
This application note focuses on intact protein measurement and peptide mapping for biotherapeutics. We demonstrate a Waters system solution and workflow that integrates ACQUITY UPLC separations, application-specific column chemistries, Xevo QTof MS mass detection, and industry-leading biopharmaceutical informatics to deliver improvements in laboratory productivity. Tasks that took two weeks to complete, such as peptide mapping, can now be accomplished in a day.
The comprehensive characterization of biopolymers as drug candidates is a requirement for safety and regulatory agencies. Challenges include the high molecular weight of biopolymers and the heterogeneous nature of protein drugs, which require extensive characterization to achieve regulatory approval. Comparability during manufacturing changes must be demonstrated, or to provide intellectual property (IP) protection against biosimilars. The analyses described here show how different analyses can be performed to provide comprehensive information more quickly to meet these business objectives. All of these analyses can be done by non-specialists on the same platform.
The combination of liquid chromatography and electrospray mass spectrometry (LC-MS) provides enabling technology for well-characterized and comparable biotherapeutics. LC-MS analysis of proteins and peptides provides high levels of detail to aid characterization. However, the ability to routinely generate and interpret LC-MS data in a timely manner has been challenging in the past because involvement of an expert was required. Additionally, a major hindrance has been, until now, the lack of automated tools to complete the analysis. In order to increase laboratory productivity, high-performance mass spectrometers need to be made more accessible.
The Waters Xevo QTof MS System is a benchtop instrument designed to provide organizations with easy access to the most sensitive, high-performance, accurate mass MS, MSE, and MS/MS analysis available. The system incorporates Waters’ design philosophy of Engineered Simplicity and features tools for automated calibration and system monitoring, allowing organizations to obtain the very best MS performance in routine analyses.
This application note focuses on intact protein measurement and peptide mapping for biotherapeutics. We demonstrate a system solution that integrates ACQUITY UPLC separations, application-specific column chemistries, Xevo QTof MS mass detection, and industry-leading biopharmaceutical informatics to deliver improvements in laboratory productivity. Tasks that took two weeks to complete, such as peptide mapping, can now be accomplished in a day.
There are good reasons to determine the intact mass of a monoclonal antibody (mAb): in addition to providing an accurate mass of the protein, intact mass analysis provides an overall view of the heterogeneity of the protein, showing relative amounts of the various forms. Minimal sample preparation and chromatographic separation is required, so intact mass analysis provides results in minutes. Furthermore, generic methods can be used for rapid analysis of many different sample types.
In an organization where there is a large increase in the number of biopharmaceutical candidates advancing through the pipeline, such an approach provides results rapidly without having to request additional headcount.
For example, if there is a need to quickly confirm that the correct protein has been made, samples may be submitted to the analytical characterization group by biochemists or biologists who are not experts in mass spectrometry. An accurate molecular weight of the protein would ensure that the overall mass matches the expected mass. Without this confirmation, the wrong protein may be tested in several expensive bioassays, increasing time and costs for the organization.
A humanized IgG1 was received as a buffered solution (21.0 mg/mL). The solution was diluted to 0.5 mg/mL with 50 mM ammonium bicarbonate in preparation for intact mass analysis.
The antibody sample was reduced with DTT at 37 °C for 20 min using a published method.1 The solution was diluted to 0.10 mg/mL (pH 3.0) with 2% (v/v) formic acid aqueous solution.
RapiGest SF (0.05% in final solution) was added to the monoclonal antibody (mAb) stock solution, and the sample was heated at
60 °C for 30 minutes. The protein was then reduced with 10 mM
DTT at 60 °C for 30 min, and alkylated with 13 mM of IAA in the
dark for 45 minutes. Trypsin digestion was performed at 37 °C
overnight (trypsin/protein ratio was 1:50). The digest was diluted
to 0.015 mg/mL with 0.1% formic acid.
