For research use only. Not for use in diagnostic procedures.
This application note describes a HILIC-based UPLC method that provides better separation of polar lipid classes than existing reversed-phase-based methods, and is faster than previous normal-phase HPLC methods.
Lipids play an important role in the energy storage, cellular signaling, and pathophysiology of diseases, such as cancer, neurodegenerative diseases, infections, diabetes, etc. Advances in LC-MS have allowed lipids to be studied with greater sensitivity and specificity. This alleviates the effects of co-eluting compounds and isobaric interference, and allows low abundance lipids to be more readily detected.1
Conventional mass spectrometric analysis of lipids is often performed by direct infusion and reversed-phase or normal-phase HPLC.2-4 However, both normal-phase and reversed-phase methods have certain disadvantages. Although reversed-phase-based separation methods provide good separation of lipids, the complete separation of lipids by class has not been shown.4,5 This is due to the fact that the mechanism of action in reversed-phase chromatography of lipids is based on their lipophilicity, which is governed by the carbon chain length and the number of double bonds.6 As a consequence, co-elution of lipids belonging to different classes in reversed-phase separations is quite common. Normal-phase methods typically allow separation and characterization of different lipid classes, but their lengthy elution and equilibration times make them very time consuming. Moreover the mobile phases typically used for normal-phase – such as chloroform or hexane – lack compatibility with UPLC systems, are difficult to handle due to their volatility and toxicity, and prove challenging for ionization and introduction into a mass spectrometer.7
Although HILIC is considered a variant of normal-phase chromatography, a reversed-phase solvent system (organic-aqueous) can be used to avoid these potential issues. Indeed, due to the highly organic mobile phase (> 80% acetonitrile) used in HILIC, electrospray ionization (ESI) may be improved through more efficient mobile phase desolvation and compound ionization. Performing HILIC using 1.7 μm unbonded BEH particles provides all the acknowledged advantages of UPLC – faster methods, with enhanced chromatographic resolution, sensitivity, and reproducibility of separation.
This application note describes a HILIC-based UPLC method that provides better separation of polar lipid classes than existing reversed-phase-based methods, and is faster than previous normal-phase HPLC methods.
Lipids were extracted from monkey plasma by adding 900 μL of methanol to 100 μL of plasma. The mixture was incubated for 15 minutes prior to centrifugation and 900 μL of the supernatant was collected. A second extraction was carried out following the addition of a further 500 μL of methanol, and the upper 470 μL of the supernatant was combined with the first extract. The extracts were dried and re-suspended in 120 μL of 1:1 chloroform/methanol. This was diluted 10 times with acetonitrile, and 3 μL injected for each analysis.
|
LC system: |
ACQUITY UPLC |
|
Column: |
ACQUITY BEH HILIC 1.7 μm, 2.1 X 100 mm |
|
Column temp.: |
30 °C |
|
Flow rate: |
500 μL/min |
|
Mobile phase A: |
Acetonitrile/water (95:5) with 10 mM ammonium acetate, pH 8.0 |
|
Mobile phase B: |
Acetonitrile/water (50:50) with 10 mM ammonium acetate, pH 8.0 |
|
Gradient: |
Linear, 100 to 80% A in 10 min |
|
MS system: |
Xevo QTof MS |
|
Ionization mode: |
ESI +/- |
|
Capillary voltage: |
2800 V (+)/1900 V (-) |
|
Cone voltage: |
35 V |
|
Desolvation temp.: |
500 °C |
|
Desolvation gas: |
1000 L/Hr |
|
Source temp.: |
120 °C |
|
Acquisition range: |
100 to 1200 m/z |
Identification of lipids is based on precursor masses and the corresponding fragment or neutral loss characteristic of each group of lipids. However, some groups, such as phosphatidylcholines (PC) and sphingomyelins (SM), not only produce characteristic fragments of the same mass (m/z 184.0738), they are only separated by 1 Da; hence their isotopes interfere with the detection and quantification of one another. This is a problem where the different lipid classes are not separated, such as by direct infusion and to an extent with reversed-phase chromatography, where the groups overlap. Class-distinct separation can be achieved by normal-phase HPLC methods, but typically require about an hour for efficient separation.8 A ‘normal-phase’-like UPLC-MS method provides effective separation of different lipid classes in a much shorter time scale (about 16 minutes).
