This application note demonstrates accurate, reproducible quantification of endogenous CRP levels in plasma using a generic kit-based approach. The broad dynamic range and specificity of LC-MS reliably measures low endogenous levels and the elevated levels expected in diseased patients.
High sensitivity quantification of a protein biomarker, demonstrated speed and reproducibility of a generic kit-based approach for protein quantification, high sensitivity of the Xevo TQ-XS mass spectrometer, mixed-mode SPE specificity, no affinity purification required.
C-Reactive Protein (CRP)1 is naturally synthesized in the liver and released into the bloodstream in response to inflammation. There is, therefore, interest in measuring CRP both in cases of chronic inflammation such as Rheumatoid Arthritis (RA)2,4 as well as to evaluate cardiovascular risk3 and inflammatory cancerous processes.4 Tissue injury or inflammation causes a > 100-fold2 increase in plasma CRP levels. While endogenous plasma levels in healthy individuals are relatively low, generally between 0.1 and 3 µg/mL (4–120 nM),2 levels present in diseased patients are so highly elevated, multiple ELISA or immunoturbidimetric3 tests are required to achieve accurate CRP quantification. The need for multiple tests arises out of the limited linear dynamic range of ligand binding assays (LBA). This inherent short coming, as well as lack of standardization, possible cross-reactivity, and expensive, difficult to reproduce reagents are just a few of the reasons that the industry is moving towards LC-MS. Mass Spectrometry detection offers many benefits for protein quantification such as sensitivity, specificity, broad linear dynamic range, fast method development times, and the ability to multiplex. However, LC-MS protein quantification still presents challenges. There is no single standardized workflow and the various workflow options can be complex and laborious, making it difficult for a scientist to achieve success. In this application note, we describe a generic, kitted approach which requires only 2 hours for digestion vs. a standard 24 hour method described in the literature.5 Digestion and subsequent peptide level purification are performed using the ProteinWorks eXpress Direct Digest (p/n: 176003688) and ProteinWorks µElution SPE Clean-Up Kit (p/n: 186008304) for the accurate and reliable quantification of endogenous CRP in human plasma. The kits and their included protocols achieve LLOQs between 0.025-0.1 µg/mL (1-4 nM) from only 35 µL of plasma.
To prepare calibration curve standards and quality control (QC) samples, CRP (human sequence) was spiked into rat or human plasma at various concentrations over the range of 0.025–100 µg/mL (1–3987 nM). CRP QC samples were prepared in one lot of rat plasma and four lots of human plasma. Calibration curve standards were prepared in duplicate, each QC level prepared in triplicate, and blank (non-spiked) plasma samples were prepared in quadruplicate. Plasma samples (35 µL) were digested for 2 hours using the ProteinWorks eXpress Direct Digestion Kit, specifically, the 3-Step (no reduction/alkylation) method included in the kit with an abbreviated denaturation time of 5 minutes (p/n: 176003688). Post digestion purification of signature peptides was done using the ProteinWorks µElution SPE Clean-Up Kit (p/n: 186008304) and included protocol.
|
LC system: |
ACQUITY UPLC |
|
Column: |
ACQUITY UPLC HSS T3, 1.8 μm, 2.1 x 50 mm |
|
Temp.: |
55 °C |
|
Sample temp.: |
10 °C |
|
Injection volume: |
5 μL |
|
Mobile phases: |
A: 0.1% Formic acid in H2O B: 0.1% Formic acid in CAN |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
Curve |
|---|---|---|---|---|
|
Initial |
0.3 |
100 |
0 |
6 |
|
1 |
0.3 |
100 |
0 |
6 |
|
5 |
0.3 |
70 |
30 |
6 |
|
5.5 |
0.3 |
10 |
90 |
6 |
|
6.3 |
0.3 |
10 |
90 |
6 |
|
6.4 |
0.3 |
100 |
0 |
6 |
|
7.5 |
0.3 |
100 |
0 |
6 |
|
MS system: |
Xevo TQ-XS |
|
Ionization mode: |
ESI+ |
|
Capillary: |
3.5 kV |
|
Source offset (V): |
60 V |
|
Source temp.: |
150 °C |
|
Desolvation temp.: |
600 °C |
|
Cone gas flow: |
150 L/Hr |
|
Desolvation gas flow: |
1000 L/Hr |
|
Collision gas flow: |
0.15 mL/Min |
|
Nebulizer gas flow: |
7 Bar |
|
Data management: |
MassLynx (v4.1); TargeLynx |
Early detection of elevated CRP levels could be imperative for proper treatment or prevention of RA or heart disease. Availability of a sensitive analytical method that can detect low levels and differentiate between small concentrations can facilitate early detection. Mass spectrometric detection, though not traditionally used for CRP quantification, has the sensitivity and reproducibility required for measuring both the low endogenous CRP levels in healthy individuals as well as elevated levels in diseased populations.
