UPLC-MSE is capable of characterizing digests of large, complex proteins such as monoclonal antibodies, with very high sequence coverage for both the heavy and light chains. Additionally, covalent modifications such as N-deamidation and N-linked glycosylation, can be identified. Peptide identifications are also supported by the high-quality MSE data.
Monoclonal antibodies are a class of recombinant proteins. They are subject to a variety of enzymatic or chemical modifications during expression, purification, and long-term storage. Liquid chromatography (LC)-based peptide mapping is the standard approach to establish protein identity, to identify post-translational modifications (PTMs), and to analyze potential impurities.
Peptide mapping with UltraPerformance LC (UPLC) Technology coupled with data-independent acquisition tandem mass spectrometry (UPLC-MSE) has been used to obtain high sequence coverage, and to identify PTMs and variants of enolase.1-2 The combination of MSE for unbiased identification3 along with the resolution and sensitivity of UPLC4-5 overcomes difficulties in protein characterization encountered with traditional LC-MS peptide mapping methods and data-dependent acquisition (DDA) LC-MS/MS sequencing tools, including co-elutions, analytical artifacts, and confusion of isobaric or nearly-isobaric peptides. MSE has the advantage over DDA in that it does not require peak detection or prior knowledge of the precursor ions.
In this application note, we demonstrate the use of UPLC-MSE to characterize a monoclonal antibody digest. The method provides high sequence coverage for both heavy and light chains of the antibody, and identifies modifications in a single analysis. MSE spectra were used to identify peptide sequences. The unexpected peptides resulting from non-specific digestion were also identified.
Commercial monoclonal antibody was in solution (21 μg/μL). Iodoacetamide (IAM), dithiothreitol (DTT) and ammonium bicarbonate (NH4HCO3) were purchased from Sigma Chemical Co. (St. Louis, Mo., U.S.). Sequence-grade trypsin was from Promega Corp. (Madison, Wisc., U.S.), Formic acid (FA) was from EM Science (Gibbstown, N.J., U.S.) and Optima-grade acetonitrile (ACN) was from Fisher Scientific (Pittsburg, Penn., U.S.). RapiGest SF was from Waters Corp. (Milford, Mass., U.S.). The water used in all procedures was from a Millipore Milli-Q purification system (Bedford, Mass., U.S.).
The monoclonal antibody solution was diluted to 5 μg/μL with 100 mM NH4HCO3 containing 0.1% RapiGest SF. Proteins in 50 μL freshly-prepared solution were digested by adding 5 μg of sequencing-grade trypsin and incubating at 37 °C overnight. Before the digestion, 5 μL 100 mM DTT was added to reduce disulfide bonds at 60 °C for 30 min. Free cysteine residues were alkylated with 5 μL of 200 mM IAM at room temperature for 30 min in the dark. Formic acid (0.5% v/v) was used to quench enzymatic reaction and degrade RapiGest SF. After adding ACN (to 30% v/v), the digest was diluted to a 1.5 pmol/μL final concentration with 5:95 ACN/water containing 0.1% FA and analyzed by UPLC-MSE.
All analyses were performed using a Waters SYNAPT MS system controlled by MassLynx 4.1 Software. An ACQUITY UPLC System equipped with 2.1 x 150 mm BEH300Å 1.7 μm Peptide Separation Technology C18 Column was used for the separation. Peptides were eluted with a 90 min gradient (0 to 50 %B). Mobile phase A was 0.1% FA in water, B was 0.1% FA in ACN. The flow rate was 0.2 mL/min, and the column temperature was 40 °C. An auxiliary pump delivered a lockmass solution (100 fmol/μL (GLu1)-fibrinopeptide B (GFP) in 50:50 ACN/water containing 0.1% FA) for mass accuracy reference.
