UNIFI enables fast, rapid, and flexible quantitation of data obtained from a wide variety of HRMS acquisition modes. This application note enables the user to collect HRMS data for quantitation, set up a simple quantitative processing method, and review the results within UNIFI. Setup and processing for both full scan and targeted Tof-MRM modes are described. Advanced parameters such as managing multiple charge states, isotopes, fragments, summing transitions, and tools for use with ion mobility modes of acquisition are covered as well.
Time-of-flight instruments offer a unique approach for the quantitation of compounds, which is complementary to traditional workflows found on tandem quadrupole instruments. With many more modes of selectivity available, analysts can quickly address complex matrix background interference and isolate peaks of interest in complex samples to generate sensitive and accurate analyses.
UNIFI enables fast, rapid, and flexible quantitation of data obtained from a wide variety of HRMS acquisition modes. It supports the construction of quantitative methods for HRMS full scan MS, MS,E MS-MS, or Tof-MRM data sets (and supports additional channels such as PDA, UV, FLR, and eSatin input from third party detectors). Flexible quantitation of single or combined adducts, multiple charge state, isotope, and ion mobility filtered extracted ion chromatograms (XICs) is also possible.
This application note enables the user to collect HRMS data for quantitation, set up a simple quantitative processing method, and review the results within UNIFI. Setup and processing for both full scan and targeted Tof-MRM modes are described. Advanced parameters such as managing multiple charge states, isotopes, fragments, summing transitions, and tools for use with ion mobility modes of acquisition are covered as well.
Many of the quantitative processing capabilities are available within the UNIFI Screening Analysis types, where a significant amount of characterization is desired (quan/qual type data). The solution and workflows described herein are comparable to many of the tools developed in TargetLynx (MassLynx) for high throughput quantitative assays.
Oxytocin (Figure 1) was used as a model compound to demonstrate quantitative modes. A dilution series of oxytocin was prepared in quenched human plasma (2:1 acetonitrile–water) at concentrations of 19 pg/mL to 1250 pg/mL. These samples were analyzed using two acquisition modes – a quan/qual HRMS approach (MSE) or a purely quantitative, HRMS Tof-MRM approach. The example datasets were processed several ways using settings described below in order to demonstrate options available for quantitation. More information on the merits (sensitivity, linearity, dynamic range, etc.) of different modes of acquisition have been described elsewhere.3
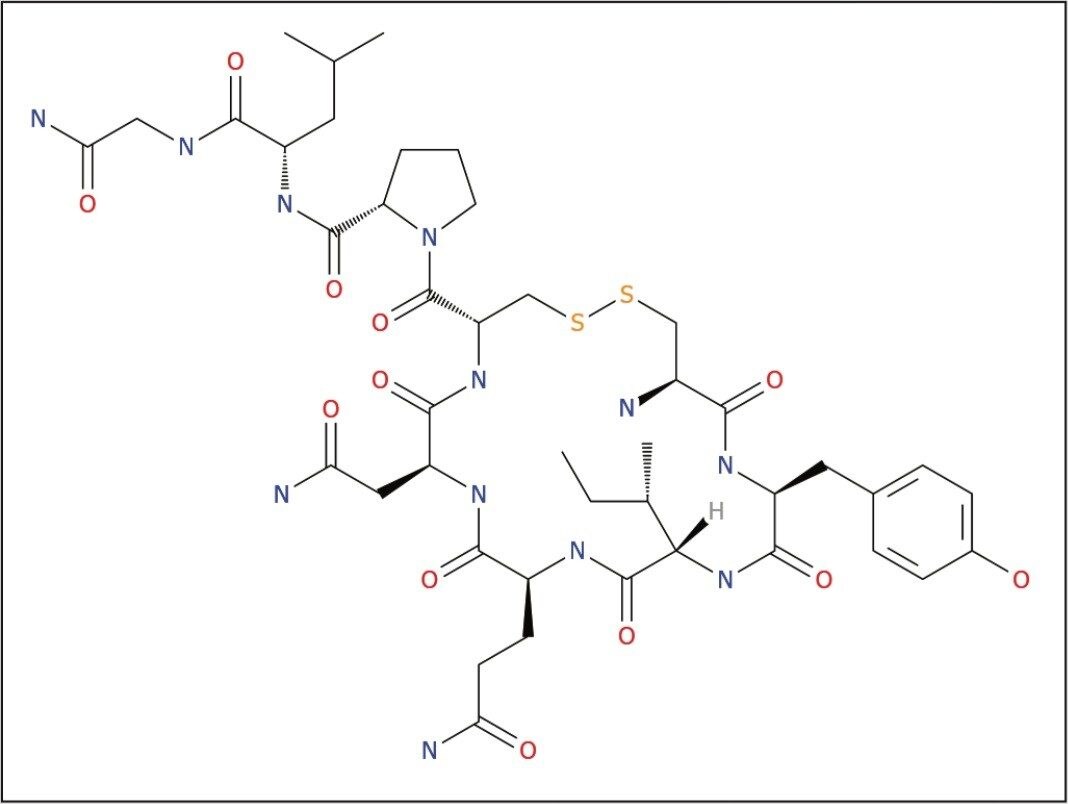
Samples were analyzed using a Xevo G2-XS QTof Mass Spectrometer collecting either MSE (generic precursor scan also collecting full fragmentation patterns) or Tof-MRM with target enhancement.
