
The UNIFI Scientific Information System offers a streamlined approach for the management of data and metadata associated with high-resolution accurate mass screening libraries.
Using purchased standards for the compounds of interest, a comprehensive scientific library may be constructed by:
Information from previous targeted screening experiments may also be of use, such as known MRM transitions for the compounds of interest.
The acquisition of information-rich datasets collected using techniques such as MSE is placing a demand on the content of the scientific libraries used to screen for a large number of compounds in complex matrices. Using more criteria with wider tolerances controls the number of false detects while ensuring that false negatives are not introduced in a high-resolution mass spectrometry (HRMS) screening experiment. In order to capture and manage the wealth of analytical information that can be gained from powerful chromatographic separations and HRMS, Waters has created the UNIFI Scientific Library.
Software functionality within UNIFI aids in the creation of a scientific library that contains all the critical detection criteria, such as retention time and the theoretical masses of fragment ions generated using structural information.
Here we demonstrate the ease with which a comprehensive scientific library can be built within UNIFI, considering two different scenarios:
For both scenarios the following four steps were taken to build the new scientific library, as described in the workflow shown in Schematic 1.
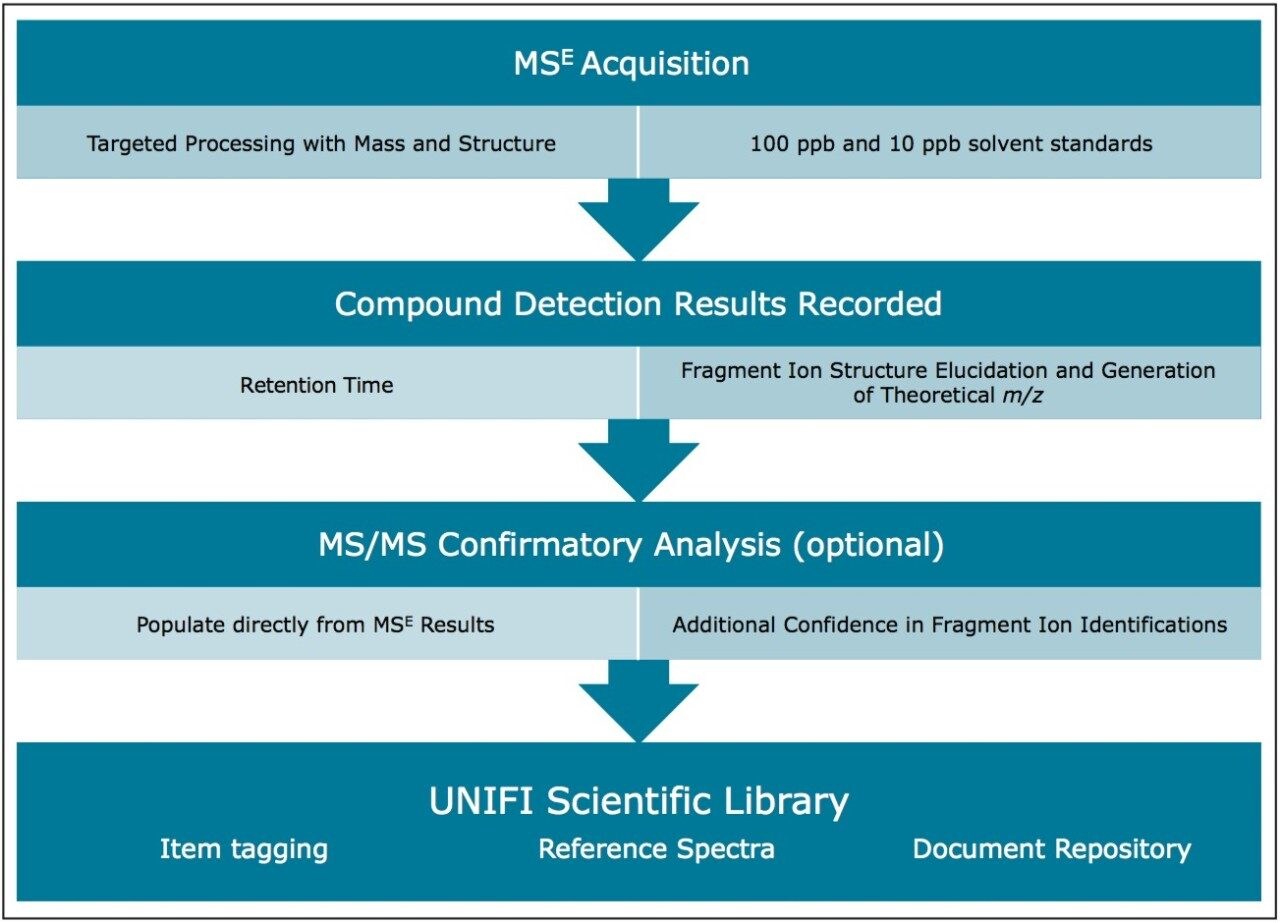
The information stored in a scientific library is a powerful tool for ensuring confident identifications in future HRMS screening experiments. Once a compound is added to the library, relevant reference and classification information can be added within the software. Due to the emergence of novel pesticides, metabolites and other newly recognized contaminants, the ability to create additions to the existing scientific library is of great utility.
