
In this application note, we describe a rapid, multi-residue method for the determination of over 200 pesticides in basmati rice using a simple QuEChERS sample preparation procedure, followed by UltraPerformance LC (UPLC) coupled to tandem quadrupole mass spectrometry. The LC-MS/MS method presented in this study can also be employed for multi-analyte screening and quantification, providing a single method for more cost-effective analysis of pesticides in rice.
Rice is one of the most widely consumed foods in the world and its demand has increased in recent decades. Basmati rice is a variety of long grain rice which is traditionally cultivated in northern part of India. To improve its production yield, the use of various pesticides in various stages of cultivation has increased. Due to the adverse effects of these pesticides on human health and to the environment, the use of pesticides must be controlled and monitored.
According to the Horizon Scan database,1 managed by the UK Food and Environment Research Agency, the most commonly found pesticides in rice exported from India between 2011 to 2014 were acephate, buprofezin, carbendazim, imidacloprid, isoprothiolane, methamidophos, pirimiphos-methyl, triazophos, tricyclazole, malathion and propiconazole. These pesticides were detected above the reporting limit.
A multi-residue pesticide method has been developed for the determination of pesticides in basmati rice. In this work, over 200 pesticides were analyzed using a simple QuEChERS sample preparation procedure, followed by UltraPerformance LC (UPLC) coupled to tandem quadrupole mass spectrometry.
|
UPLC system: |
ACQUITY UPLC H-Class |
|
Column: |
ACQUITY UPLC BEH C18, 1.7 μm, 2.1 x 100 mm |
|
Column temp.: |
45 °C |
|
Injection volume: |
10 μL |
|
Flow rate: |
0.45 mL/min |
|
Mobile phase A: |
10 mM ammonium acetate (pH 5) in water |
|
Mobile phase B: |
10 mM ammonium acetate (pH 5) in methanol |
|
Weak needle wash: |
50/50 water/methanol (v/v) |
|
Strong needle wash: |
90/10 methanol/water (v/v) |
|
Seal wash: |
90/10 water/methanol |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
Curve |
|---|---|---|---|---|
|
Initial |
0.45 |
98.0 |
2.0 |
6.0 |
|
0.25 |
0.45 |
98.0 |
2.0 |
6.0 |
|
12.25 |
0.45 |
1.0 |
99.0 |
6.0 |
|
13.0 |
0.45 |
1.0 |
99.0 |
6.0 |
|
13.01 |
0.45 |
98.0 |
2.0 |
6.0 |
|
17.0 |
0.45 |
98.0 |
2.0 |
6.0 |
|
MS system: |
Xevo TQD |
|
Ionization mode: |
ESI+ |
|
Capillary voltage: |
1 kV |
|
Desolvation temp.: |
500 °C |
|
Desolvation gas flow: |
1000 L/hr |
|
Source temp.: |
150 °C |
A mix of 204 pesticides was prepared from Waters LC Multi-residue Pesticide Standards Kit, p/n: 186007574. The additional two pesticides Pirimiphos-methyl and triazophos were purchased from Sigma Aldrich. A mixture of all pesticides at 5 μg/mL was prepared in acetonitrile.
A variety of rice samples was purchased from local supermarkets. Jasmine rice (Sample A), three types of basmati rice (Samples B, C, and E), and organic brown rice (Sample D) were analyzed in this study.
The sample preparation protocol used was slightly modified from a previously published protocol by Lucia Pareja et al..2
Briefly, rice samples were ground to a fine powder using a food processor and 7.5 grams were weighed in a 50-mL centrifuge tube. The rice samples were then homogenized in 7.5 mL of water. The tube was shaken for 30 minutes using hand-motion shaker (Eberbach corporation, EL680.Q). Subsequently 15 mL of 1% glacial acetic acid in acetonitrile was added as an extraction solvent, followed by the addition of QuEChERS AOAC3 material (Waters DisQuE QuEChERS pouches, p/n: 186006812). The tube was shaken for 4 minutes on a motion shaker and centrifuged (Eppendorf, 5810 R) at 3700 RPM for 5 minutes. A 4-mL aliquot of the supernatant was removed and dried under a stream of nitrogen. The dried extract was then redissolved in 2 mL of 50:50 water:acetonitrile and filtered through an Acrodisc Minispike Syringe (0.45 μm) PTFE Filter (WAT200559) before LC-MS/MS analysis.