The instrument was operated in the positive ion V-mode. An alternating low collision energy (5 V) and elevated collision energy (ramping from 20 to 40 V) acquisition was used to acquire peptide precursor (MS) and fragmentation (MSE) data. Scan time was 0.5 sec (1 sec total duty cycle). The capillary voltage was 3.0 kV, source temperature 100 °C, cone voltage 37 V, cone gas flow 10 L/h. Sampling of the lock spray channel was performed every 1 min.
The acquired data were processed with IdentityE Software of ProteinLynx Global SERVER 2.3. The processed data were first searched against a database consisting of monoclonal antibody light and heavy chain sequences, human, rat and mouse proteomes, with trypsin specificity and one potential miscleavage. Then, the data were searched again against the light and heavy chain sequences, without enzyme specificity. Cysteine (C) carbamidomethylation, asparagine (N) deamidation, and methionine (M) oxidation were allowed as optional modifications in these searches.
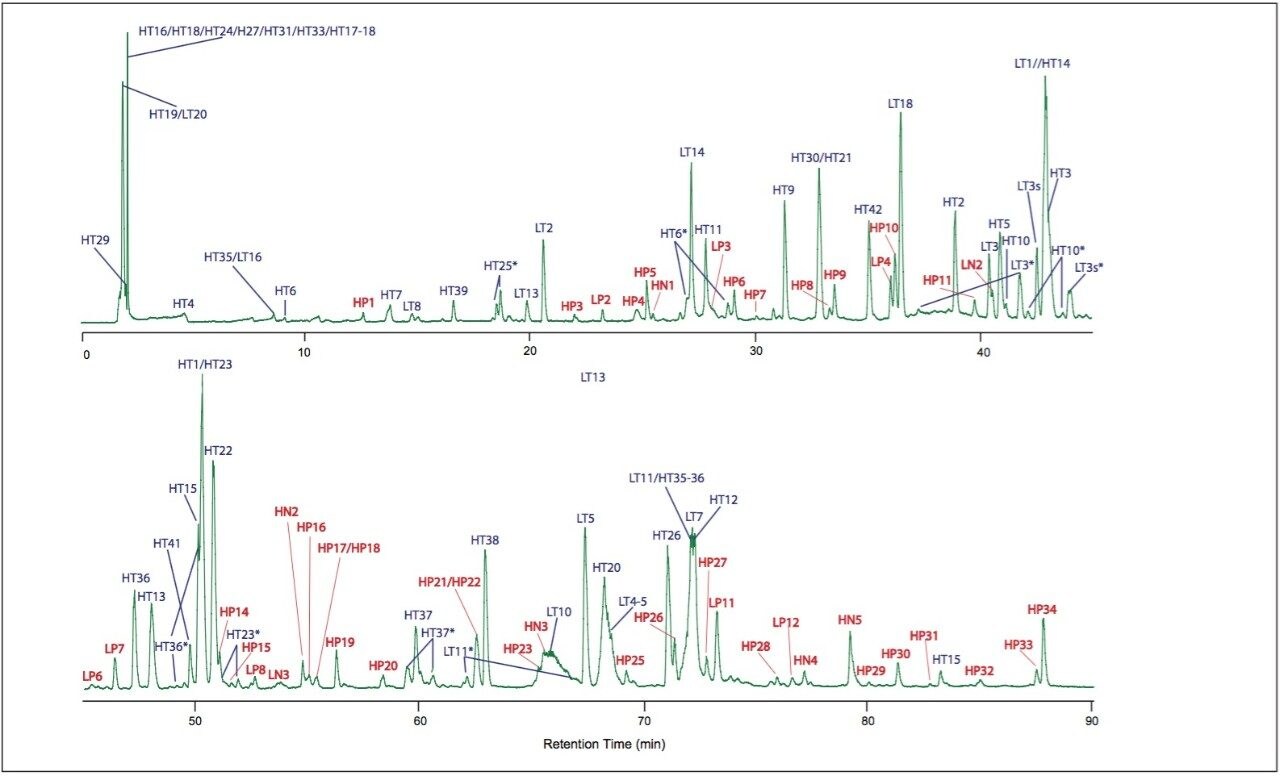
Figure 1 shows the UPLC-MSE chromatogram of monoclonal antibody digest (30 pmol injected on-column). The chromatogram features more than 100 detected peaks.