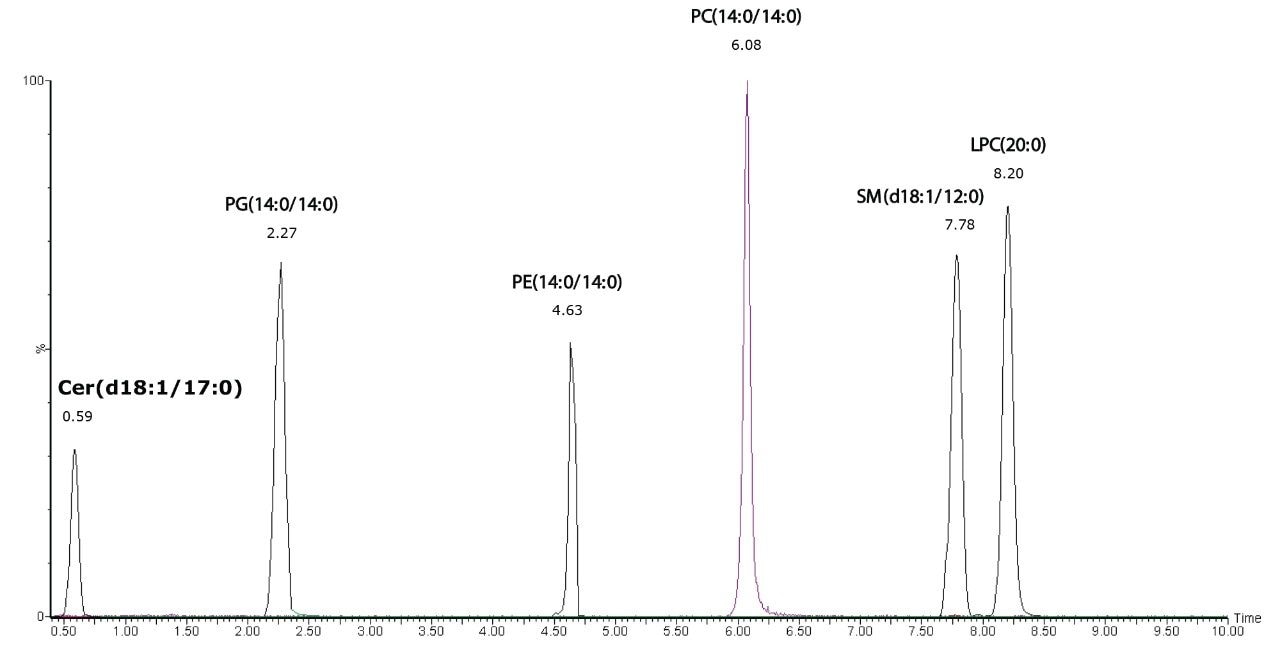
A mixture of lipid standards representing different lipid classes:
N-(heptadecanoyl)-sphing-4-enine (Cer(d18:1/17:0)),
dimyristoyl-phosphatidylglycerol (PG(14:0/14:0)),
dimyristoyl-phosphoethanolamine (PE(14:0/14:0)),
dimyristoyl-phosphatidylcholine (PC(14:0/14:0)), lauroyl sphingomyeline (SM(d18:1/12:0)), and 1-eicosanoyl-sn-glycero-3-phosphocholine (LPC(20:0)) were used to demonstrate the separation of the lipid classes, as shown in Figure 1. Extracted ion chromatograms of the characteristic fragment ions in the elevated energy MSE data provided confirmation of the lipid classes.