To study linearity, solvent and matrix matched standards (MMS) calibration curves were created by spiking the pesticide mix from 0.00125 mg/kg to 0.32 mg/kg (1.25 ppb to 320 ppb) in solvent and the rice sample respectively.
For the standard addition work, six test portions of ground Basmati rice (Sample B) were spiked with the pesticide mix from 0.00125 mg/kg to 0.32 mg/kg. Each of the pre-spiked samples were then extracted as described above and analyzed to create a standard addition curve.
All of the pesticides were successfully detected at or below their EU maximum residue level (MRL) in the spiked rice sample. According to the EU legislation, the MRL level for the most commonly found pesticides in rice exported from India were between 0.01 to 8 mg/kg.
The vast majority of the pesticides (~85%) showed linearity with R2 values >0.99. The remaining pesticides showed R2 values above 0.98 in both the matrix-matched and solvent calibration curves. Examples of two pesticides in both solvent and matrix are shown in Figure 1.

To assess the method recovery, rice samples were spiked at 0.01 mg/kg and extracted according to the described protocol. The samples were spiked in triplicate and each extract was analyzed in triplicate (total = nine injections). The recoveries of all of the pesticides together, along with standard deviations are shown in Figure 2.
As shown in Figure 2, the recoveries for most of the pesticides (92%) fell within the acceptable tolerance of 70% to 120% range (SANCO/12571/2014). Only 5% showed recoveries between 50% and 70%, and another 3% showed recoveries between 25% and 50%. The relative standard deviations (RSDs) were less than 20% except for nine compounds which had RSDs higher than 20%.

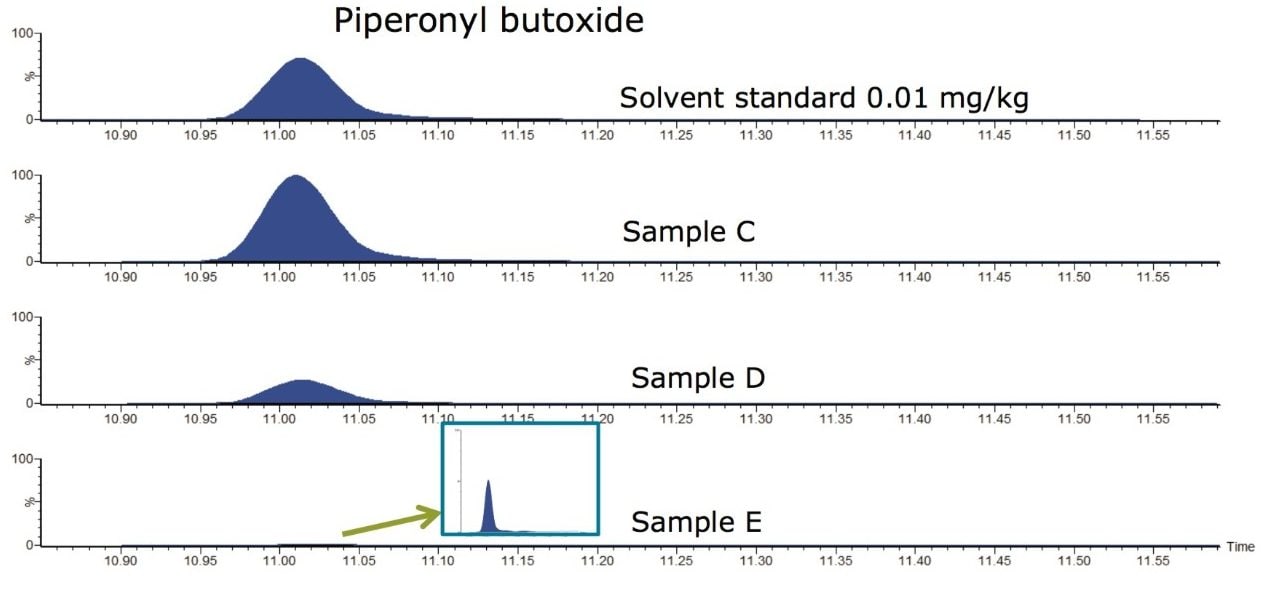
For the remaining samples, quantification was performed against a matrix-matched calibration curve. Piperonyl butoxide was detected in Sample C (0.0236 mg/kg) and Sample D (0.0054 mg/kg). Other pesticides were detected at concentrations <LOQ (e.g. imidacloprid in all samples, carbendazim in Sample C and Sample D, and piperonyl butoxide in Sample E. Figure 3 shows the MRM chromatogram for piperonyl butoxide in Samples C, D, and E, along with the solvent standard at 0.01 mg/kg.
