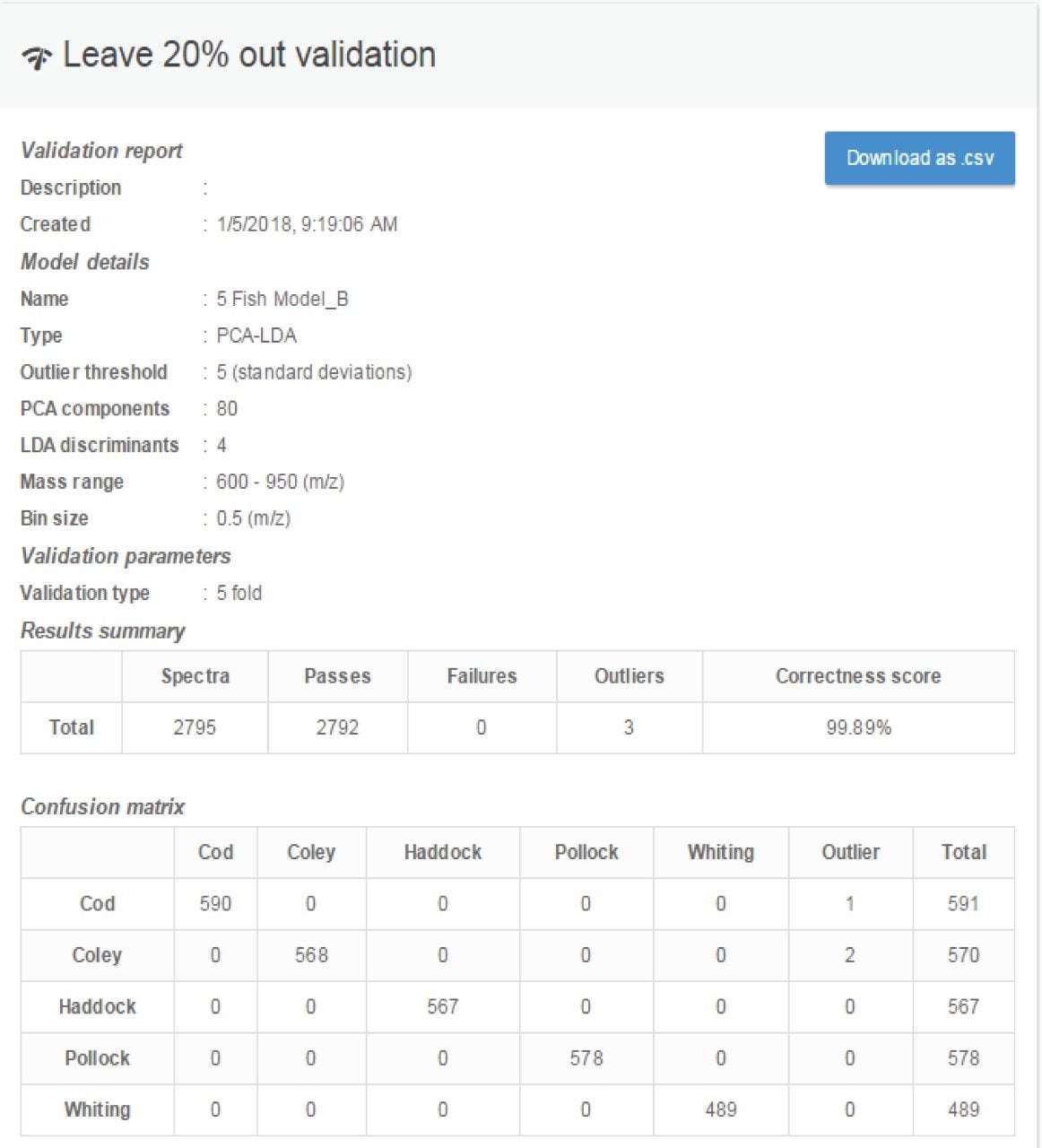
The multi-species fish classification PCA/LDA model was subject to in silico cross validation using the “leave 20% out” method (Figure 3). The validation resulted in a 99.9% correct classification with no misclassifications, with only one cod and two coley samples classified as outliers. The PCA/LDA model was also validated according to the “leave one file out” method whereby each of the training data files was systematically excluded from the model and classified as an independent sample; in this case a 98.9% correct classification with no misclassifications was achieved. A higher number of outliers were observed (data not shown).