|
LC system: |
ACQUITY UPLC I-Class |
|
Column: |
ACQUITY UPLC BEH C18 1.7 μm, 2.1 x 100 mm |
|
Column temp.: |
45 °C |
|
Injection volume: |
10 μL |
|
Flow rate: |
0.45 mL/min |
|
Mobile phase A: |
10 mM ammonium acetate (pH 5) in water |
|
Mobile phase B: |
10 mM ammonium acetate (pH 5) in methanol |
|
Sample manager purge: |
90/10 water/methanol |
|
Sample manager wash: |
50/50 water/methanol |
|
Seal wash: |
90/10 water/methanol |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
Curve |
|---|---|---|---|---|
|
Initial |
0.45 |
98 |
2 |
6 |
|
0.25 |
0.45 |
98 |
2 |
6 |
|
12.25 |
0.45 |
1 |
99 |
6 |
|
13.00 |
0.45 |
1 |
99 |
6 |
|
13.01 |
0.45 |
98 |
2 |
6 |
|
17.00 |
0.45 |
98 |
2 |
6 |
Table 1. UPLC method for the Pesticide Screening Application Solution.
|
MS system: |
Xevo G2-S QTof |
|
Ionization mode: |
ESI + and - |
|
Capillary voltage: |
1 kV |
|
Desolvation temp.: |