|
LC system: |
Waters ACQUITY UPLC I-Class System (FTN and BSM) |
|
Column: |
ACQUITY UPLC BEH C18, 1.7 μm, 2.1 mm x 50 mm (P/N 186002350) |
|
Column temp: |
60 °C |
|
Sample temp: |
10 °C |
|
Injection volume: |
5 μL |
|
Flow rate: |
0.6 mL/min |
|
Mobile phase A: |
Water + 0.1% Formic Acid |
|
Mobile phase B: |
Acetonitrile + 0.1% Formic Acid |
|
Gradient: |
2 to 50% B over 3 min, 50% to 95% from 3 to 3.5 min, hold 95% from 3.5 to 4 min, return to 2% at 4.1 min.Total run time 5 min. |
|
MS system: |
Xevo G2-XS QTof |
|
Ionization mode: |
ESI+, sensitivity mode |
|
Acquisition range: |
50–1200 Da |
|
Capillary voltage: |
1 kV |
|
Collision energy: |
MSE (4 eV low, 20–30 eV high) |
|
Collision energy: |
Tof-MRM (settings described in results) |
|
Cone voltage: |
40 V |
UNIFI Scientific Information System 1.8.1
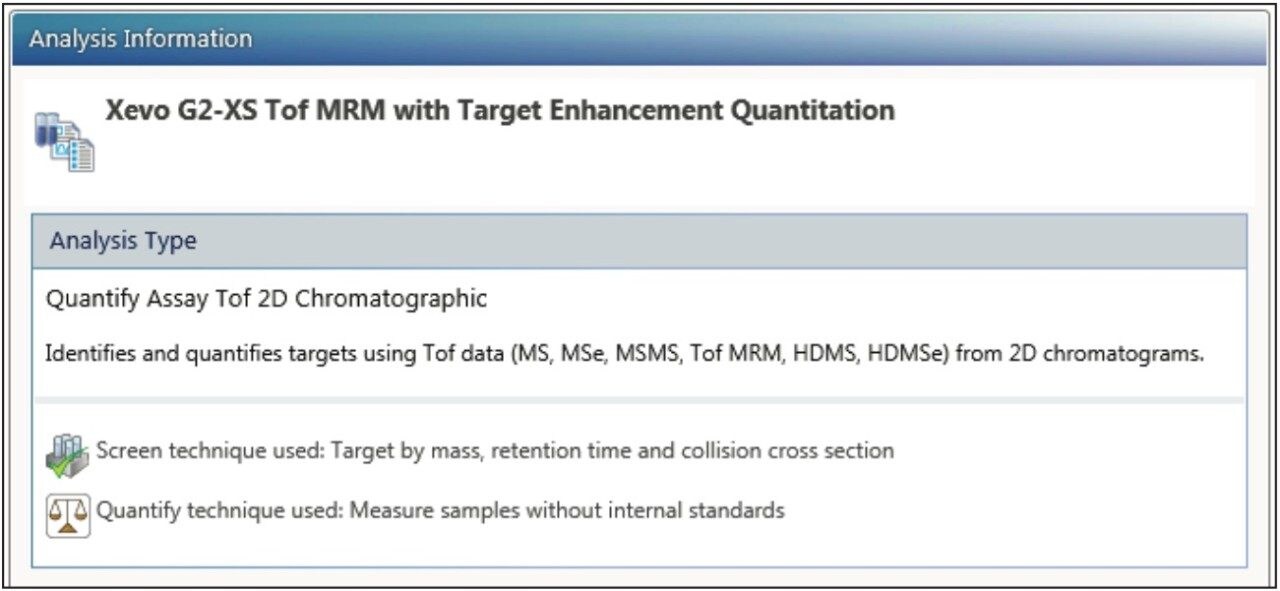
Acquisition and processing of data directly through the UNIFI Quantify Assay Tof 2D Chromatographic Analysis method option are described for multiple modes of acquisition (Figure 2). Unlike screening or biopharm workflows that focus on identification, this mode does not perform any characterization or qualitative processing. It is an optimized workflow for generating HRMS XIC (or ion mobility XICs) trace(s) and performing quantitation analogous to a SIR (single ion recording) or MRM (multiple reaction monitoring) type workflow.

The experiments and acquisition/processing modes are presented in four sections; the contents are described briefly:
1. Full Scan Data, Acquisition, and Processing Parameters
2. Targeted Data, Acquisition, and Processing Parameters
3. HRMS Quantitation Analysis Processing Parameters
4. Reviewing Quantitative Results
Full Scan Data, Setting Up Acquisition
Any mass range supported by the Tof platforms may be processed. This also includes HD (ion mobility enabled) types. A typical instrument setup for MSE data acquisition is shown below in Figure 3.