An independent validation was carried out to ensure the validity of the results from the in silico validation. The raw data acquired from a set of validation samples was subjected to a cross validation similar to that of the leave 20% out in silico validation. The model was created using a reduced training set of samples (n=379) excluding 99 samples assigned as the validation set. Each validation sample was then assigned a fish species classification. An overall correct classification rate of 98.9% was obtained in perfect agreement to the classification rate obtained using the LiveID cross-validation tool.
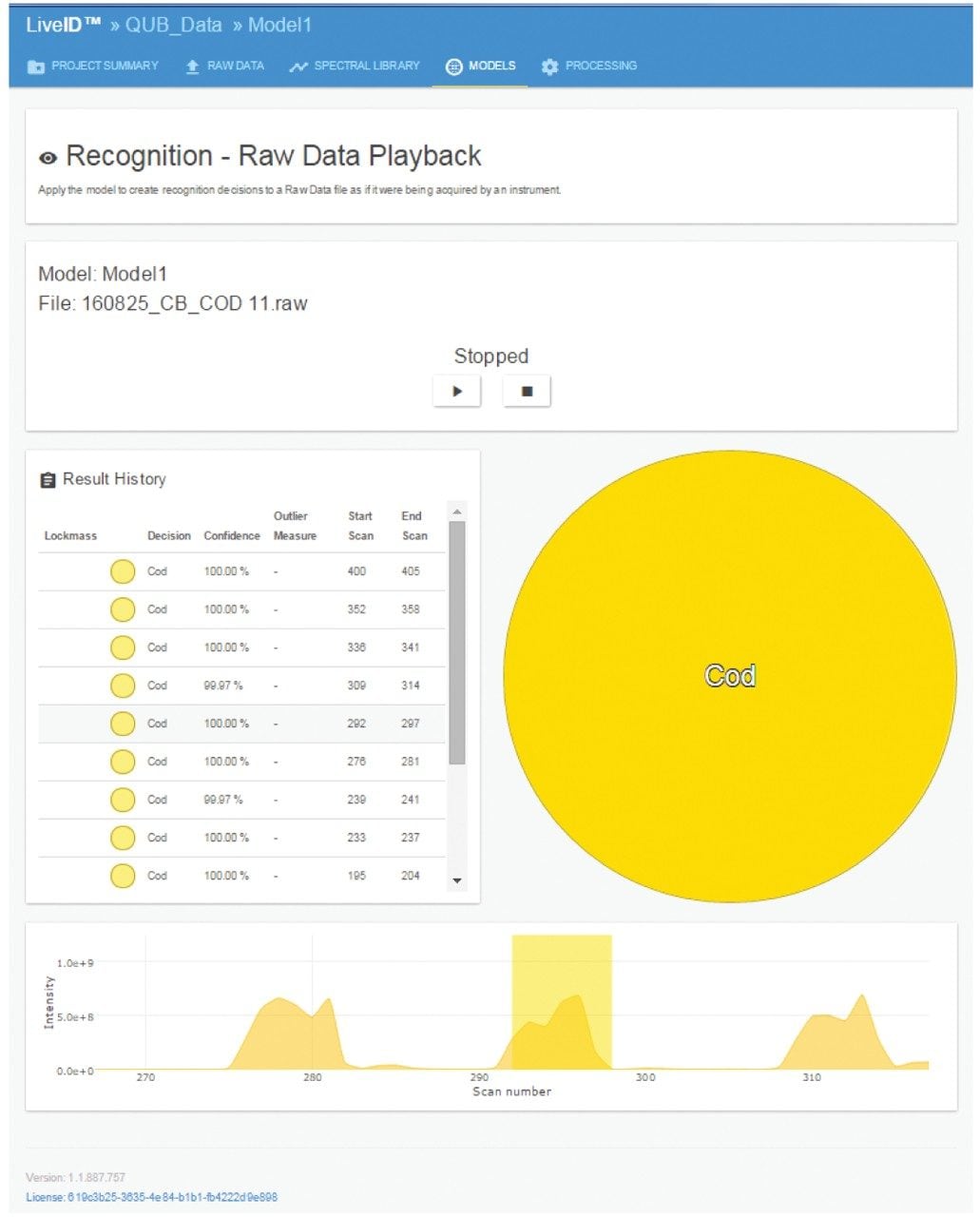
Following a successful build and validation, the PCA/LDA white fish model was used for realtime identification of fish samples. Raw data files were acquired and run live though the software providing a nearly instantaneous identification (Figure 4), excluding the delay between sampling appearance of a signal of approximately 2 s. A standard deviation of 5σ was used for class assignment. The spectral intensity limit was set at 1e8 counts thus ensuring that only the cuts were assigned a species classification and not any background noise. In all cases, the sample was correctly identified.
As a test of the inter-laboratory repeatability of the model, a subset of 68 of the training samples (representing approximately 14% of the total population) were sent to a second laboratory facility and analyzed using a different REIMS instrument. The second site's data was classified using the training set data generated at the primary site and resulted in a 95.6% correct classification rate which was due to three haddock samples being misclassified as cod.