|
LC system: |
Waters ACQUITY UPLC System |
|
Columns |
• For intact IgG protein: MassPREP Micro desalting column 2.1 x 5 mm, 20 μm, 1000Å • For separation of IgG heavy/light chains: ACQUITY UPLC C4 BEH300 1.7 μm, 2.1 x 50 mm • For peptide mapping: ACQUITY UPLC C18 BEH300 1.7 μm, 2.1 x 150 mm |
|
Column temp.: |
• Intact IgG protein: 80 °C • Reduced monoclonal antibody: 80 °C • Peptide mapping: 60 °C |
|
Mobile phase A: |
0.1% Formic acid (Water) |
|
Mobile phase B: |
0.1% Formic acid (ACN) |
|
Flow rates: |
0.2 mL/min |
|
Gradients: |
• Intact monoclonal antibody: 10-90% B in 1.5 min. • Reduced monoclonal antibody: 25-35 %B over 15 min. • Peptide mapping: 2% B - 40B% in 90 min. |
|
MS system: |
Waters Xevo QTof MS |
|
Ionization mode: |
ESI positive |
|
Capillary voltage: |
3.0 kV |
|
Cone voltage: |
25 V (peptide) / 45 V (protein) |
|
Desolvation temp.: |
350 °C |
|
Source temp.: |
150 °C |
|
Desolvation gas: |
800 L/Hr |
|
Acquisition range: |
m/z: 50 to 1990 (peptide) / 600 to 4500 (protein) |
BiopharmaLynx Application Manager, v. 1.2, of MassLynx Software
Analysis of humanized IgG was used as an example to demonstrate the general LC-MS workflow as well as Xevo QTof MS’s performance. UPLC Technology provides significant advantages for all of these analyses by avoiding the need for flow splitting and by providing increased sensitivity for MS detection
The therapeutic was analyzed in three ways:

Information on heterogeneity is provided by intact mass measurement. Incomplete characterization of a candidate protein could lead to a delay in achieving regulatory approval, costing the company millions. In the worst case, if a protein is approved and later found to be incompletely characterized because of the presence of an undesired form, the product may have to be removed from the market, costing the company millions of dollars and hurting its reputation.