In order to investigate potential protein contaminants in the antibody, the data were searched against a database consisting of human, rat, and mouse proteomes, plus the light and heavy chain sequences. Only the light and heavy chains of the antibody were identified, demonstrating that the sample was not contaminated with other proteins. It has been shown previously that UPLC-MSE can identify protein contaminants at levels above 0.1 to 1%.1
The tryptic peptides identified are listed in Table 1 and assigned in Figure 1. They comprise 97% sequence of heavy chain and 97.2% sequence of light chain.
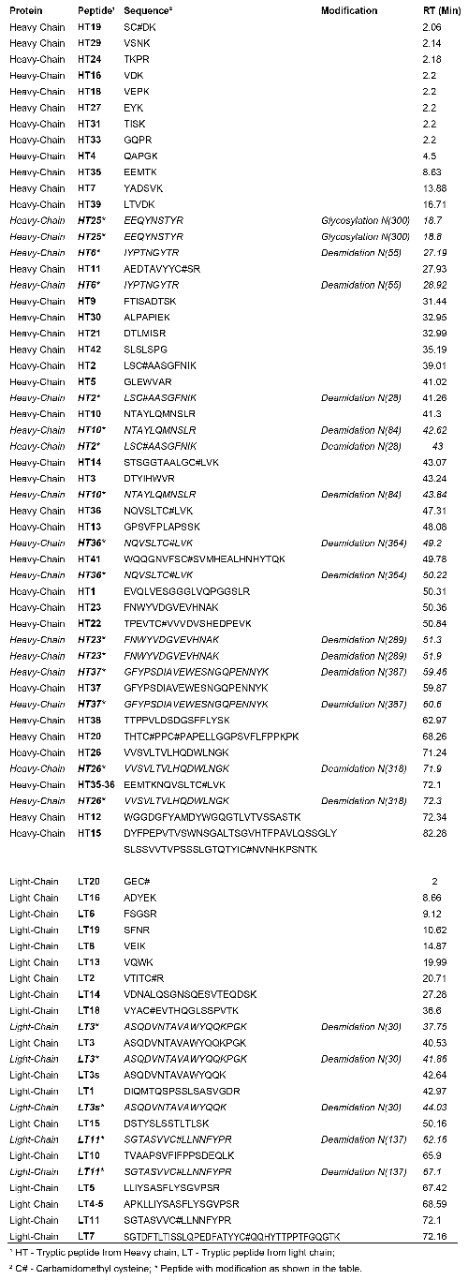
Peptide HT25 was identified as N-linked glycosylated forms. In addition, nine deamidated N sites (seven in heavy chain and two in light chain, two isoforms each as shown in Table 1) were also identified.
After assignment of the tryptic peptides, unknown peaks remain in the chromatogram. These peaks are partially tryptic and non-tryptic peptides, as identified in a subsequent database search against light and heavy chain sequences without enzyme specificity. The identified partially/non-tryptic peptides are presented in Table 2 and shown in Figure 1.