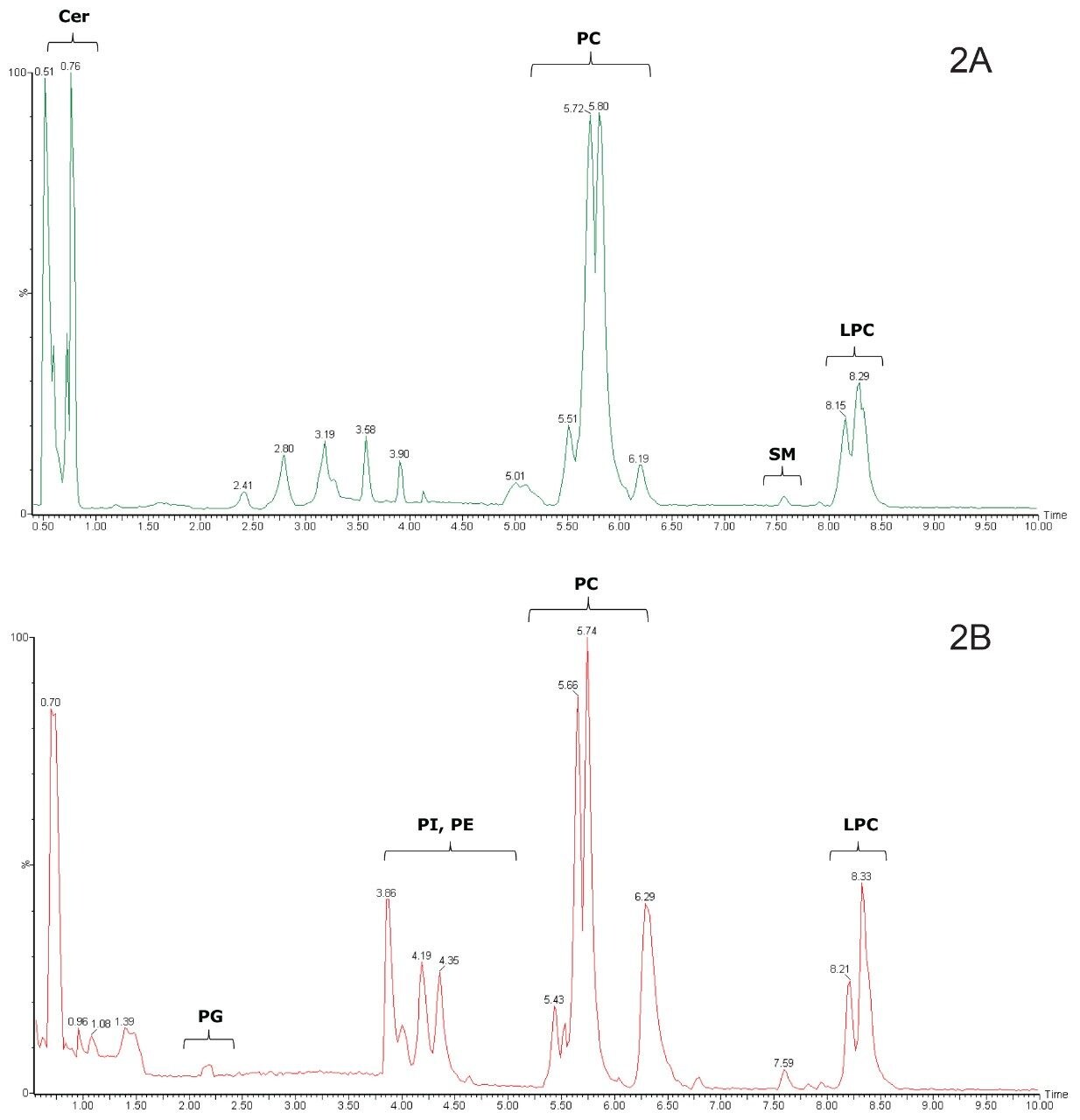
Although the developed HILIC method resulted in a much narrower spread of peaks within each class of lipids as compared to the reversed-phase UPLC method,9 the separation between different classes was greatly increased, as shown in Figure 2. This is clearly illustrated by the separation of the phosphatidylcholines (PC), sphingomyelins (SM), and lysophosphatidylcholines (LPC) in the positive ion chromatograms using the HILIC UPLC method, shown in Figure 2A. The complete separation of these classes was not possible using the reversed-phase UPLC method.9,10 The triacylglycerols and diacylglycerols are not expected to be retained by the HILIC column and were not observed, whereas the ceramides (Cer) eluted early. In the negative ion mode, phosphatidylglycerols (PG), phosphatidylinositols (PI), and phosphatidylethanolamines (PE) were distinct from the PCs and eluted earlier, as shown in Figure 2B.