Two examples are described:
a) Setting up a single transition for quantitation, [M+H]+
b) Summing two transitions for quantitation, [M+H]+ + [M+2H]2+
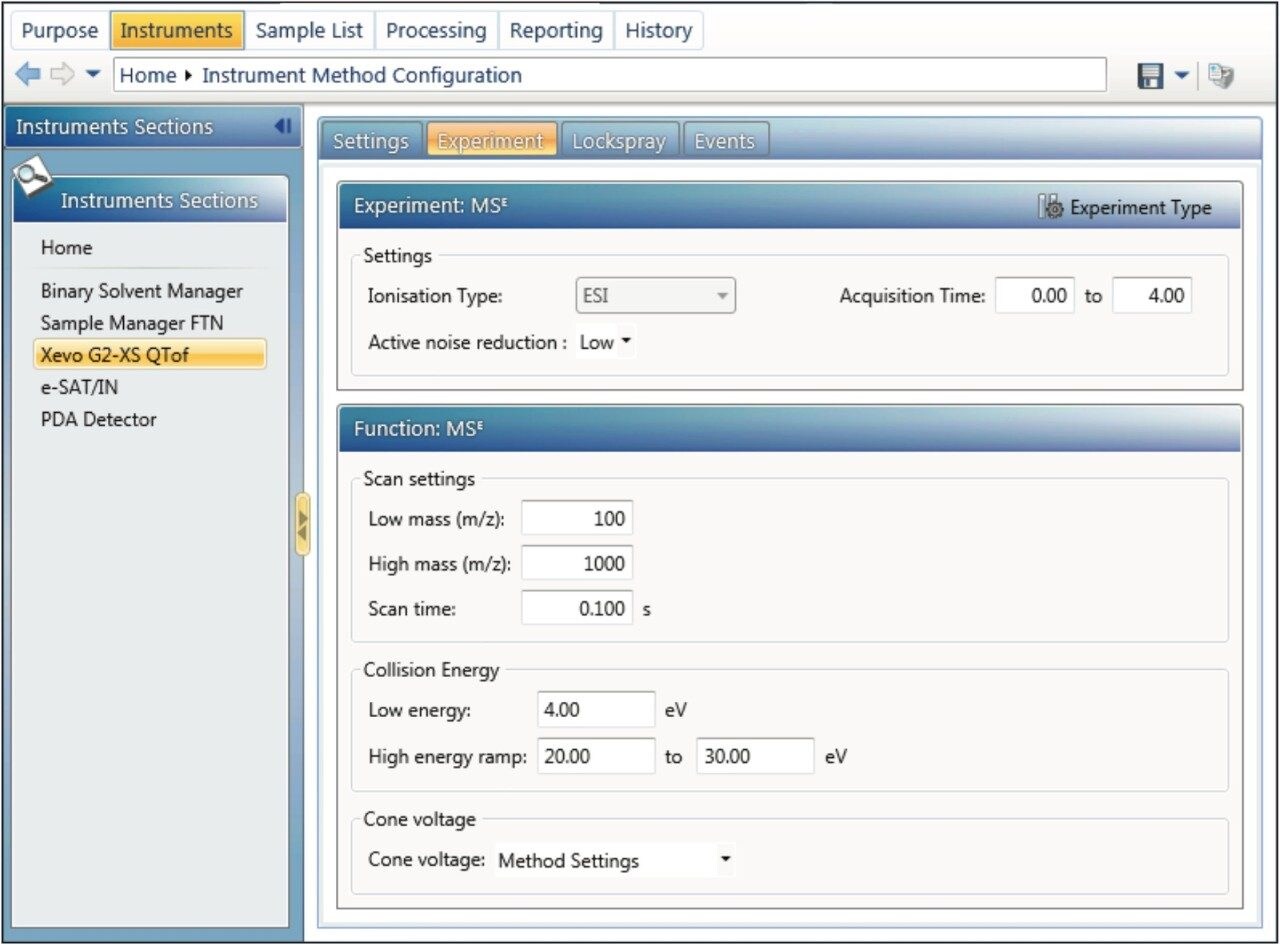
In order to correctly process the data, the processing method must contain values needed to generate the desired XICs. The XIC trace may need to be defined as low energy or high energy. Alternatively a function number may be specified. If no specific energy level or function is described, then all channels in the sample which contain the m/z value for the component are processed.
It is possible to quantitate either the primary ion of interest [M+H]+ (Figure 4a), or to sum the responses from both singly and doubly charged ions (Figure 4b/c). The target values for the ions may be imported from UNIFI’s Scientific Library by clicking the “Import” button. If the compound has detection results (has been analyzed and the MS-MS results have been previously sent to library), then the actual ions of interest can be selected, and the masses for quantitation (precursors and/or fragment ions) can be imported directly. Utilizing the library helps reduce transcription errors for setting up HRMS transitions. In this example, the XICs for oxytocin are extracted with a 50 mDa window (retention time window is 2 min, so the XIC +/– 1 min with respect to the expected retention time is displayed in the results; if no time window is specified, the entire XIC range is displayed). Tolerances that are narrower (<50 mDa) or wider (>50 mDa) may provide different degrees of selectivity and can be fine-tuned for each compound and matrix.