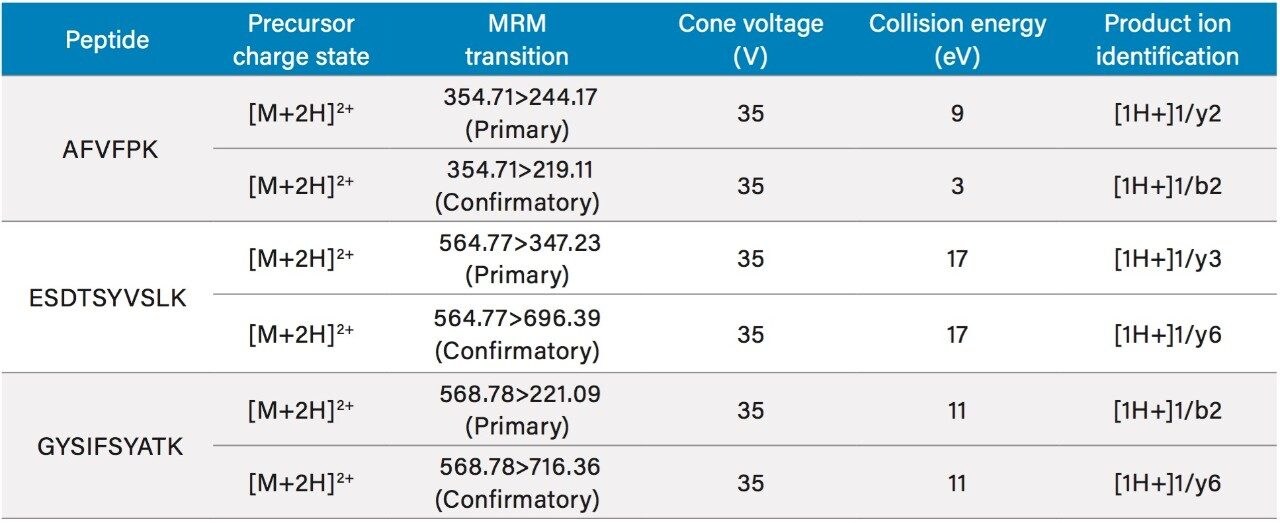
Protein quantification by LC-MS is typically performed via the bottom up approach which uses enzymatic digestion (usually trypsin) and relies on analysis of resulting (tryptic) peptides. To identify an appropriate representative or surrogate peptide for human CRP (Figure 1), an in-silico digestion was performed using Skyline Software (MacCoss Labs, University of Washington).6 In addition to identifying potential tryptic peptides, Skyline facilitated MS method development by selecting and optimizing precursors and collision induced fragments for MRM (multiple reaction monitoring) analysis. The full amino acid sequence of CRP7 and the unique signature peptides: AFVPFK, ESDTSYVSLK, and GYSIFSYATK (highlighted in blue), are illustrated in Figure 2. Optimized MS conditions and MRM transitions for the CRP tryptic peptides are listed in Table 1. Both primary and confirmatory (secondary) transitions are included for each peptide.