Sample D was labeled as organic brown rice and was found to have low but detectable levels of two pesticides. The USDA National Organic Program4 includes a list of synthetic compounds that may or may not be used in organic crop production. This list does not include piperonyl butoxide or imidacloprid. The pesticides detected however were not at concentrations that exceeded the U.S. Food and Drug Administration (FDA) or the Environmental Protection Agency (EPA) regulatory tolerances and as such, no action level has been exceeded.
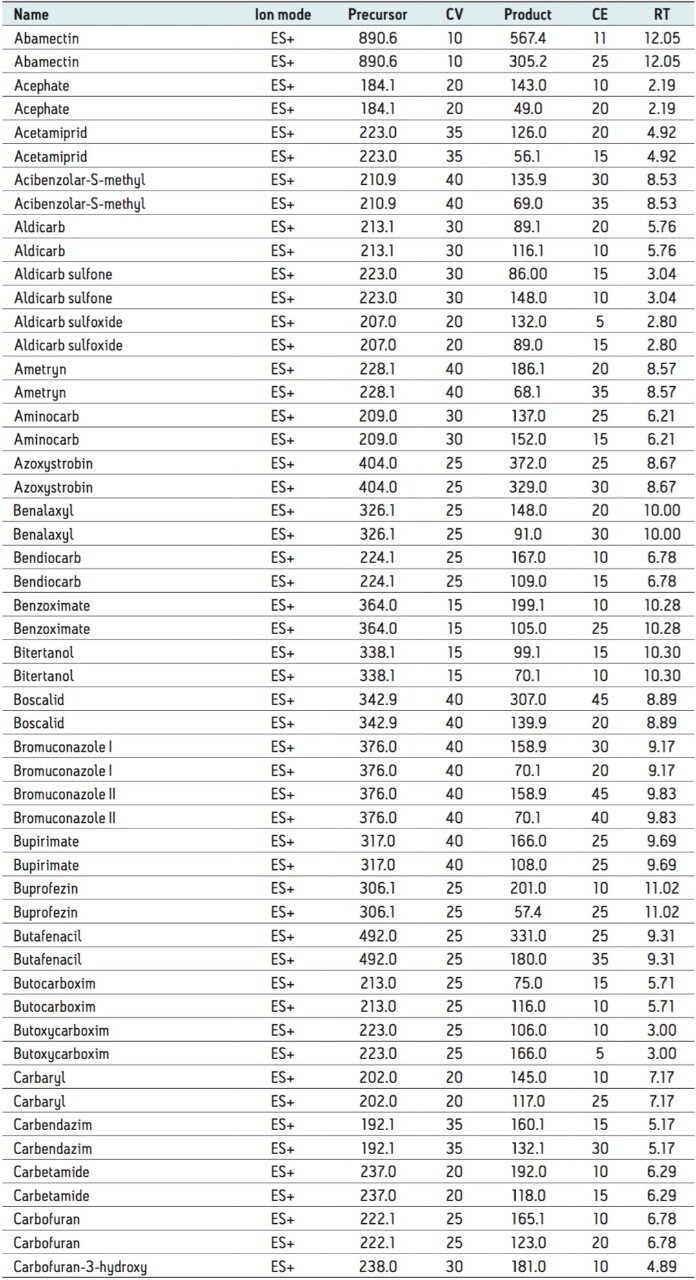
All of the incurred pesticide residues detected were identified from comparison of the retention time and ion ratio data with that from reference standards and found to be within the tolerance limit specified by the European Commission (Document No. SANCO/12571/2014).
A total of five rice samples were extracted and analyzed using the described method. One of the basmati rice samples (Sample B) was found to contain seven different pesticides. For accurate quantification of this sample, a standard addition approach was taken. The incurred residues were then automatically quantified using TargetLynx. Figure 4 shows the quantification of buprofezin using the new standard addition feature in TargetLynx.
