This is an Application Brief and does not contain a detailed Experimental section.
This technology brief demonstrates the workflow for peptide MRM optimization using the Xevo TQ-XS Triple Quadrupole Mass Spectrometer, Skyline software and phosphorylation B peptide standard.
Moving from discovery to targeted proteomic experiments involves MRM transition optimization and refinement of many peptides from a protein. Compounding the process is the need to utilize peptides from a protein digest leading to a relatively complex sample, which can only be injected on-column to perform these optimization experiments. Similarly, quantification from a purified or synthetic peptide can have many of the same complexity challenges. Such iterative injection and review can be time consuming. The use of software to facilitate MRM assignment and visualization of the results can speed up this process to a few injections on-column. Skyline is one such, open source software that aids in the development of SRM /MRM transition optimization by on-column injection of sample. The software can perform in silico digestion of a protein using a variety of enzymes. Skyline works with MassLynx Mass Spectrometry Software to create MRM methods, then visualize the raw data, and finally, automatically refine MS methods for routine analysis. This technology brief describes how to set up the software and perform such an experiment.
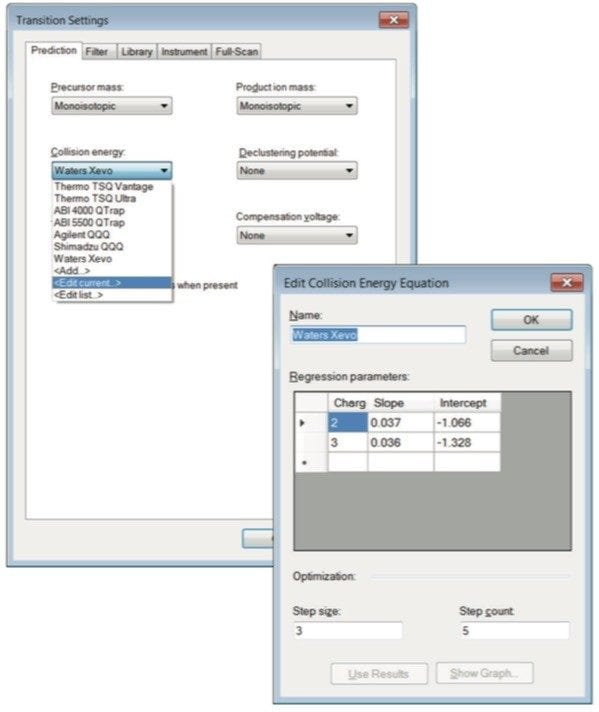
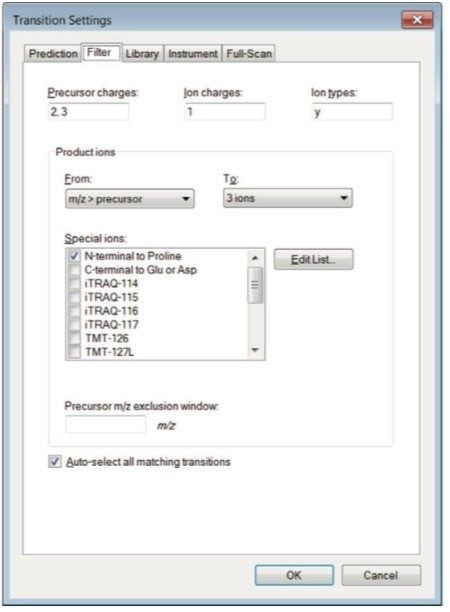
Step 1. Configure Skyline for Waters mass spectrometers by changing the collision energy to “Waters Xevo” in Settings > Transition Settings > Prediction tab (Figure 2). Additionally, the number of collision energy steps and intervals to investigate can be modified by clicking on ”Edit current” under “Collision energy.” The optimization ”step size” can be set to 2 and the ”step count” 5. Step count is the number of different collision energies to try above and below a predicted value, and step size is the difference between those collision energies. Under the Filter tab, change ”precursor charges” to 2, 3. The “ion charges” (charges on the product ions) value could be 1 or 1,2 –useful for triply charged precursors, and the ”ion types” to fit your experiment, typically y or y,b. Select the product ions for Skyline to display. We have chosen m/z > precursor, 3 ions, and N-terminal to Proline as shown in Figure 3.
Step 2. When performing in silico digestion, the parameters should be set to match your experiment. For example, under Settings > Peptide Settings > digestion tab the default choice is Trypsin as the enzyme and zero Max missed cleavages (Figure 4).