To determine classification fidelity, raw spectrometric data obtained from authenticated samples of fish species not represented in the model [seabass Dicentrarchus labrax (n=6) and seabream Sparas aurata (n=8)] were run through the LiveID playback recognizer feature to obtain classification results. Of the 14 samples analyzed, 13 (92.8%) were correctly recognized as “outliers” with one sample being classified as both an outlier (with 66% predictive certainty) and coley (34% predictive certainty) within the multiple burn regions.
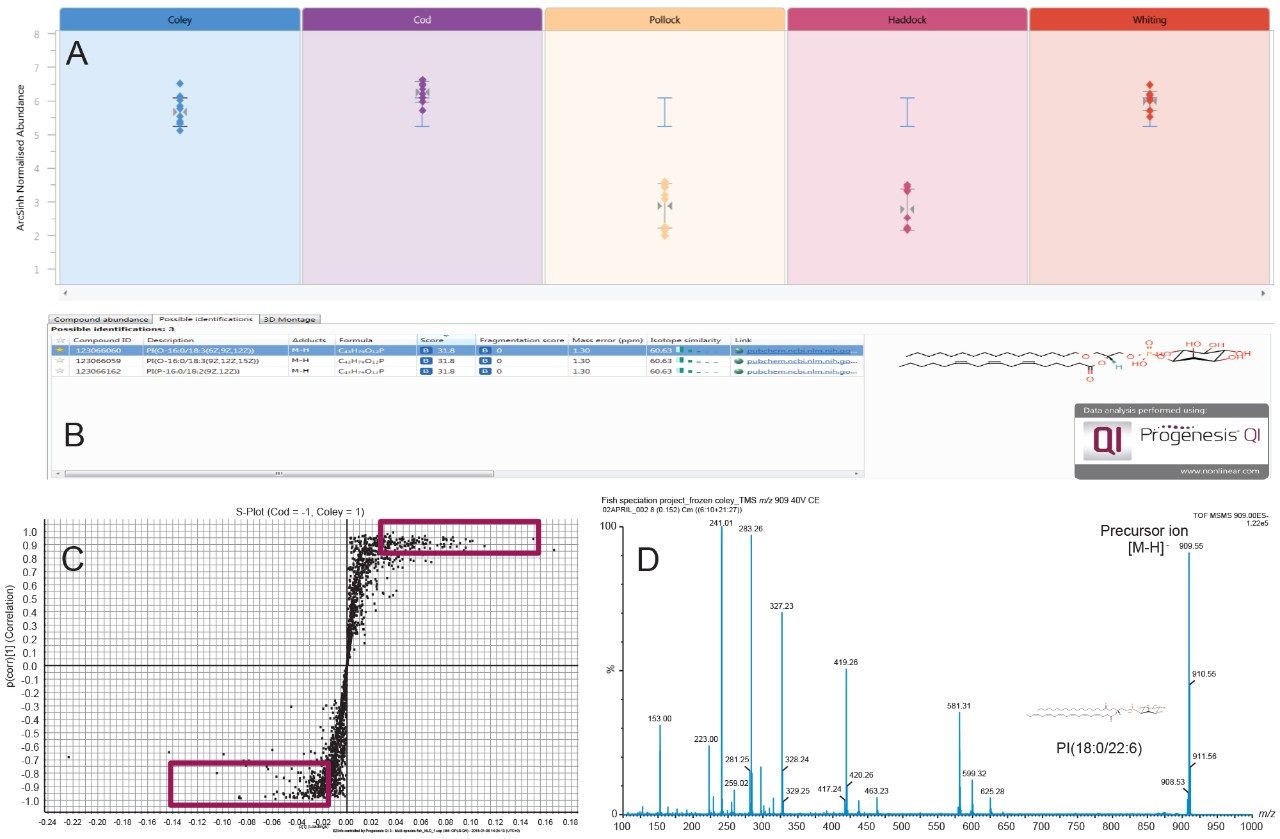
MS data files were processed through MassLynx Software's Sample List using the Progenesis Bridge application to convert the files that contained multiple sampling events (burn regions) into an individual file per burn region in the format of a Gaussian peak. Lock mass correction and ABS were also performed during this step. The pre-processed data files were subsequently imported into Progenesis QI Software (v.2.4) and a direct analysis workflow was followed to generate multivariate statistical models and feature abundance plots (Figure 5A).
EZInfo (v.3.0.0.0) was used to create a series of OPLS-DA S-plots to determine the significant ions responsible for species level separation in the PCA/LDA model. Ions present at the upper and lower extremity regions of the S-plots (highlighted in the red boxes Figure 5C) were deemed to be the significant ions involved in species classification and were selected for database searching within Progenesis QI using ChemSpider and LipidMaps databases (Figure 5B). Subsequent REIMS MS/MS experiments were performed whereby the precursor ion was isolated in the quadrupole