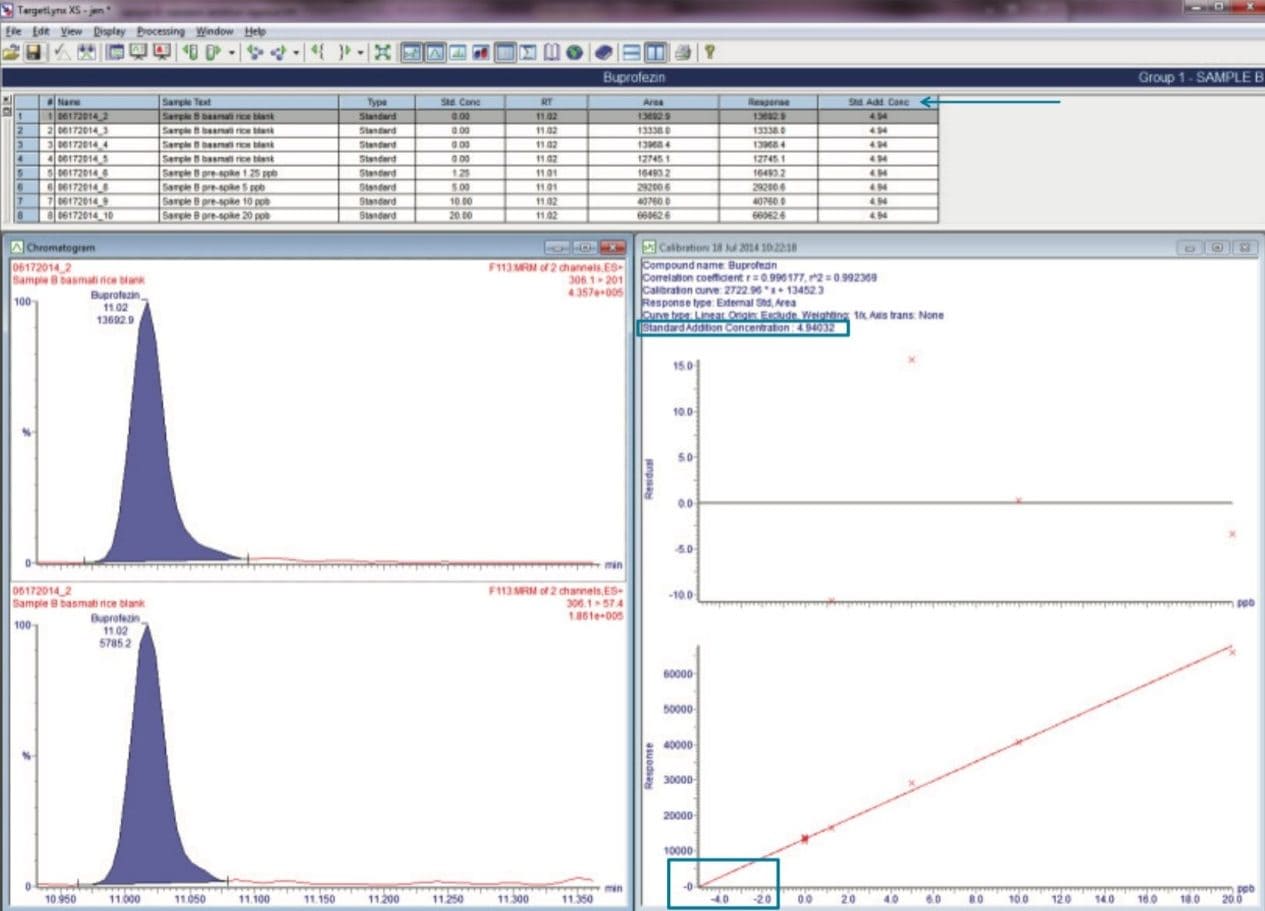
Using the standard addition method, seven pesticides were determined in basmati rice (Sample B) four of which were at concentrations greater than the lowest calibration point (0.00125 mg/kg); tricyclazole (0.0113 mg/kg ), propiconazole (0.0099 mg/kg), buprofezin (0.005 mg/kg), and triazophos (0.0015 mg/kg). Acetamiprid, carbendazim, and imidacloprid were detected at concentrations below the lowest calibration point (0.00125 mg/kg). The detection of these compounds corresponds to those that have been listed in the Horizon Scan database. All the pesticides detected in basmati rice Sample B were at concentrations that were below the European MRLs.
All of the pesticides of interest were successfully analyzed using a simple QuEChERS sample preparation procedure and UPLC-MS/MS determination in rice samples.
720005295, January 2015