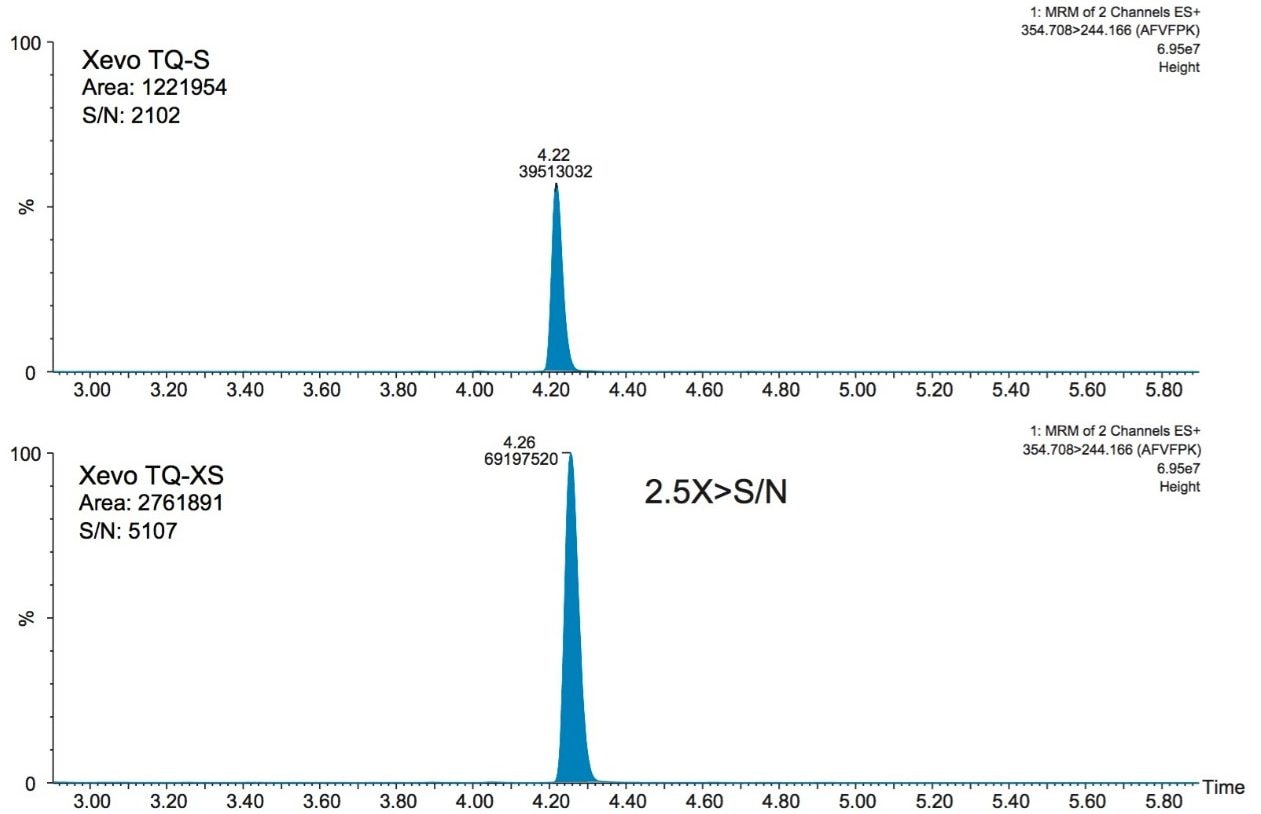
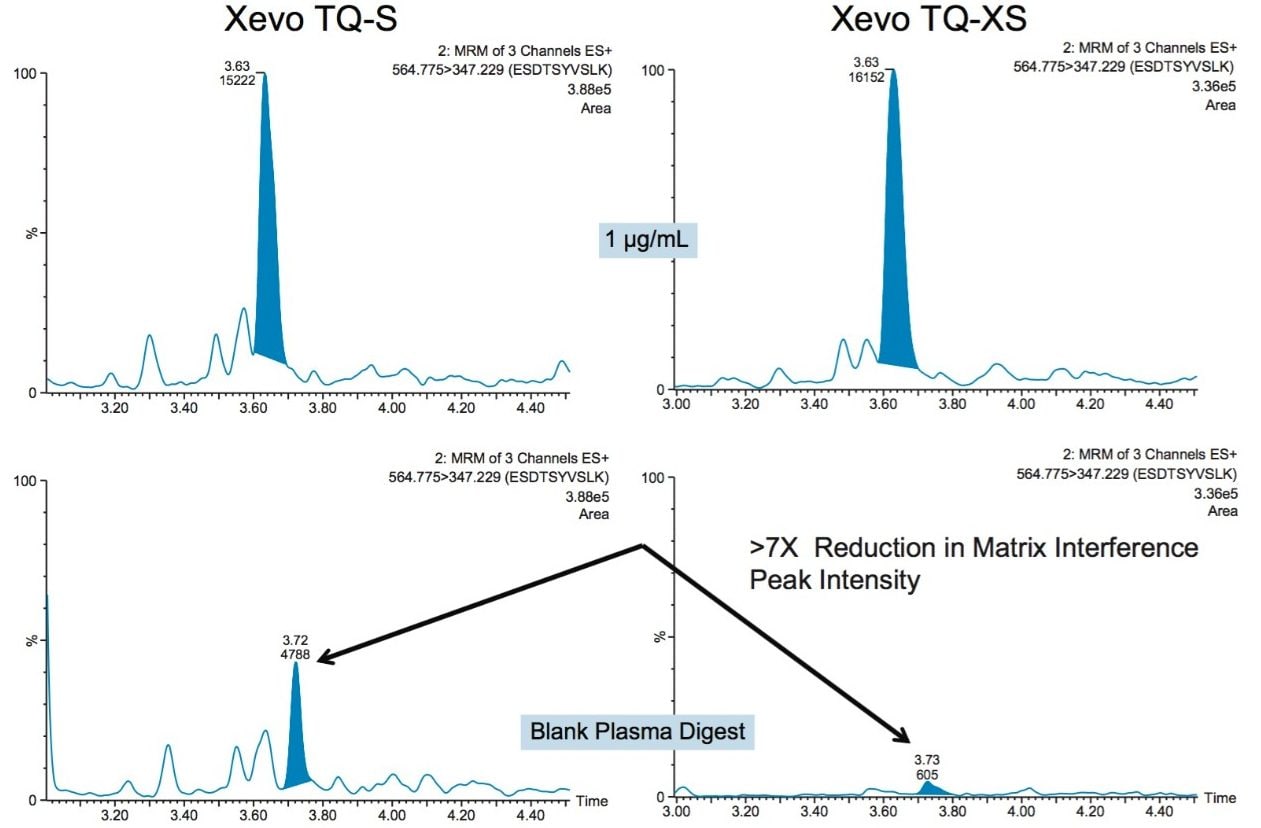
In an earlier study, CRP quantification was performed on a Xevo TQ-S MS. Quantification was subsequently transferred to a newer generation platform, a Xevo TQ-XS triple quadrupole mass spectrometer, which provided the benefit of improved sensitivity. Signal-to-noise (S/N) for the three CRP peptides used for quantification increased 2.5 X (AFV), 3.5 X (ESD) and 1.4 X (GYS) using the Xevo TQ-XS vs the older platform. This improvement in sensitivity for the AFV peptide is demonstrated in Figure 3. For the ESD, peptide (RT 3.63 minutes), the intensity for an interfering matrix peak (RT 3.72 minutes), present as a shoulder on the main peak, was decreased 7 X. This is shown in Figure 4. The reduced matrix interference facilitated easier peak integration and resulted in improved limits of detection.
