For research use only. Not for use in diagnostic procedures.
This application note describes the use of AccQ•Tag Ultra reagent combined with UPLC-ESI-MS/MS for the analysis of amino acids and biogenic amines in human plasma or serum via a high-throughput, sensitive, and specific method.
Blood concentrations of amino acids and other amino-containing compounds change in response to different conditions, including toxicity and metabolic diseases (inborn errors, metabolic syndrome, diabetes, obesity, cancer, etc). As such, there is a need for a rapid, targeted method for their quantification in biological fluids to support metabolic phenotyping studies. Currently, mass spectrometry- (MS) based methods for the quantification of amino compounds using MS for detection include procedures based on GC-MS, CE-MS/MS, and LC-MS/MS.1–3 Reverse-phase liquid chromatography (RPLC)-based methods are complicated by the polar, amphoteric nature of these analytes, which can result in poor retention and make direct analysis impractical for all but a few analytes. While alternative modes of liquid chromatography (cation exchange, HILIC, ion-pairing) can be used, they are not without disadvantages. As a result, modifying the chromatographic properties of this class of compounds to form more retentive derivatives for RPLC remains popular. 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate (AccQ•Tag reagent) was originally used to analyze primary and secondary amines via fluorescence detection, but can also enable detection of these analytes via LC-MS. This application note describes the use of AccQ•Tag Ultra reagent combined with UPLC-ESI-MS/MS for the analysis of amino acids and biogenic amines in human plasma or serum via a high-throughput, sensitive, and specific method.
Human plasma/serum (10 µL) is mixed with Optima™ grade water (10 µL), 5 µL of a 10 µg/mL internal standard (IS) mixture, and 40 µL of cold isopropanol–0.1% formic acid (v/v). Next, it is vortexed to precipitate proteins. After 20 minutes (-20 °C), samples are centrifuged (≥10,000 g, 10 minutes). For derivatization, 10 µL of supernatant mixed with 70 µL of borate buffer (pH 8.6) was treated with 20 µL of the AccQ•Tag Ultra reagent solution (prepared according to Waters® instructions) with vortex mixing, heated at 55 °C (10 minutes), then diluted 1 in 100 with Optima grade water for UPLC-MS.
The method was validated, as far as practicable; using the approach outlined in the FDA “Guidance for Industry” Bioanalytical Methods – which covers aspects such as intra- and inter-assay precision, specificity, carryover, recovery, matrix effects, and stability.
|
LC system: |
ACQUITY UPLC I-Class |
|||
|
Detection: |
Xevo TQ-S |
|||
|
Column: |
ACQUITY UPLC HSS T3, 1.8 μm, 2.1 mm x 150 mm |
|||
|
Column temp.: |
45 °C |
|||
|
Sample temp.: |
4 °C |
|||
|
Injection volume: |
2 μL |
|||
|
Flow rate: |
0.6 mL/min |
|||
|
Mobile phase A: |
0.1% formic acid in H2O (v/v) |
|||
|
Mobile phase B: |
0.1% formic acid in ACN (v/v) |
The starting composition was 4% B held for 0.5 minutes, before increasing to 10% over 2 minutes, then to 28% over 2.5 minutes, increasing to 95% for 1 minute to wash the column, and finally returning to 4% B for a 1.3 minute re-equilibration step.
|
MS system: |
Xevo TQ-S |
|||
|
Ionization mode: |
ESI+ |
|||
|
Acquisition range: |
MRM detection |
|||
|
Capillary voltage: |
1 kV |
|||
|
Collision energy: |
Analyte-dependent |
|||
|
Cone voltage: |
50 V |
|||
|
Data management: |
MassLynx Software with TargetLynx Application Manager |
The AccQ•Tag Ultra reagent reacts with primary and secondary amines, giving derivatives with good chromatographic and mass spectrometric properties. MS conditions for positive ESI were obtained via direct infusion of the individual derivatives for determination of the parent ion and best multiple reaction (MRM) transitions, dwell times, cone voltages, collision energies, etc.