550 °C |
|
Desolvation gas flow: |
1000 L/Hr |
|
Source temp.: |
120 °C |
|
Reference mass: |
Leucine enkephalin [M+H]+=556.2766 |
|
Acquisition range: |
m/z 50 to 1200 |
|
Acquisition rate: |
4 spectra/s |
|
Low CE: |
4 eV |
|
High CE ramp: |
10 to 45 eV |
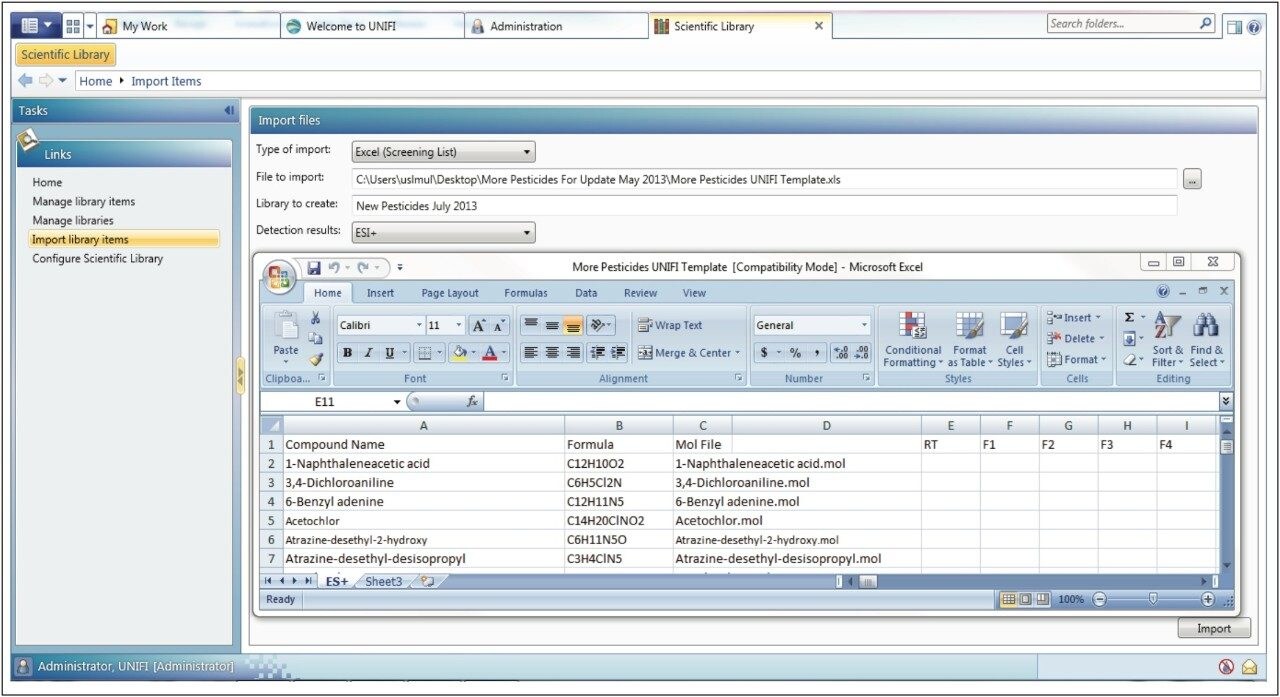
In order to include compound structures in the UNIFI Scientific Library, .mol files are required for each of the compounds of interest. These can be downloaded from ChemSpider(www.chemspider.com). An Excel spreadsheet containing the names of the compounds of interest, their molecular formulae, and the name of the corresponding .mol file were saved in the directory containing the .mol files. This spreadsheet was then imported into UNIFI as a scientific library container (ULC), to be used as the initial screening list, shown in Figure 1.