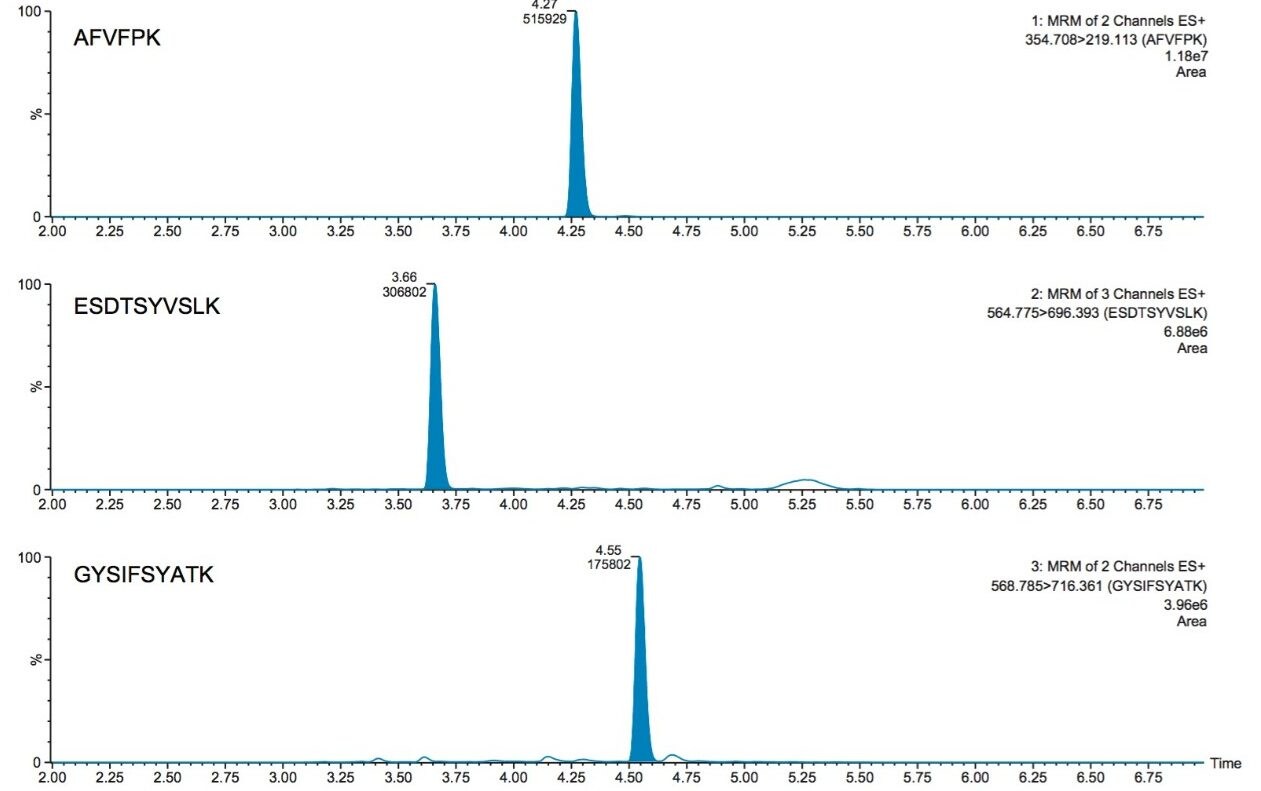
Reversed-phase chromatographic retention can be challenging due to the relatively small size and polar nature of tryptic peptides. Use of an ACQUITY HSS T3 Column, 1.8 µm, 2.1 x 50 mm (p/n: 186003538) for the CRP peptides afforded improved retention as compared to a BEH C18 column and facilitated resolution from endogenous matrix interferences. Representative chromatograms of the three peptides: AFVPFK, ESDTSYVSLK, and GYSIFSYATK are illustrated in Figure 5. Peak widths were <4.5 seconds wide.