Targeted Data, Setting Up Acquisition for Tof-MRM
Two examples are described:
a) Setting up an untuned Tof-MRM – 1007.4>1007.4437
b) Setting up a tuned Tof-MRM for a fragment ion – 1007.4>723.2589 CE 28 V
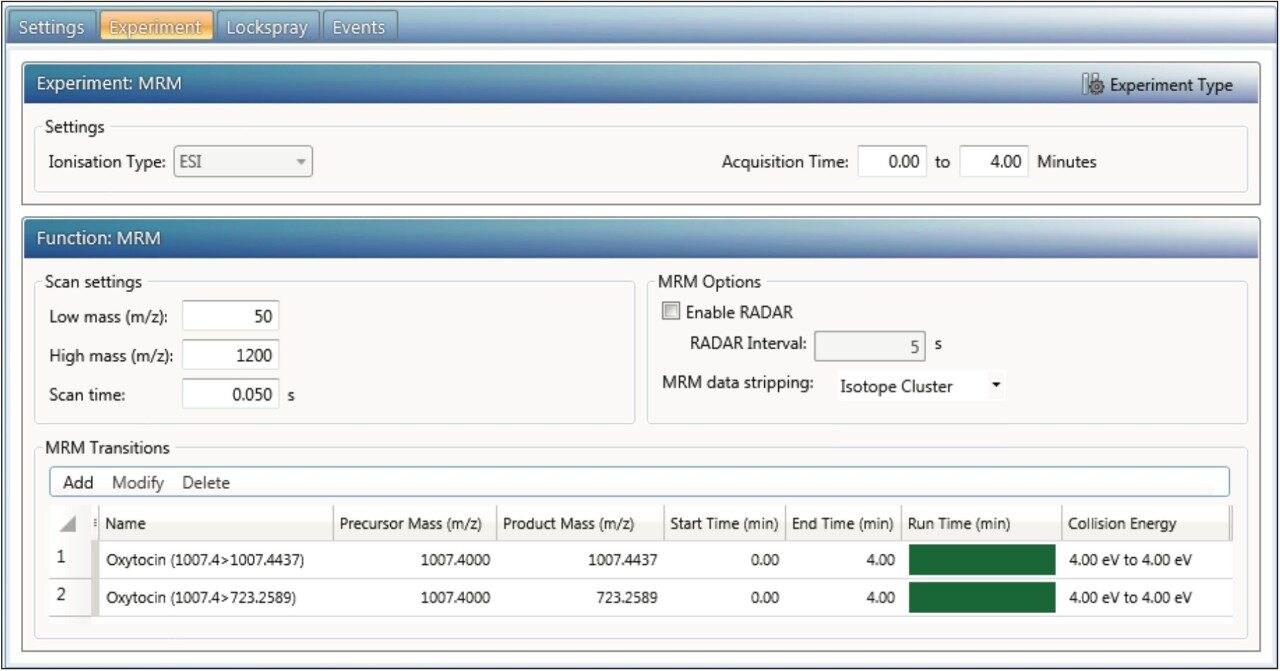
Tof-MRM setup is straightforward. A precursor mass (Q1, nominal set mass) and product mass (HRMS product ion) are specified in the method as shown in Figure 5. RADAR is an acquisition mode in which both MRM and full scan MS are acquired simultaneously. It may be used in method development to determine whether the regions under the MRM signals coelute with highly concentrated or interfering background matrix ions. The interval at which full scan spectra are collected (by default, every 5 seconds) can be specified. MRM data stripping determines what data are ultimately recorded and there are three options: “none,” “single isotope,” and “isotope cluster.” About 50 Da of m/z are enhanced around the ion of interest in the target enhancement phase of Tof-MRM and if fragment and/or isotope ions for monitoring are near the target mass, more than one HRMS peak may be quantitated using this transition. Selecting “none” collects the entire range (~50 Da), “isotope cluster” collects 5 Da (i.e. a region of isotopic peaks around the product ion that may also potentially be quantitated), and “single isotope” only records a 1 Da region centered on the ion of interest. Selectivity can be further increased by selecting a narrow HRMS XIC mass tolerance to isolate the HRMS signal from matrix noise; this is described in the processing section.