For choosing the optimum cell line, determining the protein intact mass analysis is a powerful, high-throughput method used to show changes in the protein’s heterogeneity with different cell lines, helping to reduce development and manufacturing costs. In QC functions, an intact mass analysis would give general confirmatory information.
Figure 2 shows mass spectra of an intact IgG from a traditional QTof mass spectrometer and the Xevo QTof MS, respectively. Both spectra show a charge envelope with a distribution of multiply-charged ion peaks. Although great similarity between the two spectra can be found, the Xevo QTof MS data show a six-fold sensitivity increase in comparison to traditional QTofs.
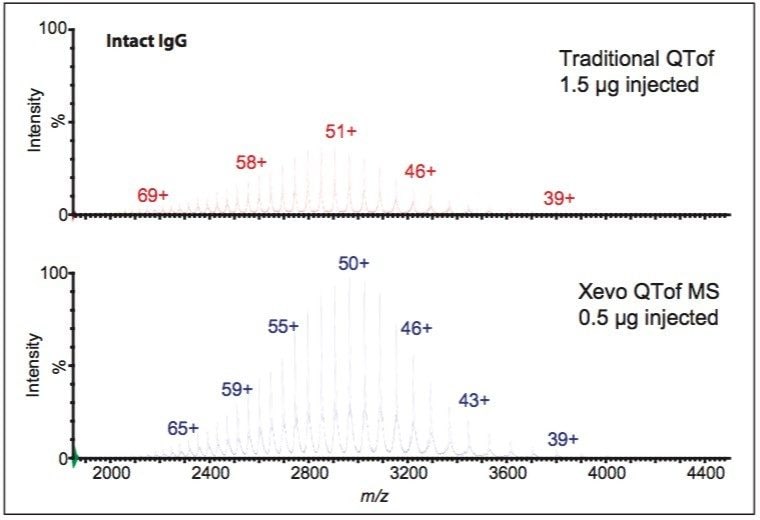
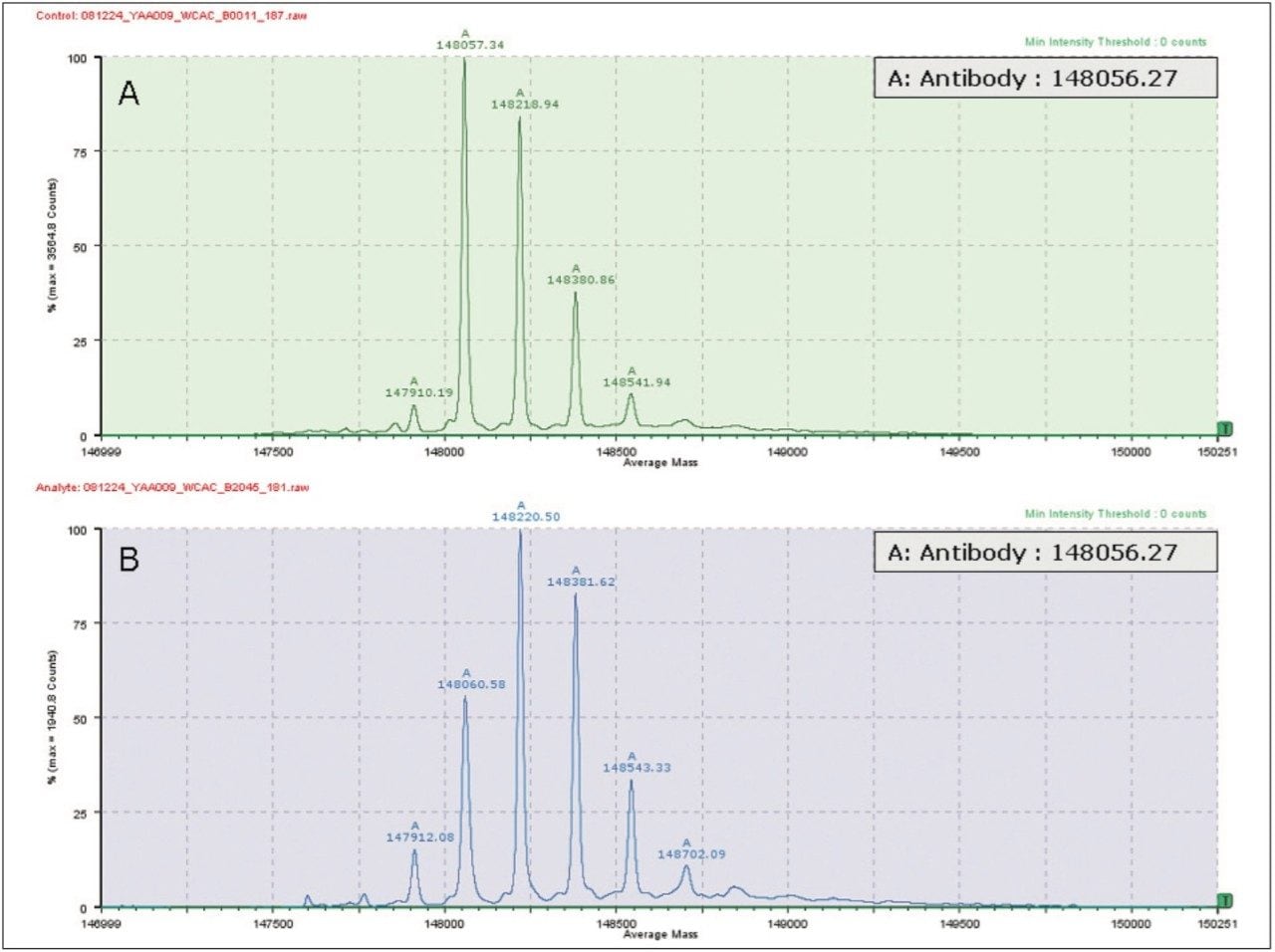
In a manual analysis, a scientist typically transfers this data to a software package to deconvolute the multiply-charged states to a zero charge molecular mass or distribution for the intact protein. With BiopharmaLynx 1.2 software, spectral deconvolution is performed automatically on the acquired data. The user is provided with an interactive browser page that displays a comparison of a control sample and an analyte so that differences are easily seen (Figure 3). The use of such software removes the need for intermediate manual transfers and avoids human bias. A large number of samples can be rapidly compared to a control and reports can be generated automatically for external users.
Chromatographic separation of the heavy and light chains adds further detail to the characteristics of an mAb, confirms that the glycoprotein profiles of candidate expression clones match the expectation, or checks whether there are unusual distributions of glycoforms. When specific glycoforms are of interest, the separation of light and heavy chains helps to examine glycoforms more closely.