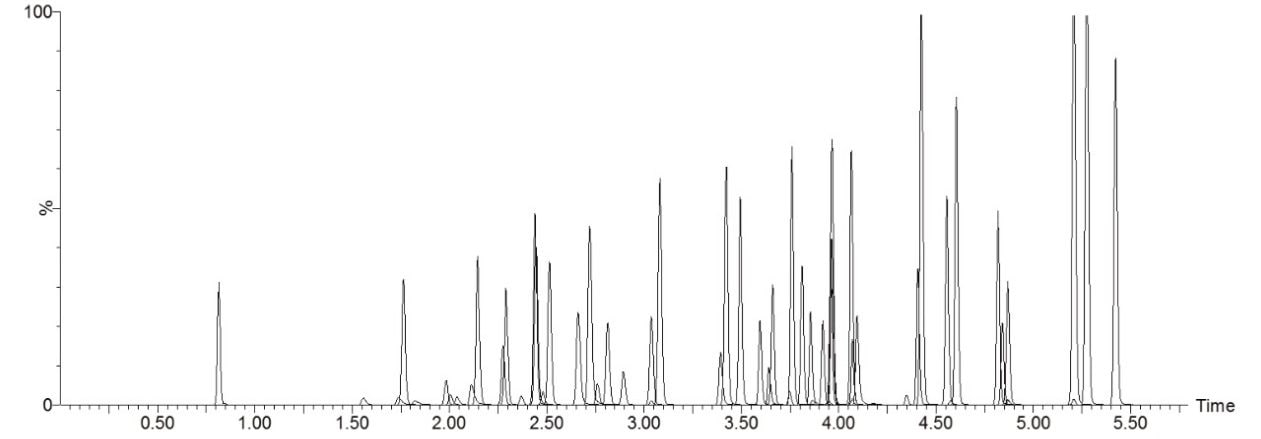
The AccQ•Tag Ultra reagent gives rise to a common fragment ion at m/z 171, generated by a loss of the aminoquinoline (AMQ) moiety; in most – but not all – cases the combination of parent ion and this common fragment was selected for detection. Monoamines form a mono-ACQ derivative, giving the [M+AccQ+H]+ ion, while polyamines (e.g. cystine, lysine) can form derivatives containing more than one ACQ unit. The chromatographic method requires only 7.5 min/sample including column re-equilibration, allowing high-throughput analysis (Figure 1). A good separation is provided for a number of isomers/isobars such as 1- and 3-methylhistidine; sarcosine, β-alanine, and alanine; γ-amino-n-butyric acid, β-aminoisobutyric acid, and α-amino-n-butyric acid; isoleucine and leucine.
Because obtaining true biological blanks for these analytes is not possible, standard curves and QC samples were prepared in 50:50 water–methanol (v/v). Investigated were the linearity; lower and upper limits of quantification (LLOQ, ULOQ); inter- and intra-day accuracy and precision; specificity; carry over; recovery; matrix; and analyte and derivative stability. Standard curves were linear over the range 1–400 µm/mL (effectively 2–800 µm/mL for plasma/serum due to sample dilution prior to derivatization) with correlation coefficients (R) of 0.993 or better determined over the three days of the validation. Stock solutions of the underivatized analytes were stable when either left for six hours at ambient temperature or frozen at -20 °C for 48 hours. The derivatized analytes were stable at either ambient temperature or in the autosampler at 4 °C for up to one week. Carryover was found to be acceptable at 11% or less for all of the analytes tested while carryover of the internal standards was less than 1%.