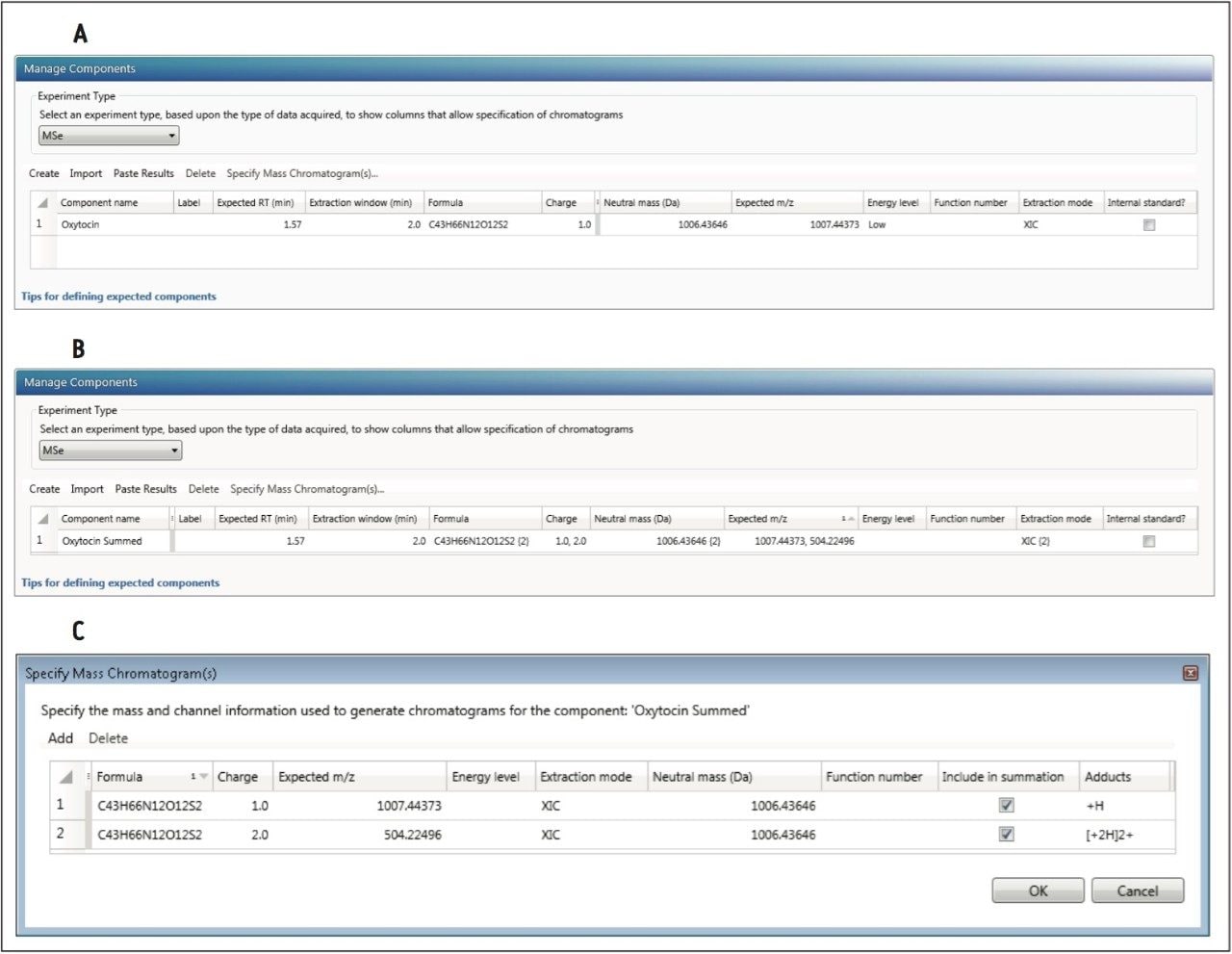
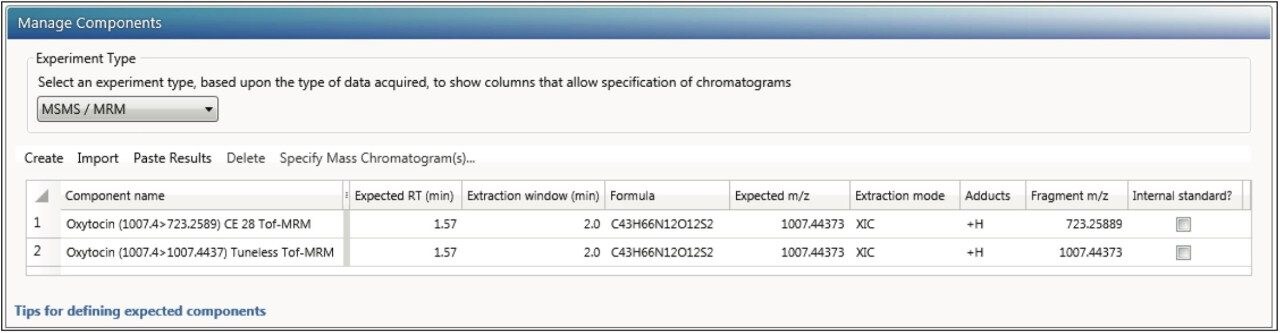
Once the method is defined, the processing settings need to be defined as shown in Figure 6. Specify the same values here as in the instrument method, so that processing generates the correct XICs for quantitation.