The separation of light chain and heavy chain also allows the relative component quantitation to be achieved, and modifications specific to the light or heavy chains can be characterized. Mass changes associated with specific residues (such as succinimide formation) can be detected by analysis of partially reduced monoclonal antibodies (or of peptides from enzymatic digestion as discussed below) with chromatographic separation.
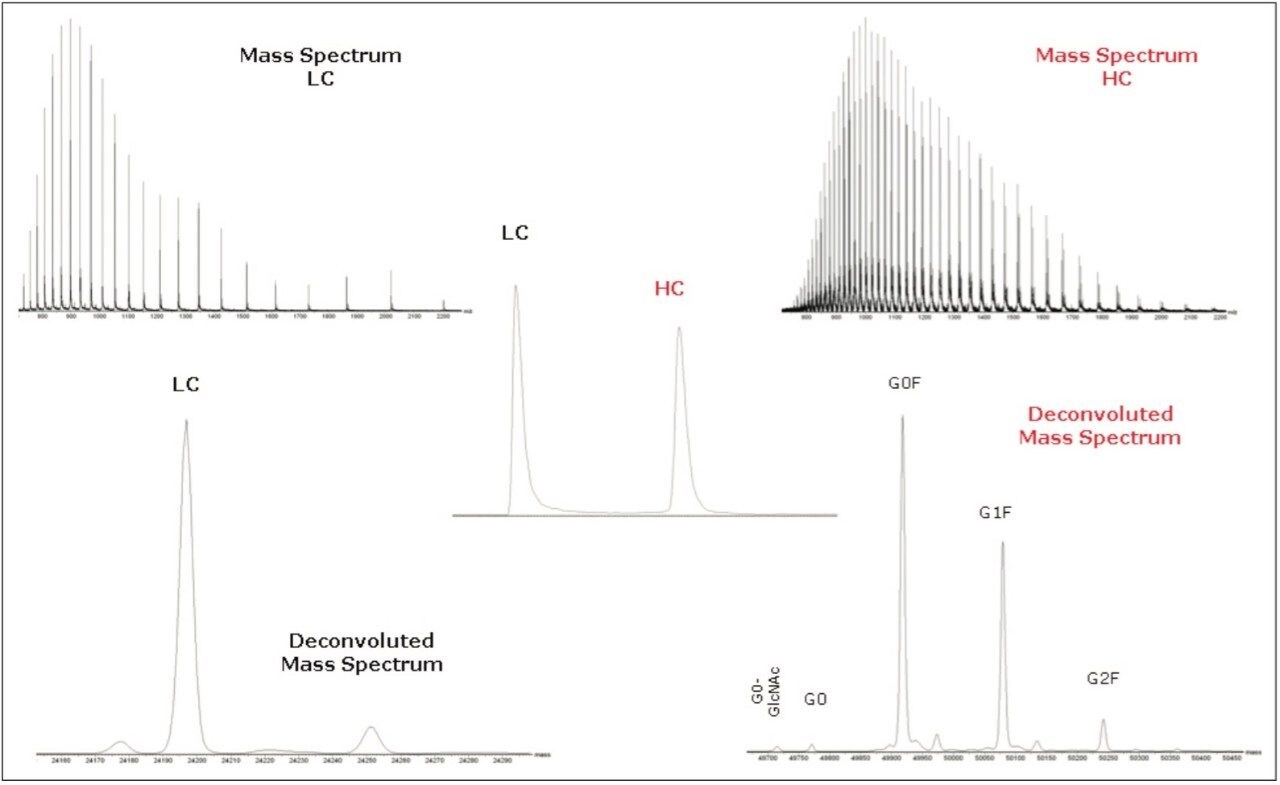
Figure 4 displays separate spectra for the heavy and light chains from a reduced monoclonal antibody. The corresponding deconvoluted spectra for the light chain and heavy chain are also shown. The chromatogram shown in inset demonstrates that high resolution separation by an ACQUITY UPLC BEH C4 column enables the differentiation of minor isoforms of either heavy chain or light chains, providing enhanced assessment on the heterogeneity of the sample.
In intact mass analysis, the overall heterogeneity of the protein is obtained. In order to characterize modifications that lead to small or no mass changes in the intact protein mass (e.g., deamidation or structural isomers of glycans), peptide mapping with MS detection provides a valuable technique for detailed and comprehensive modification coverage in a simple manner.