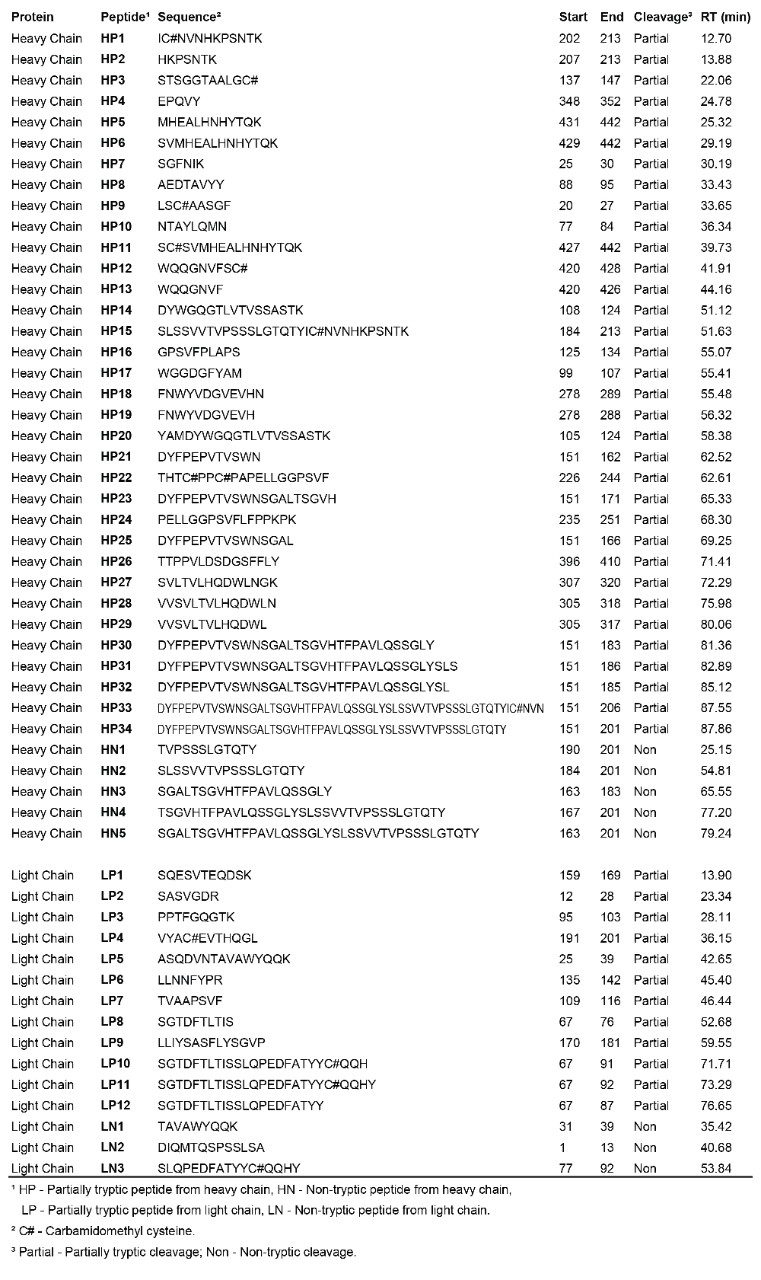
The second search suggests that the longest tryptic peptide HT15 in the heavy chain (63 amino acids; 6712.3 Da) was fragmented into a series of partially tryptic peptides (HP1, HP2, HP15, HP21, HP23, HP25, HP30, HP31, HP32, HP33, HP34) and non-tryptic peptides (HN1, HN2, HN3, HN4, HN5). All the MSE spectra of peptides listed in Table 2 were manually inspected, confirming the validity of identifications.