region of the Xevo G2-XS QTof and a collision energy of 25 eV was applied to yield fragmentation spectra to assist with the chemical elucidation and tentative identification process (Figure 5D). Interpretation of the spectra revealed that members of the diacylglycerophosphoethanolamine (PE), phosphatidylinositol (PI), sphingomyelin (SM), and free fatty acid classes had a significant involvement in the differentiation of fish species.
REIMS with LiveID Technology requires no sample preparation and provides accurate and nearly instantaneous results. The reduced amount of time required for analysis and data interpretation that REIMS offers in contrast to the current PCR-based methods represents a significant improvement in operational efficiency. REIMS with LiveID has been demonstrated as a complementary technique for the detection of commercial fish fraud.
Along with speciation, REIMS is able to detect multiple aspects of fish fraud e.g. the separation of line and trawl caught haddock samples. By employing this technique, we may also be able to differentiate other aspects such as geographic origin and wild fish versus farmed fish; areas where genomic profiling alone would not be useful.
For further details on this study please refer to the journal article: Connor Black, Olivier P, Chevallier, Simon A. Haughey, Julia Balog, Sara Stead, Steven D. Pringle, Maria V. Riina, Francesca Martucci, Pier L. Acutis, Mike Morris, Dimitrios S. Nikolopoulos, Zoltan Takats, Christopher T. Elliott. A real time metabolomic profiling approach to detecting fish fraud using rapid evaporative ionisation mass spectrometry. Metabolomics 2017, DOI 10.1007/s11306-017-1291-y.
720006205, February 2018