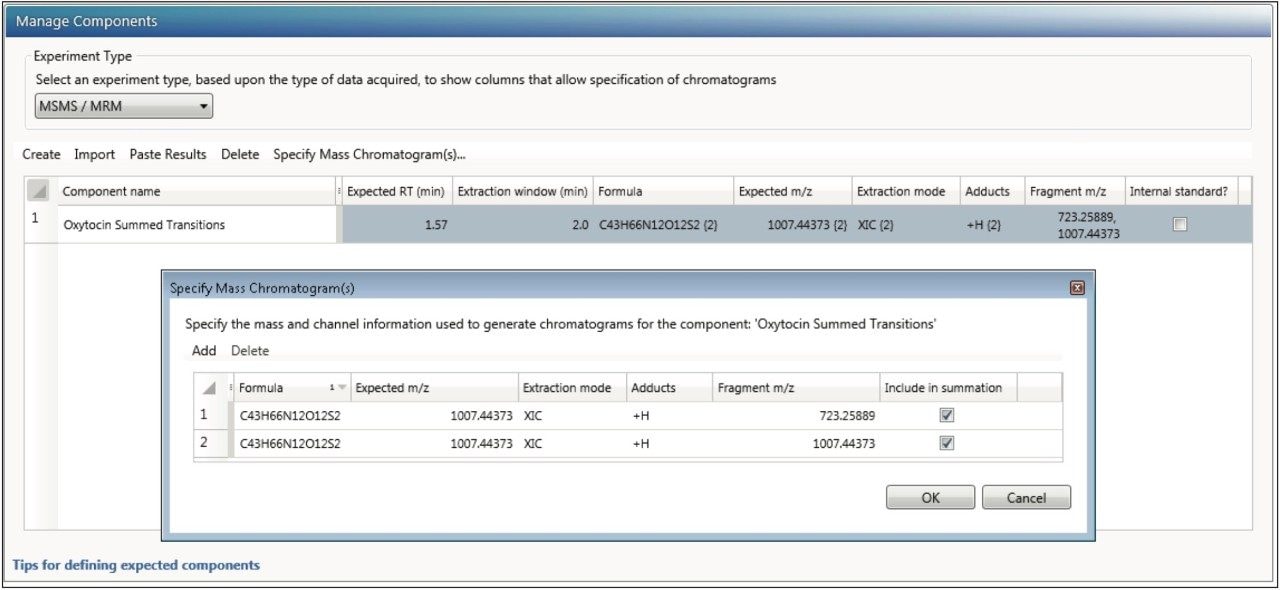
In order to create a summed transition, create a single component. Then select “Specify Mass Chromatogram(s)” (Figure 7). This opens a window in which the precursor (“Expected m/z”) and “Fragment m/z” must be specified. Traces may be summed for the final response in the final column by clicking the column labeled “Include in summation.” Once the information is entered press OK. The information is summarized in the “Manage Components” table.
Data can be collected at this point. The sample list for quantitation curve is set up and acquired. The next section details the steps required to process the collected data.
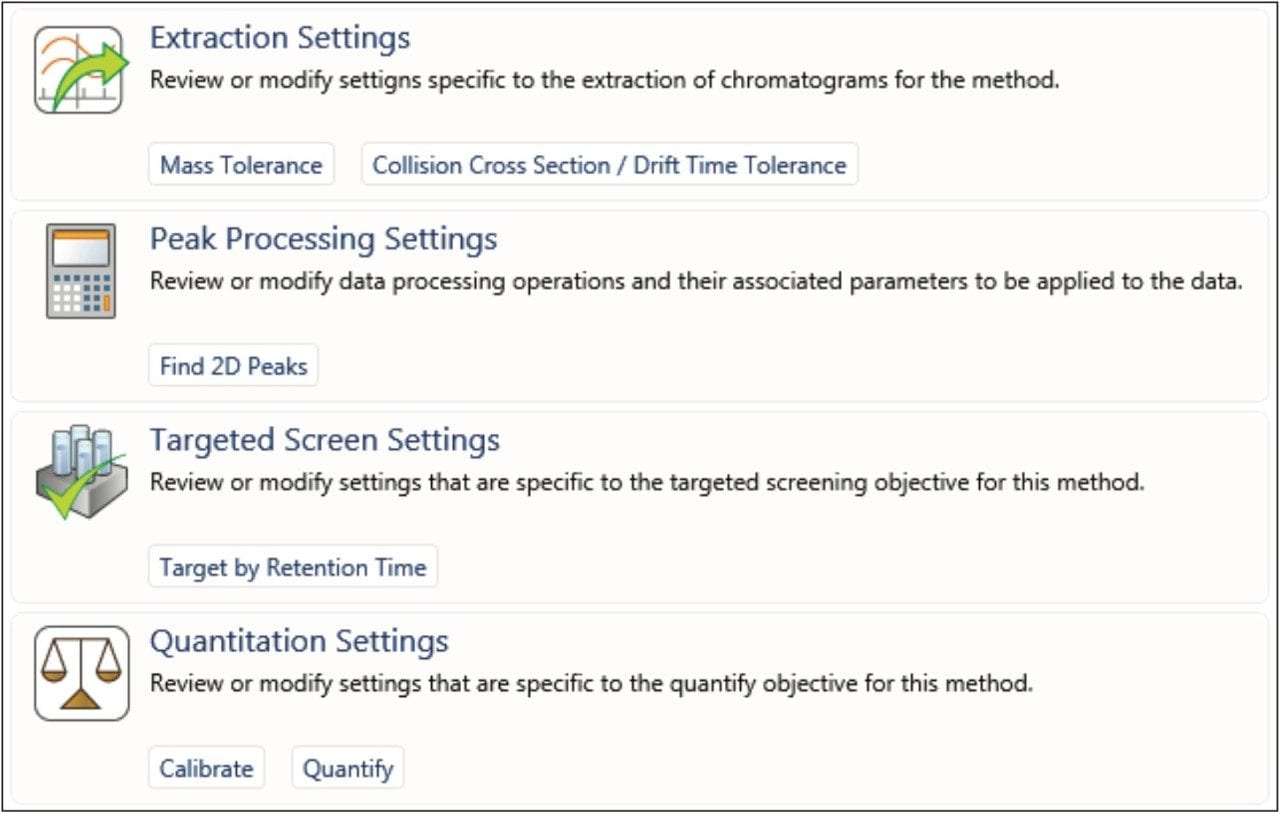
Once the data are acquired, many settings for processing are identical for all HRMS types, regardless of the acquisition format. Figure 8 shows the key parameters for processing data for HRMS quantitation; these include “Mass Tolerance,” “Find 2D Peaks,” “Target by Retention Time,” and “Quantitation Settings.” In most cases, the default settings are an appropriate starting point. Other advanced quantitative settings – such as those relating to chromatographic performance, how to address system suitability, and US and EU Pharmacopeia regulations – are not covered in this document, but have comprehensive help sections in UNIFI.