Typical sample preparation workflows for protein quantification are often complex and laborious. In this application, we used the ProteinWorks eXpress Direct Digest Kit (p/n: 176003688) to directly digest plasma samples in only 2 hours. Subsequent purification of the peptides with the ProteinWorks µElution SPE Clean-Up Kit (p/n: 186008304) and protocol removed buffer salts, phospholipids, and excess digestion reagents post digestion. The SPE kit relies on a mixed-mode sorbent (Oasis MCX, reversed-phase and ion-exchange) to provide enhanced specificity. Tryptic peptides are bound to the ion-exchange moiety of the SPE sorbent, thereby imparting orthogonality, and thus greater specificity, into the method as a whole. In addition, very polar peptides are more efficiently trapped by ion exchange than by traditional reversed-phased only sorbent. This was particularly important for the most polar peptide, ESD. Recoveries for all three CRP peptides were excellent, with greater than ≥90% recovery using the generic protocol provided.
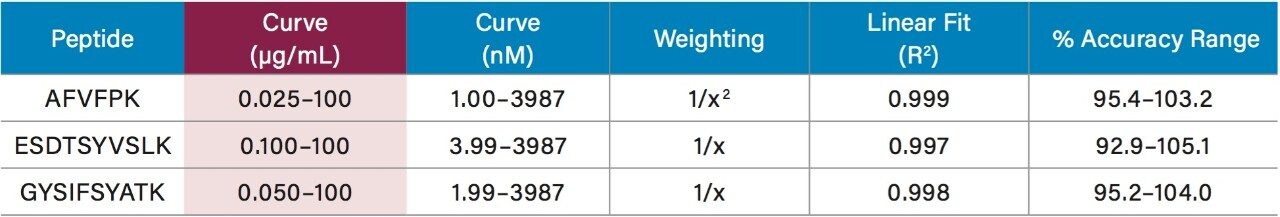
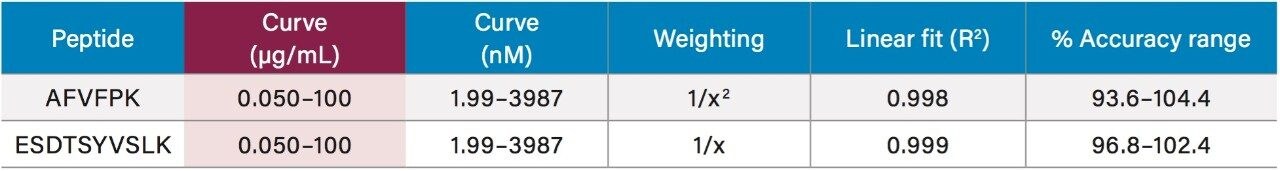
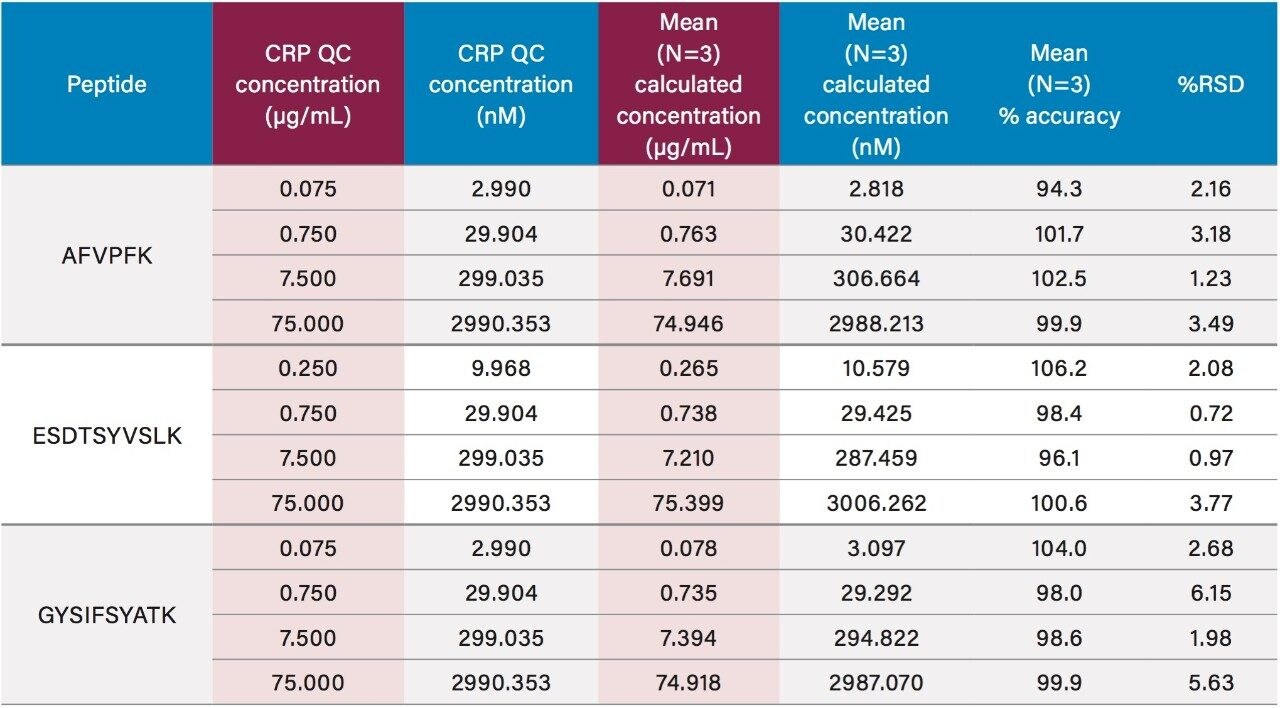