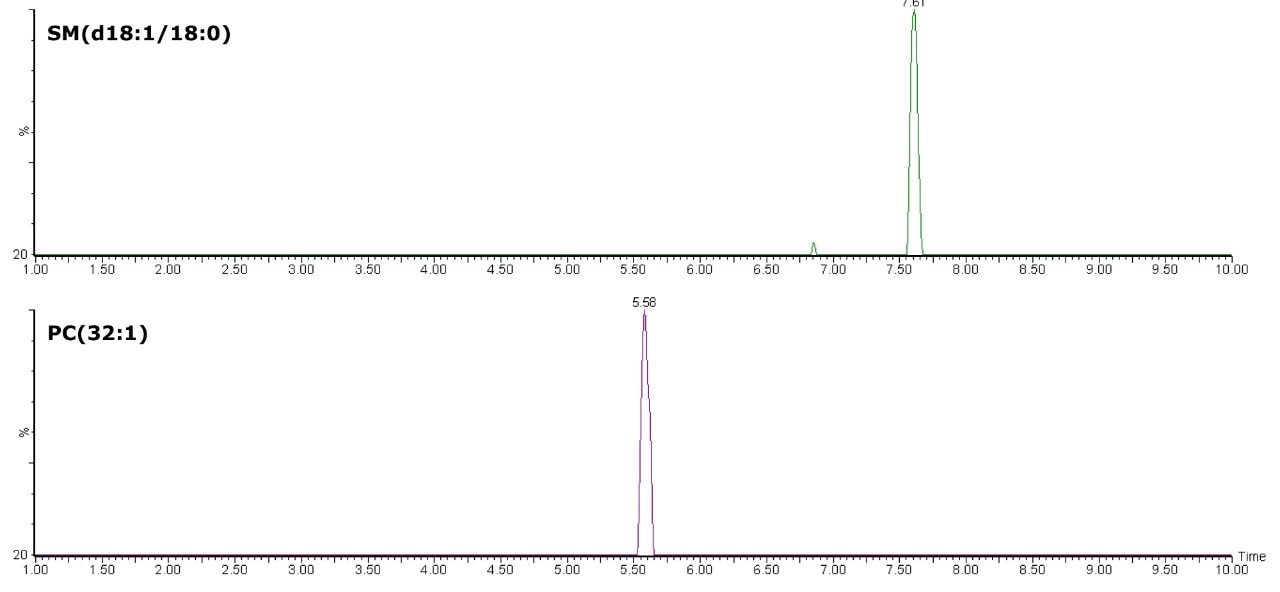
The separation of PCs from SMs using the HILIC is exemplified by PC(32:1), m/z 732.5545 and SM(d18:1/18:0), m/z 731.6069. These eluted closely using reversed-phase chromatography, but were clearly separated in approximately around 2 minutes, as shown in Figure 3. The separation of these two classes of lipids into distinct regions of the chromatogram allows unambiguous detection and measurement.
This HILIC-based UPLC method provides a simple and effective means to separate lipids by class in a similar fashion to normal-phase, and it offers a complementary separation technique to the reversed-phase methods we have previously described.9 This method is much faster than conventional normal-phase methods7 and it involves the use and disposal of less volatile and toxic solvents.
720004048, July 2011