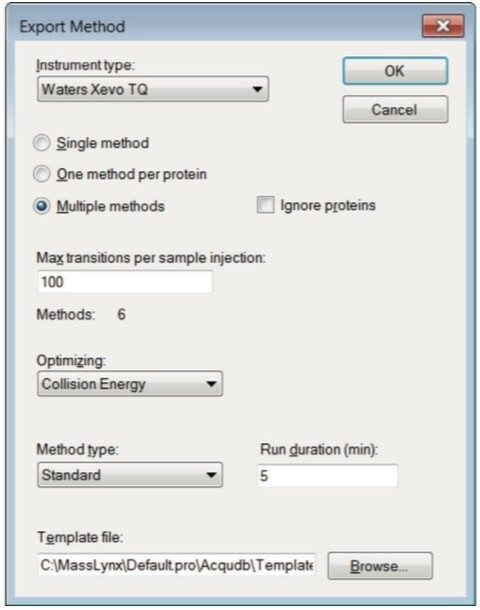
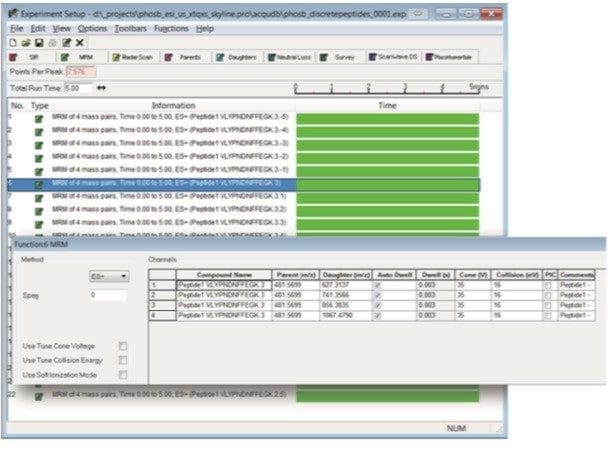
Step 3. Populate the peptide list by copying and pasting peptides individually in a list, or by importing a FASTA file. If you are working with proteins, you may paste the protein sequence to allow Skyline to perform in silico digestion. Create the MRM methods for injection under MassLynx by selecting File > Export > Method. Choose Waters Xevo for instrument type, select “Multiple methods”, “Max transitions per sample injection” = 100, select “Collision Energy” for “Optimizing,” “Method type” is “Standard”, and browse in an MRM method template (Figure 5). With these settings, our optimization will be complete in 6 injections taking a total of 30 minutes. Figure 6 shows an example of the MS method created in Skyline. In this method there are 22 functions with a total of 88 transitions to be monitored.
Additional notes: “Max transitions per sample injection” is a parameter that can be adjusted according to the quality and concentration of sample as well as length and resolving power of the LC method. Choosing too high of a number of transitions can impact the data quality by reducing the number of points per peak in the optimization. Additionally, the step size and step count can also be modified to ensure there are enough points per peak in the acquisition method created by Skyline. A consideration for shortening the number of injections is to use Scheduled chromatography in the method type. Scheduling will create time windows for the MRM transitions around the predicted retention time of a peptide and therefore more transitions can be added into a single MS method without compromising data quality.
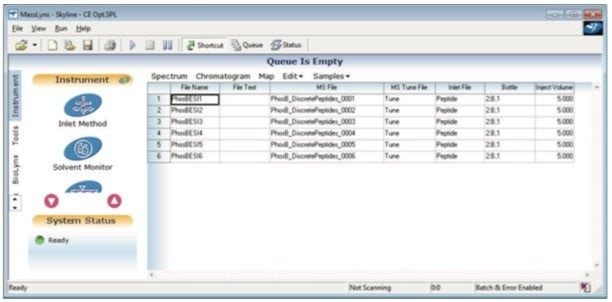
Step 4. Create the Sample List in MassLynx, browse in the individual MS methods and perform the injections (Figure 7).
Once the injections have finished, the data can be imported into Skyline for processing and viewing.