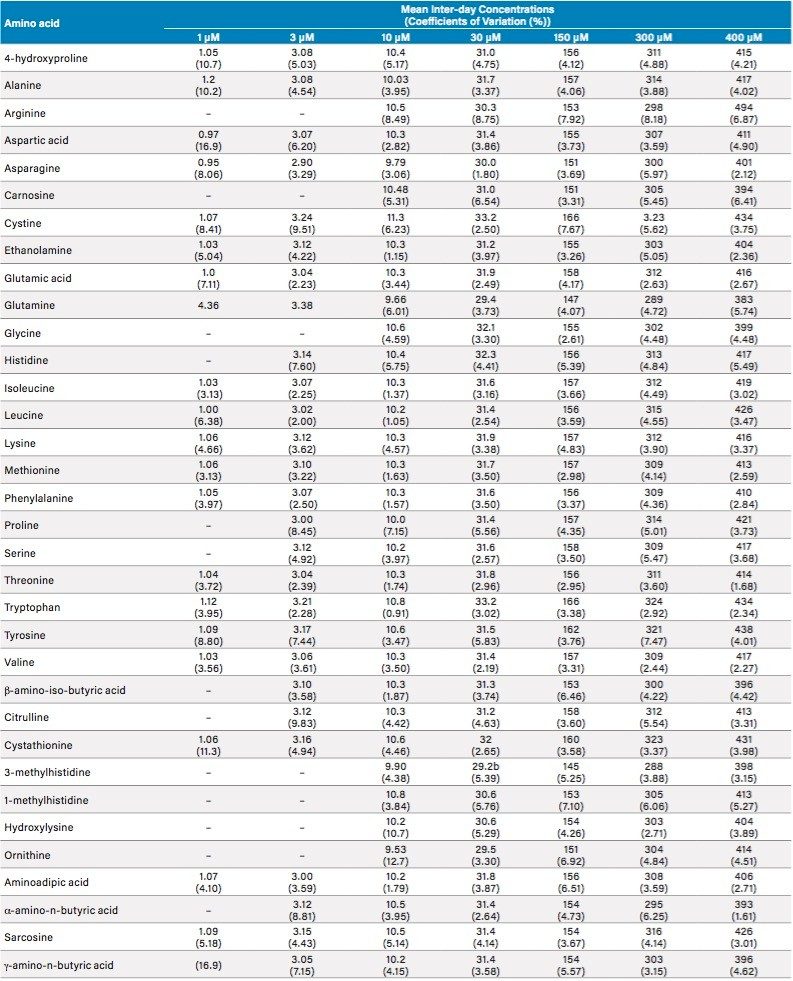
The intra- and inter-day performance of the assay was evaluated by the analysis of samples on three separate occasions on three separate days over the concentration range of 2–800 µm, and the method was found to be reproducible and accurate. For both intra- and inter-day analysis, a lower limit of quantification of (LLOQ) of 2 µm was obtained using isotopically labeled internal standards for 17 of the amino acids (inter-day data in Table 1). A further eight of the target gave acceptable results using a LLOQ of 6 µm, while the remaining seven compounds considered for absolute quantification had LLOQs of 20 µm (Table 1). The ULOQ for all analytes was 800 µm. The additional 20 amino compounds monitored here for relative quantification were not included in the validation study.
The intra- and inter-day precision of the assay ranged from 0.91–16.9% for the intra-day determinations and 2.12–15.9% for the inter-day comparison. The corresponding values for intra- and inter-day accuracy were 0.05–15.6% and 0.78–13.7% respectively. Specificity, in terms of matrix and other interferences, was assessed and found acceptable for matrix to analyte (at the LLOQ), matrix to internal standard, analyte to internal standard, and internal standard to analyte interference, etc.
The use of 6-aminoquinolyl-N-hydroxysuccinimidyl carbamate (AccQ•Tag reagent) as a derivatizing reagent for the targeted analysis of amino acids and other amino-containing compounds provides these analytes with good reverse-phase liquid chromotography and mass spectrometry properties, and allows the sensitive and specific analysis of this class of compounds in human serum and plasma in methods that are well suited to high-throughput analysis using UPLC.
720005882, February 2017