Using only 35 µL of sample and the aforementioned ProteinWorks kits, quantification limits of 0.025 and 0.05 µg/mL (1–2 nM) were achieved in rat and human plasma, respectively. Calibration curves from the tryptic peptides, in both rat and human plasma, were linear with R2 values >0.99 using 1/x or 1/x2 weighted regressions. A summary of standard curve performance in rat and human plasma is shown in Tables 2 and 3. Linearity, accuracy and precision data met typical method validation requirements. For both rat and human, standard curves were linear over 4 orders of magnitude with mean accuracies ranging from 94–105%. The precision and accuracy for the QC samples was excellent with mean % RSDs all <5% and % QC accuracy ranges of 94.3–106.2 (rat) and 89.4–103.5 (human). Rat QC performance is highlighted in Table 4. Human QC performance for the AFV and ESD peptides is shown in Table 5, Panels A and B, respectively. Demonstration of this performance is illustrated in Figure 6, Panels A and B.
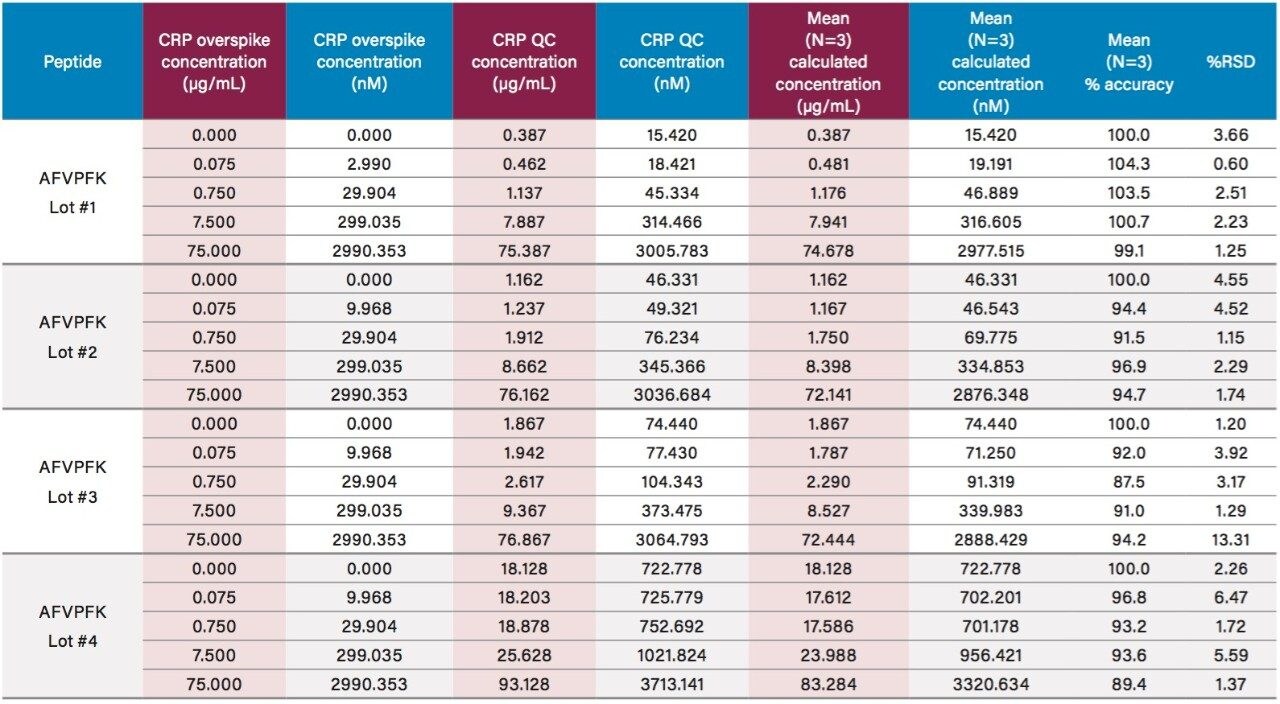
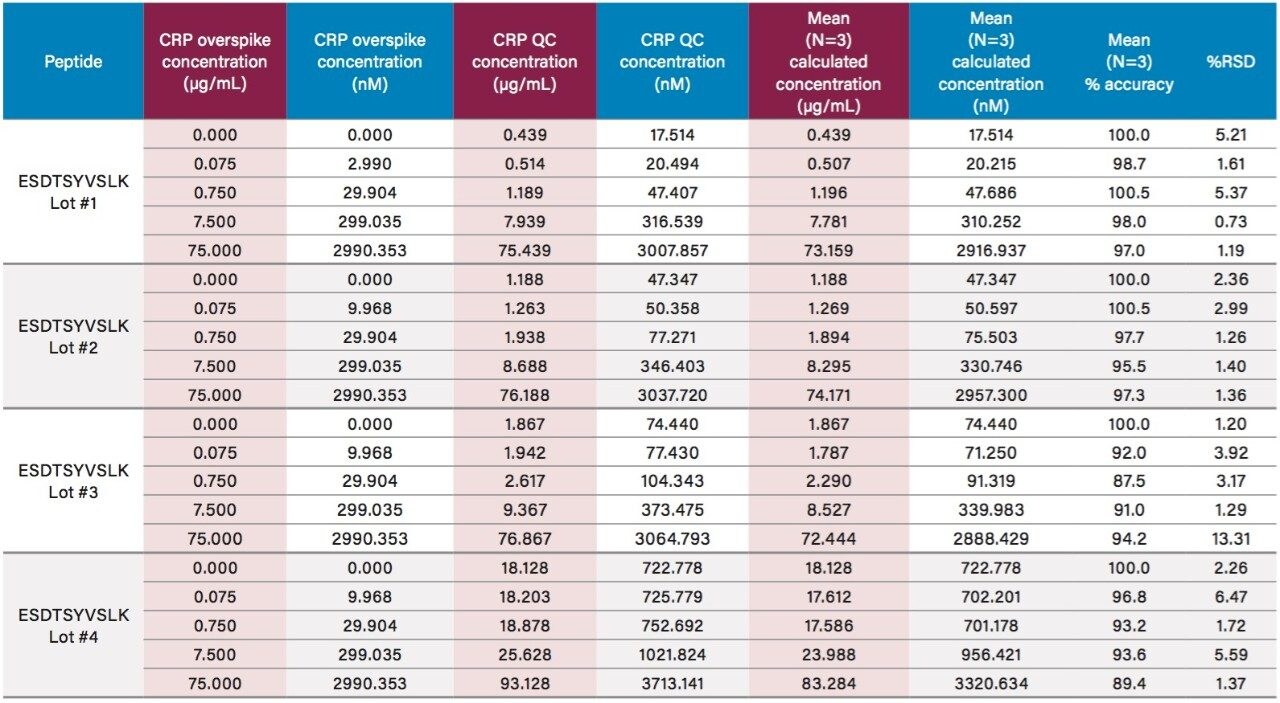
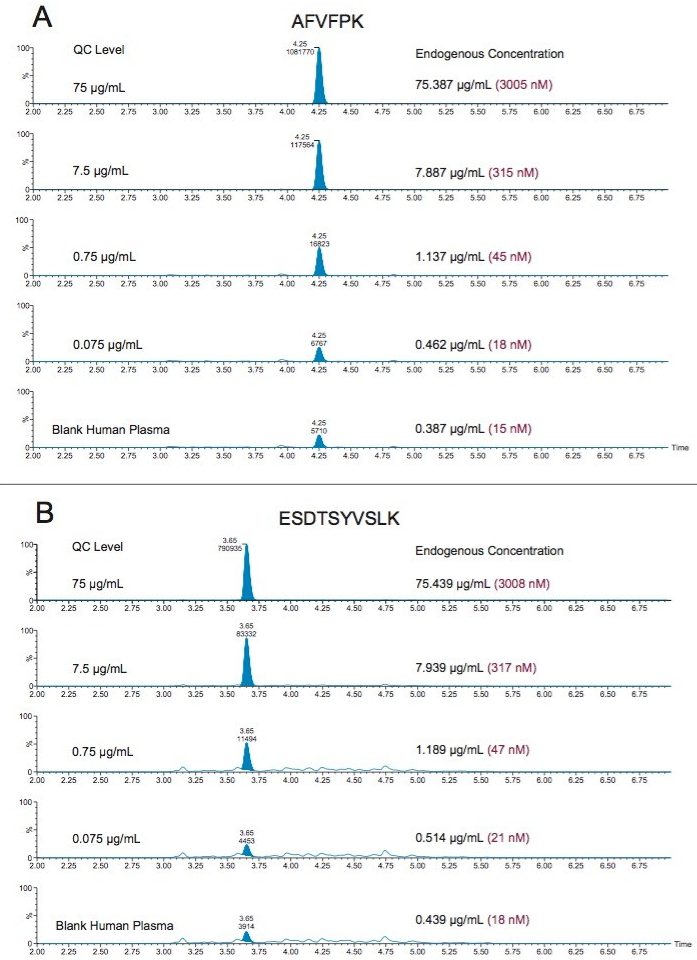