In LC-MS this can be done with ease using only one enzyme to digest the protein and obtain sequence coverage above 95%.The ability of a peptide map to highlight small changes in the primary structure of a protein makes it valuable for establishing the identity, purity, and composition of a protein. For example, in shelf life studies the ability to detect modifications is vital. In the optimization of in vitro folding processes the arrangement of disulfide bridges is crucial.
LC-MS analysis has greatly expanded the capability to develop and interpret data from peptide mapping experiments, compared to traditional LC-UV/VIS detection. However, single-stage MS analysis may not be capable of resolving peptides that are isobaric, nor can suspected peptide modifications be identified and localized from MS data alone. MS/MS data are therefore used to generate fragment ion mass spectra. In the past, MS/MS data were acquired with selected peptides using a traditional data-dependent (or directed) MS/MS acquisition (DDA). We present here an approach that is a significant improvement on DDA by using MSE analysis to confirm sequence information.
MSE provides a comprehensive MS/MS picture without any prior knowledge of the sample and is applicable across all charge states. (See references 2-7 for details of MSE.) This means that the peptide map can be performed in an unbiased, systematic manner and provide consistent results without the need for an expert. The methodology also means that the same dataset could potentially be re-analyzed at a later stage for additional information without having to re-run the samples – saving significant costs, time, and capital investment.
In the BiopharmaLynx browser, the user is able to interactively annotate data that might previously have been overlooked. Modifications are annotated automatically and disulfide bridges can be shown on the peptide map. Increased sensitivity provided by the Xevo QTof MS enhances the information available with no compromise in productivity.