A new analysis method was created using the Accurate Mass MSE Experiment type. The instrument settings of the Pesticide Screening Application Solution (PSAS) were used for this analysis. For QC purposes, the Pesticide Screening Mix (p/n 186006348) compounds were also added to the component list. Prior to analysis of the new compounds, a test injection of 10 ppb of the pesticide screening mix was performed. The mass error and observed retention times of the pesticide screening mix components were checked against the expected values, according to our QC requirements, to provide consistency in the library updates (Figure 2). This step ensures uniform results between new library entries and those created in the past, by different users, or on a different system.

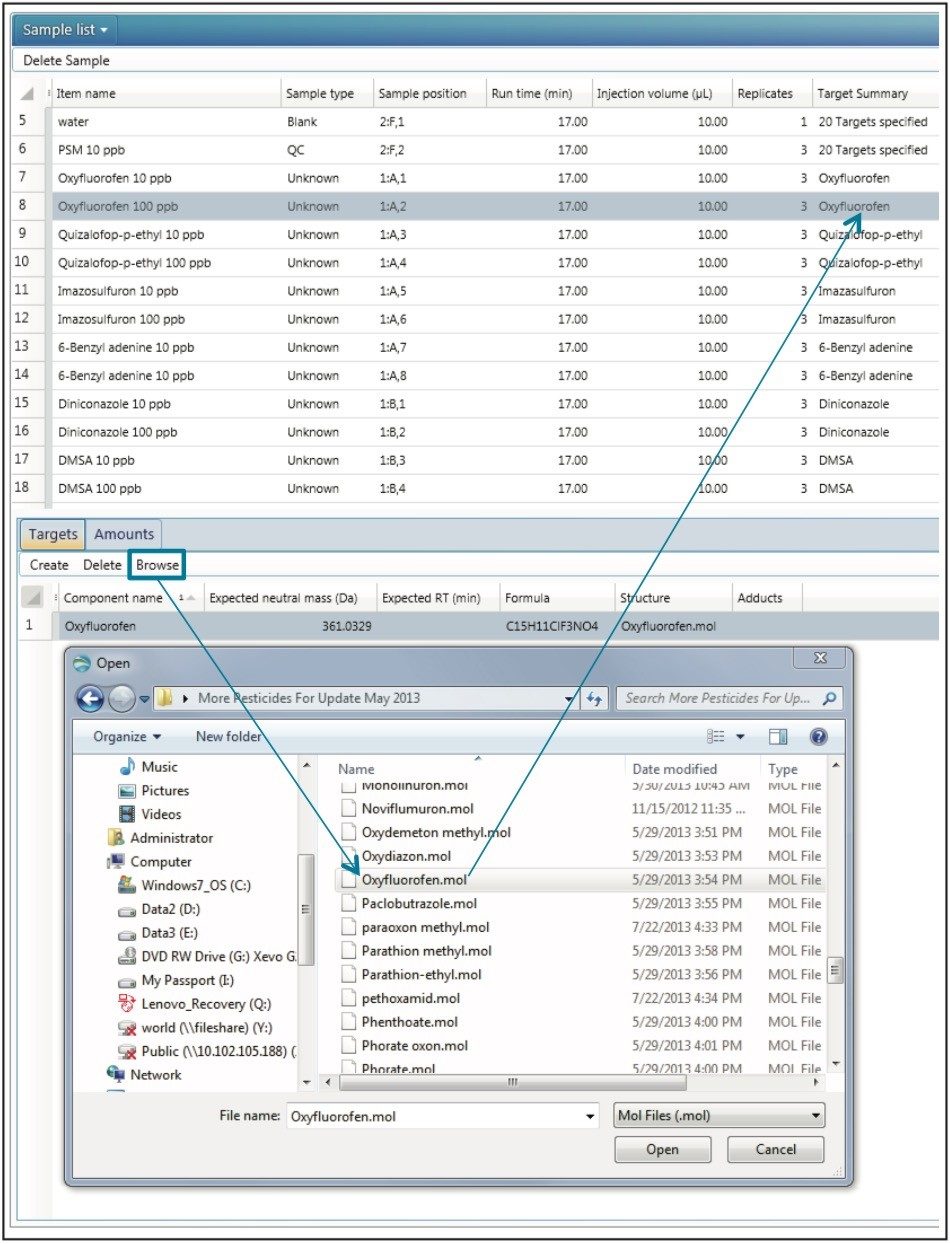