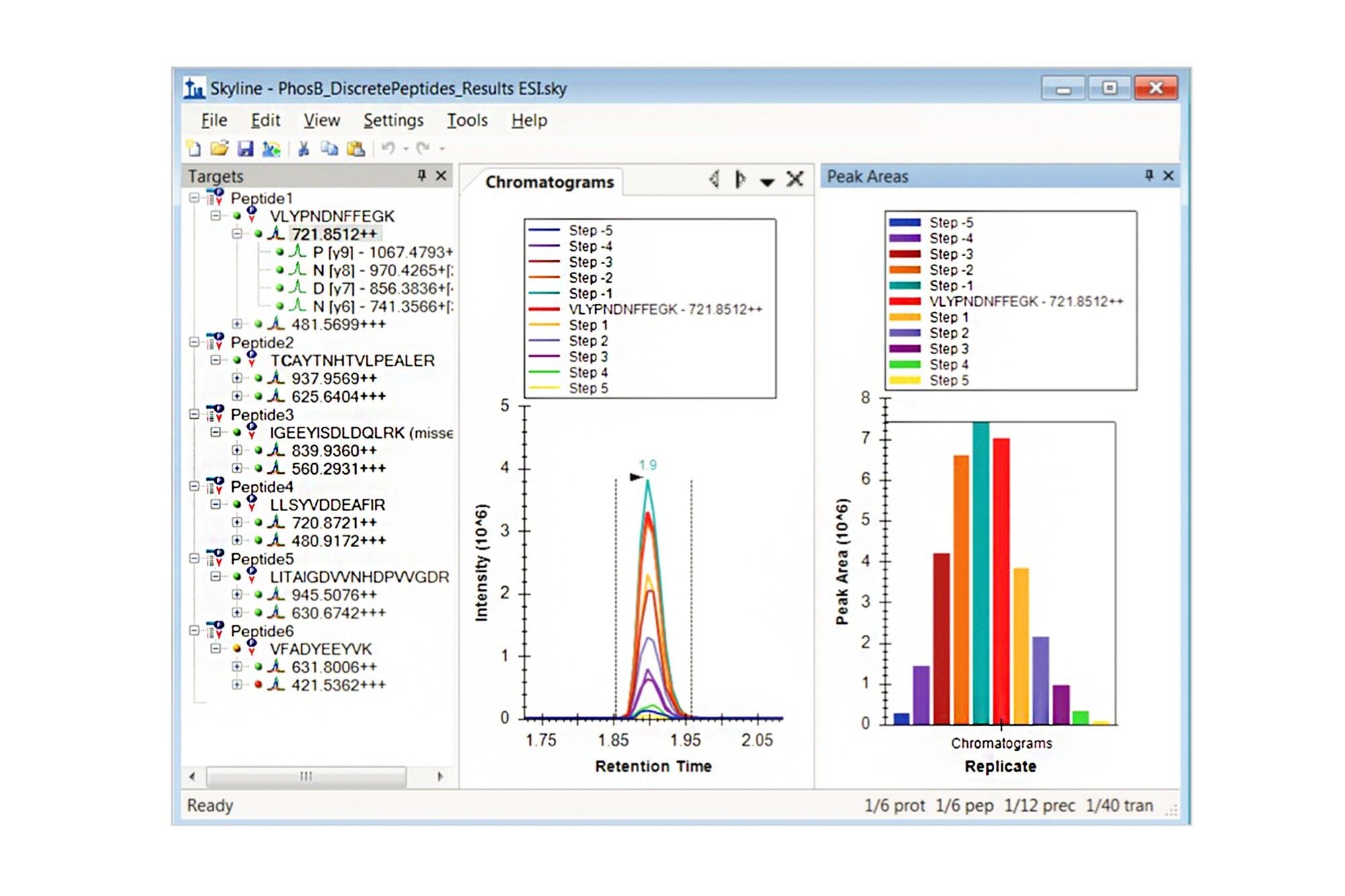
Step 1. In Skyline, go to File > Import > Results Add one new replicate and select “Collision Energy in Optimizing. Shift”, click to select all of the optimization injections and import. Once all data is in, you will need to change your view settings. Select View > Transitions > Single (or F10), View > Auto-Zoom > Best peak (or F11), and View > Peak Areas > Replicate comparison (or F7). Figure 1 shows the collision optimization review in Skyline for VLYPNDNFFEGK. The chromatogram overlay shows the results for the various collision energies and the bar graph is the normalized peak area for the injections.
Step 2. Data can be reviewed and decisions on appropriate collision energy can be manually determined by scrolling through each transition and taking note of the collision energy with the highest signal. In a large dataset where you will be working with peptides from many proteins, this manual review can be time consuming. In such a case, automatic refinement can be performed. Select Edit > Refine > Advanced, and on the Results tab you can automatically remove poor transitions by setting the Min and Max peak found ratios to keep or discard, set “Max transition peak rank” to the number of transitions per peptide you want, and select “Remove nodes for missing results” (hover over each field for an explanation). You can then delete the peptides or individual transitions you do not want to measure. Finally go to Settings > Transition settings and select the box ”Use optimization values when present” and “Optimize by Transition”. Now, when exporting the MS Method under File > Export, the best values for collision energy will be automatically written into the method.
Peptide quantitation has become a routine laboratory process. However, peptide MRM transition optimization can be more complex and, therefore, more time-consuming than small molecule optimization. The use of Skyline visualization software with the Xevo TQ-XS streamlines this optimization and acquisition.
720005926, February 2017