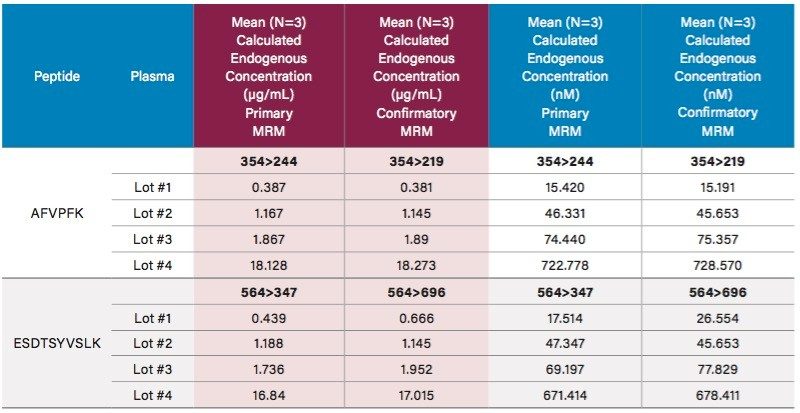
Endogenous human CRP concentrations were accurately quantified in a total of four lots of human plasma. The lowest level of CRP that could be accurately and precisely quantified above the endogenous level in human plasma was 0.05 µg/mL (2 nM). For the blank human plasma digest (Figure 6), there is a strong signal from the human CRP peptide, due to endogenous CRP concentrations in the range of 0.4–0.666 µg/mL. In rat plasma blank digest (Figure 4), human CRP peptide is not detected. This is expected as the rat CRP amino acid sequence is different than human CRP sequence. Calculated endogenous CRP levels for the four lots of human plasma are summarized Table 6 and illustrated in Figure 7, panels A (AFV peptide) and B (ESD peptide). Within each plasma lot, the calculated endogenous CRP concentrations derived from either the AFV or ESD tryptic peptides were within 10% agreement. Confirmatory transitions for each peptide were used to verify endogenous concentrations of each plasma lot. With the exception of Lot 1, the concentrations calculated from the primary and confirmatory transitions were within 10% of each