“Mass Tolerance” may be specified in Da, mDa, or ppm (default is 0.1 Da). For both full scan and Tof-MRM examples, an XIC mass tolerance of 50 mDa was used because it generated an XIC with optimal selectivity (Figure 9). Different XIC tolerances may be extracted for each component by using the “Specify Component” dialog in the navigation panel. All components not specified in the “Specify Components” section will be processed using the default value.
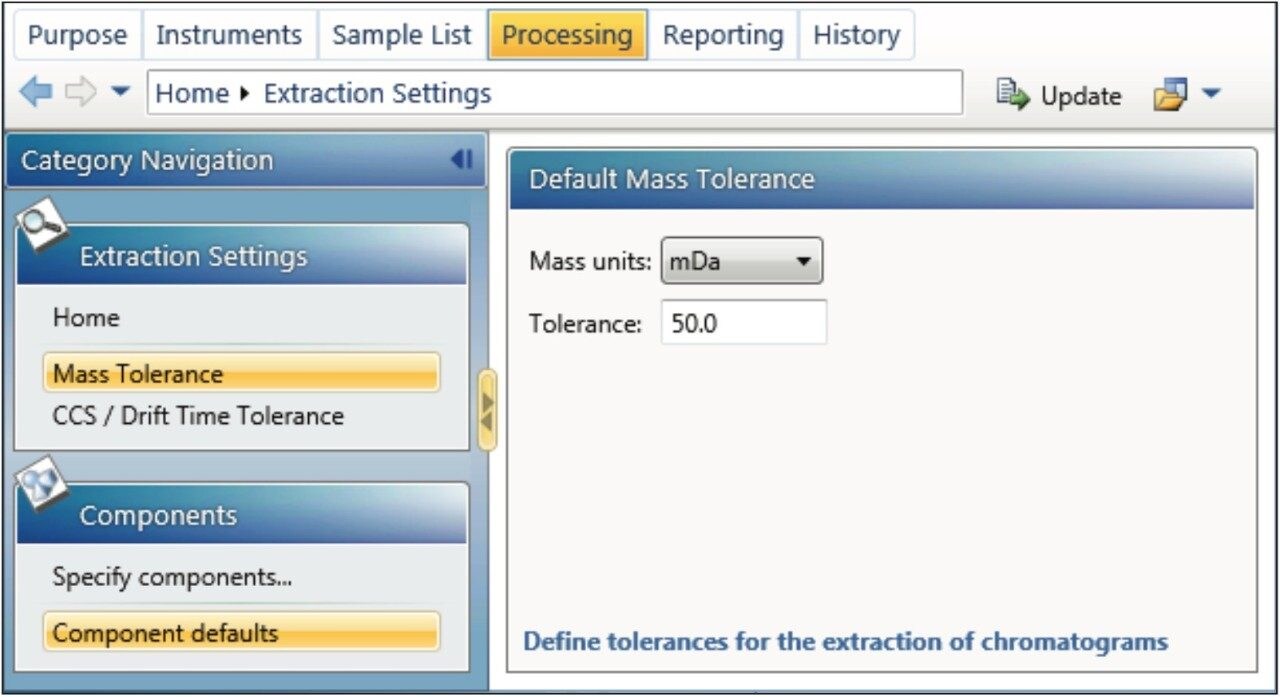
In this section, “CCS/Drift Time Tolerance” parameters may also be adjusted (dialog not shown). The value is specified as the percentage of the “Drift Time Range.” A value of 100% would use the entire DT / CCS range and is thus unfiltered data. Between 2% and 10% is generally a good starting point and removes any interfering matrix peaks that have the same mass (isobaric) but a different CCS to the targeted component of interest. A CCS value needs to be specified in the component table and may be obtained from a screening analysis.
“Peak Processing/Find 2D Peaks” settings can be modified if desired. Global integration settings may be adjusted or defined for the analysis. Alternatively, different integration parameters per component may be specified by using the “Specify components…” dialog.