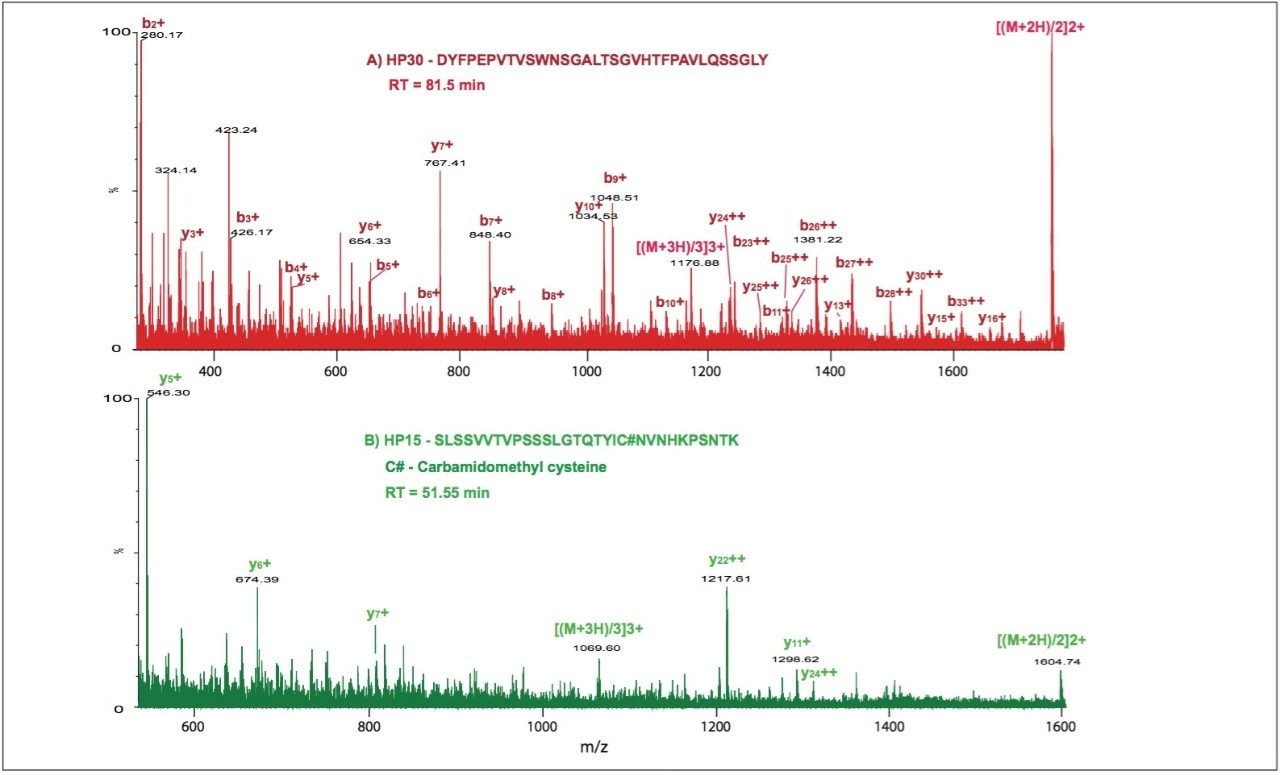
Figure 2 shows the MSE spectra for partially tryptic peptides HP30 and HP15, products of chymotryptic cleavage of tryptic HT15 peptide. When combined, they cover the entire sequence of the HT15.
Two broad peaks were observed in the chromatogram eluting between 65 and 70 minutes (Figure 1). They are proline-rich peptides LT10 (65.5 min) and HT20 (68.3 min). Their peak shapes could be improved by using higher column temperature. MSE allows for identification of peptides without complete resolution because it acquires fragment ion data of multiple peptide precursors in parallel.
The results of the analysis of a monoclonal antibody, with very high sequence coverage for both the heavy and light chains, illustrate that UPLC-MSE is capable of characterizing digests of large, complex proteins such as monoclonal antibodies. The unexpected peptides in the sample resulting from non-specific digestion were also identified with high confidence.
MSE enables sampling low-abundant components and acquires indiscriminately MSE spectrum. The specific conclusions from this study show that:
MSE collects precursor and fragment ion data of peptides within a single LC run, which enables the method to sequence the LC-eluted peptides. This ensures identification of unknown protein contaminants and PTMs in a single analysis. Therefore, the method may be used for detailed characterization of such proteins without additional MS/MS analyses and sacrificing the analytical efficiency. The method should increase the analytical efficiency in the characterization of therapeutic proteins.
720002821, January 2009