In Figure 5, an Enolase digest was analyzed by UPLC-MSE. The comparison between a traditional QTof and the new Xevo QTof MS highlights the improvements in sensitivity for this workhorse instrument. In BiopharmaLynx 1.2, additional information on peptide sequence is also provided. This confirms the sequence of the peptides from the protein. Figure 6 shows a screenshot from the tabular browser display presented to the user. In the right-most column shown, the y- and b- ions are listed to indicate exactly which backbone fragments were automatically identified from the MS/MS spectra. The user is also able to select alternative sequences where they are presented or reject them if the user prefers a different interpretation. Figure 6.
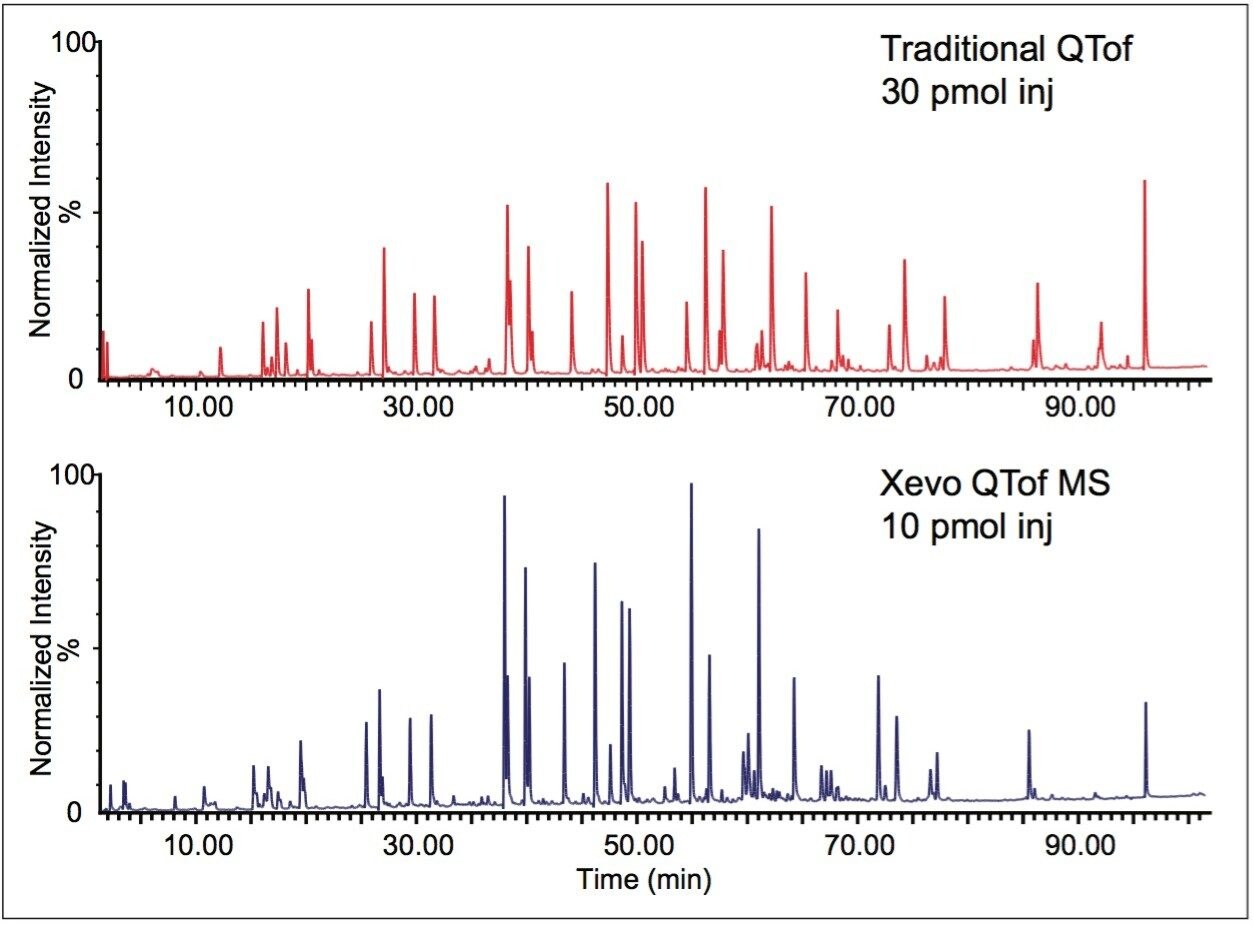
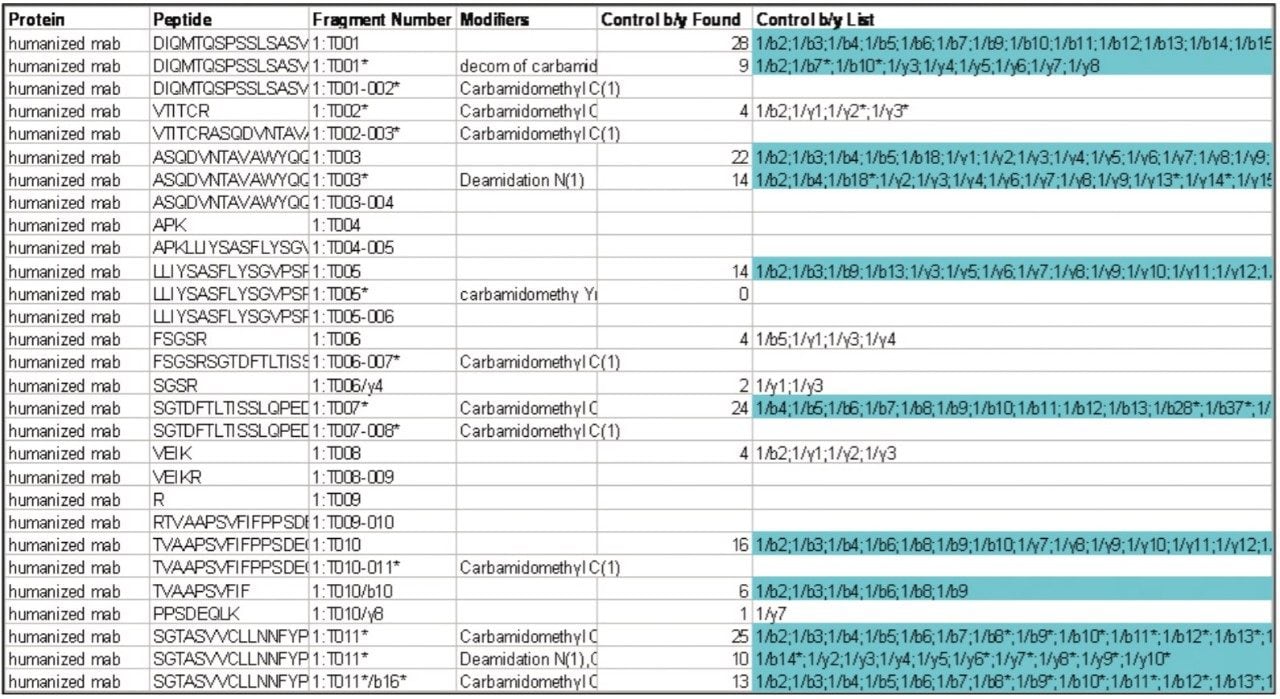
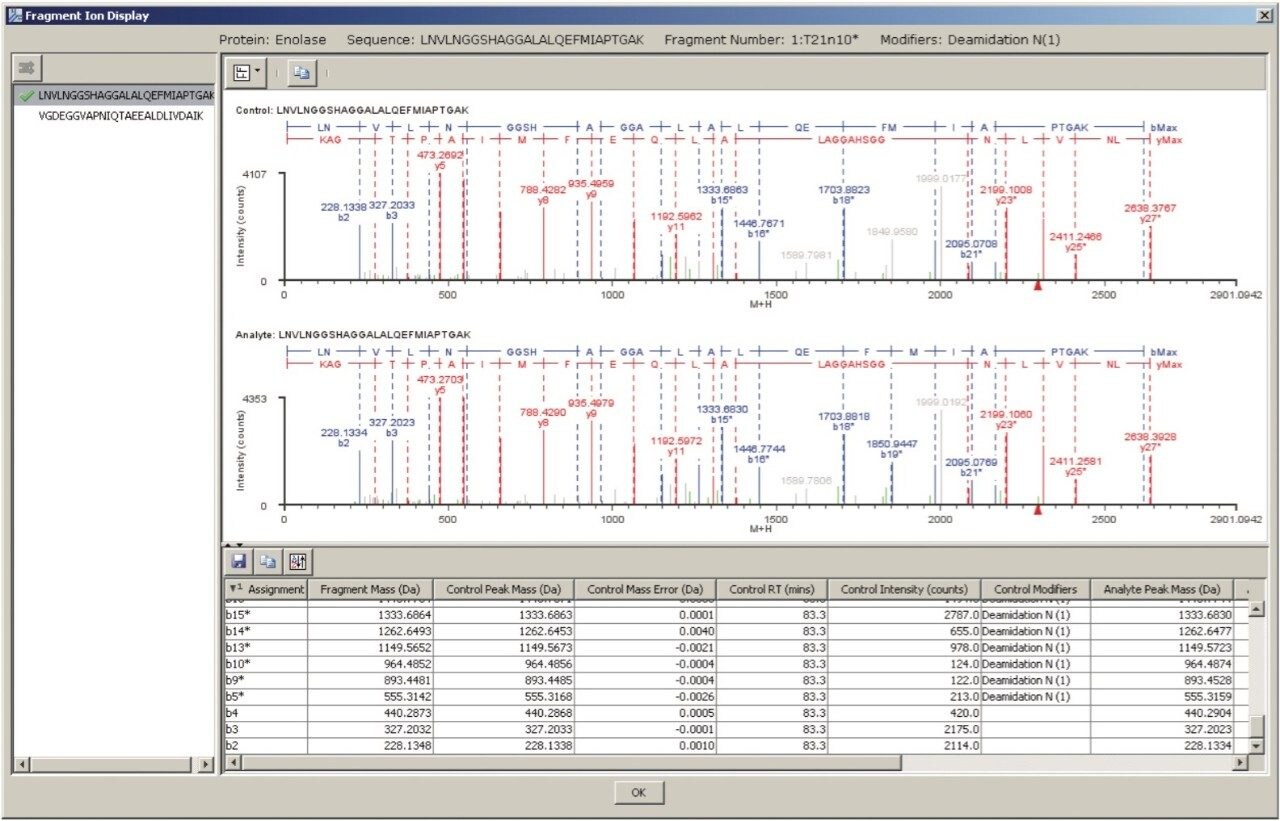
Figure 7 shows the sequence information superimposed on a spectrum in the BiopharmaLynx browser. The y-series amino acid sequence information is shown in red underneath the b-series sequence in blue.
720003046, May 2009