“Targeted Screen Settings” determines how the peak is identified in the XIC. The default is “closest to RT” and picks the peak that has the closest RT specified in the component table (shown in Figure 4a-c; in this example 1.57 min). Other peak-picking rules are available (maximum peak area, maximum height, first peak, etc.) which may be useful for XICs that are more complex and contain multiple peaks within a very small time window. Different rules per component may be specified by using the specify component dialog.
Under “Quantitation Settings”, there are two tabs: “Calibrate” and “Quantify.” “Calibrate” is used to specify the standard curve options such as curve fit type, weight type, and units. In this example a linear 1/x fitting is used; the values are reported as a concentration (amount is also available) and the units are reported in pg/mL. These defaults may be changed as needed for the assay. Under the “Quantify” tab, the user may select whether the area (default) or peak height is to be used for quantitation.
After processing the data (in “Review,” select “Process, Full Application Processing”), the data is reviewable. Opening the “Quantitation Calibration” panel and selecting the component of interest enables data review. Clicking on the “Ignore” check box for a sample or right-clicking a data point (round circles) on the calibration plot allows the point to be removed. To change the integration of a peak, simply left click the peak start or stop mark and then adjust it to the desired position. The peak modifications are stored via the audit trail for use in highly regulated applications.
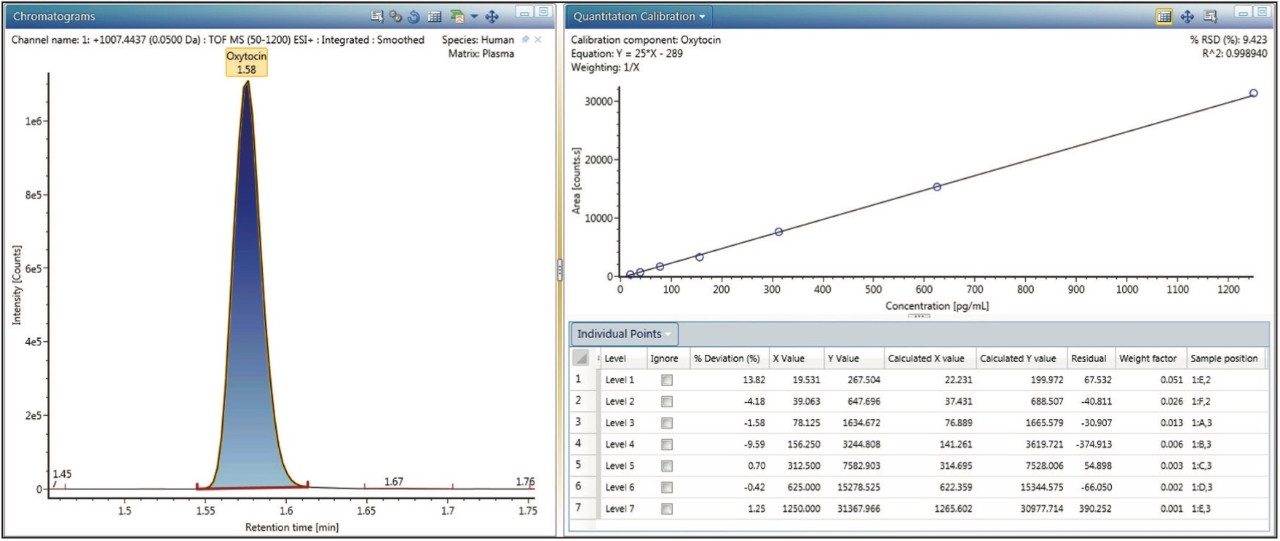
Data collected from other UNIFI analyses may be processed with the Quantify Assay Tof 2D Chromatographic Analysis method type. Any previously acquired data from UNIFI HRMS workflows (Biopharm, Accurate Mass Screening, Met ID Screening, Natural Products Screening, etc.) can be analyzed. It has been tested using imported MassLynx (continuum data, G2 series, and later mass spectrometers) files (MS,E MS/MS, Tof-MRM, HDMS, HDMSE types) and can support data acquired from many more HRMS quantitative platform configurations (including instruments configured with non-UNIFI native inlet configurations such as ACQUITY UPC2 and ionKey data).
HRMS data for quantitative applications are becoming more and more widely accepted. UNIFI software enables collection of multiple modes of data and flexible quantitation workflows. UNIFI enables users to develop, process, and analyze HRMS bioanalysis assays with workflows that fit traditional tandem discovery or regulated bioanalytical workflows.
This application note outlines the acquisition of either full scan data (MSE) or newer sensitive targeted Tof-MRM data types. It then outlines processing these data types for a quantitative assay. Both data types (along with most other MS and ion mobility data types) may be processed using the Quantify Assay Tof 2D Chromatographic Analysis method type.
Using a comprehensive toolset, analysts can build methods to acquire and process data to resolve matrix background from peaks of interest in complex samples. This is achieved using multiple modes of selectivity and HRMS filtering to provide sensitive and accurate results.
The new HRMS Tof quantitatation analysis method is fast, flexible, and compliant. It enables time-of-flight data to be collected and analyzed in a bioanalytical laboratory.
720005605, March 2016