other. Endogenous CRP concentrations calculated from the GYS peptide did not correlate with those calculated by the AFV and ESD peptides. It is speculated that an underlying, co-eluting interference impeded accurate quantification using the GYS peptide.
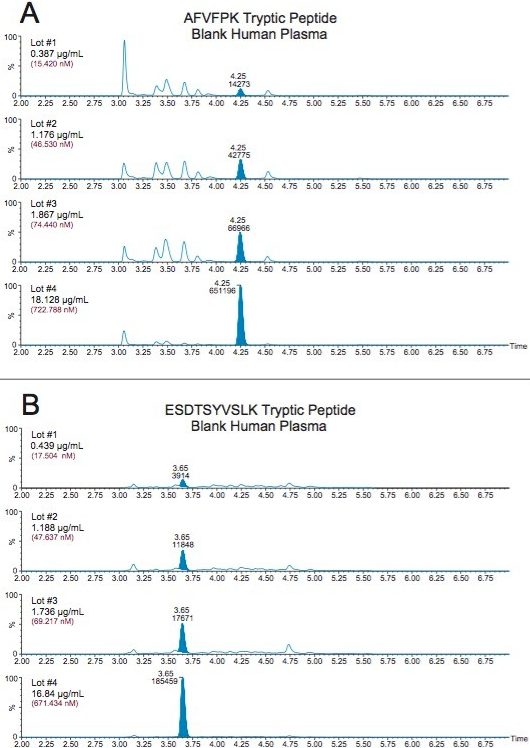
Endogenous CRP was reliably quantified down to 0.025 µg/mL (1 nM) using commercially available digestion and purification kits. Plasma (35 µL) was directly digested (no affinity purification necessary) and subsequent tryptic peptides purified using the generic protocols provided in the kits. The combination of mixed-mode SPE and protocol resulted in >90% recovery for the three peptides. Total sample preparation time, including SPE, was <3 hours. Updating the MS platform, Xevo TQ-S to Xevo TQ-XS, improved sensitivity and S/N, consequently improved the LOQ. This method demonstrates the accurate, reproducible quantification of endogenous CRP levels in plasma using a generic kit-based approach. The broad dynamic range and specificity of LC-MS reliably measures low endogenous levels and the elevated levels expected in diseased patients.
720005819, October 2016