Standards of the new compounds in Milli-Q grade water were injected at two concentrations, 10 ppb and 100 ppb, to benchmark the sensitivity and response of each compound. For each of the standards injected, a target list of the .mol files for the compound(s) in that standard can be added directly to the sample list, as shown in Figure 3. For the pesticide Screening Mix, all of the compounds were added to the target list (first line, Figure 3 “20 targets specified”). Using a target list can be used as an alternative to the aforementioned Excel spreadsheet. Upon analysis, retention times were easily determined, based on identifications made using the exact mass, as can be seen for the rodenticide Coumachlor in Figure 4.
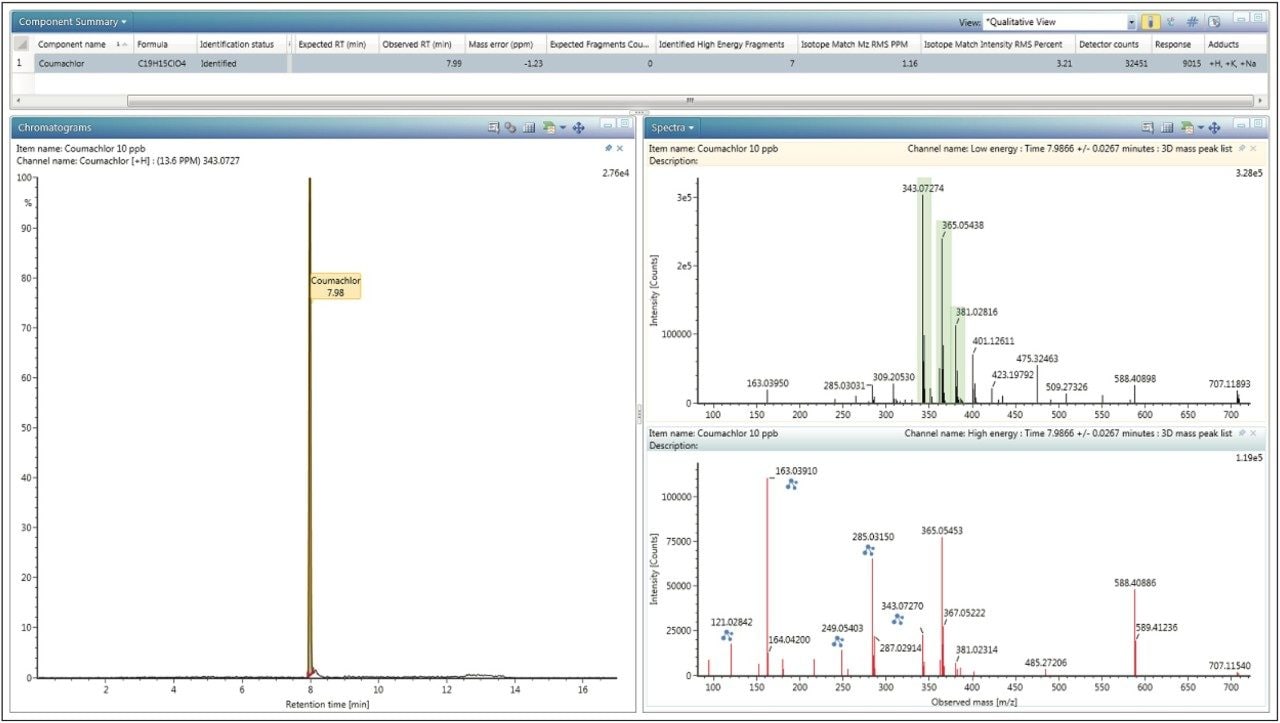
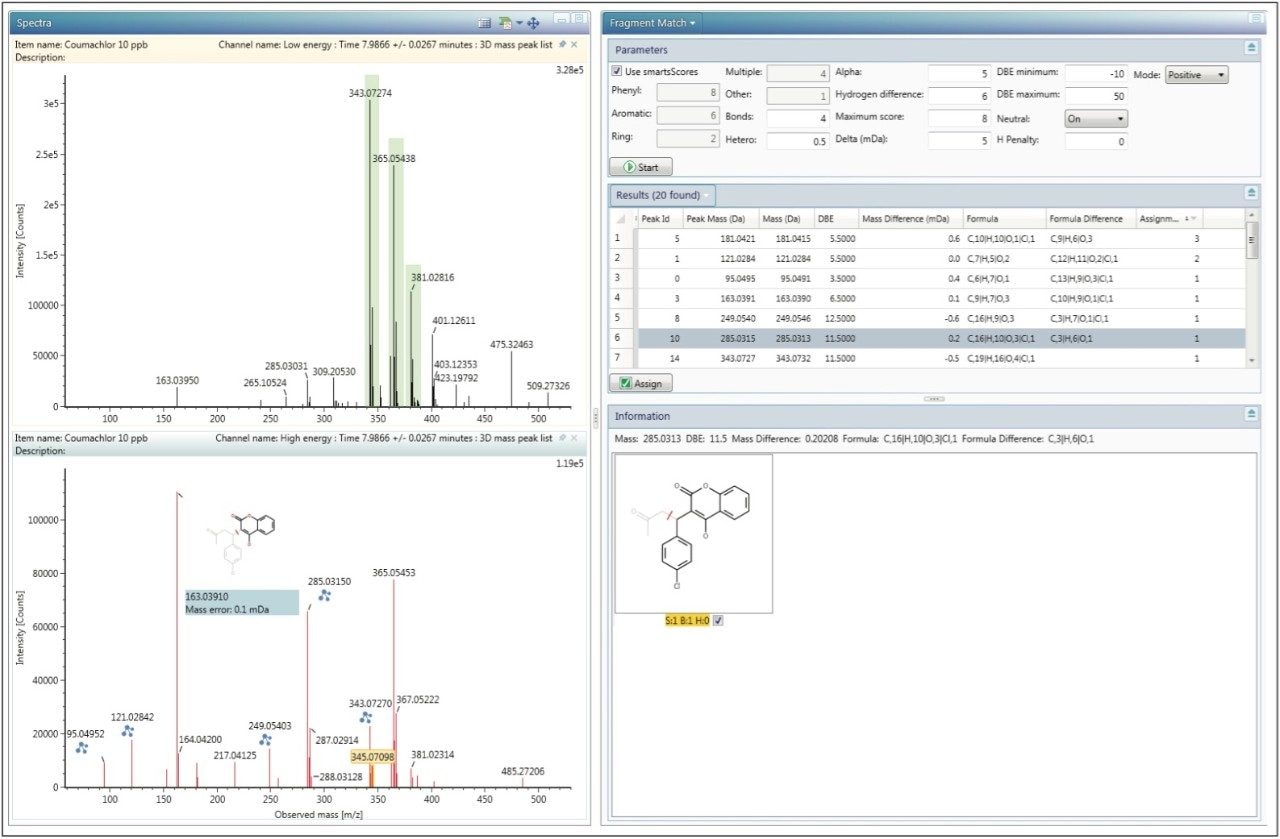
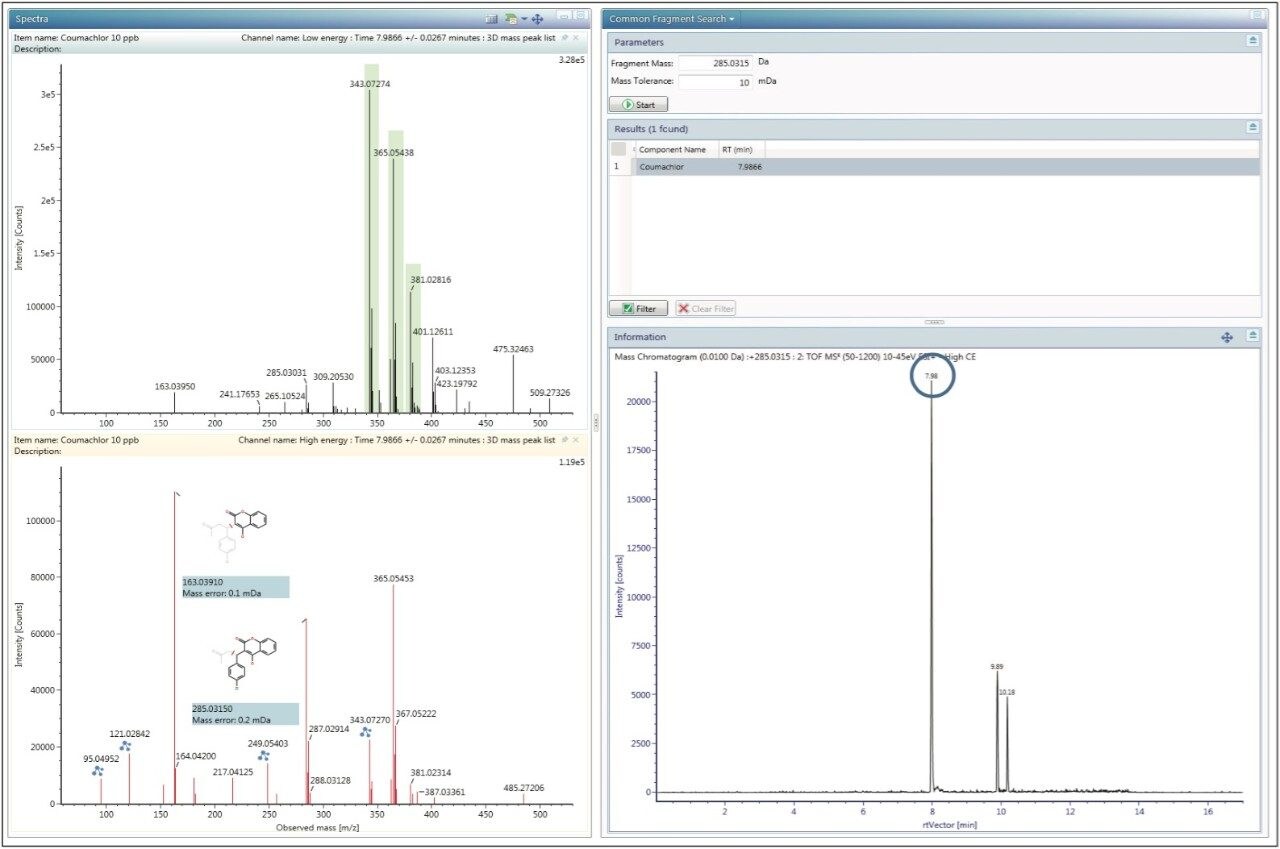
Continuing with the Coumachlor example, the Fragment Match results (Figure 5) were displayed by right clicking on the name of the identified compound and selecting Elucidate. The most ideal fragment ions to include in the scientific library were assigned based on their intensity and match scores. Common Fragment Match functionality was used to bring up an XIC of the proposed fragment mass, shown in Figure 6. The chromatogram provides more confidence in correct fragment ion assignment and also gives an indication of the selectivity of that particular fragment ion. If prior knowledge of MRM transitions for a specific compound is available, referring to those would offer additional data for fragment ion selection. Using this approach, theoretical fragment masses were determined and were used to populate the new library, either by adding to the Excel spreadsheet, or entering them directly into the UNIFI Scientific Library. Once the information for all compounds was added, entries from the newly created scientific library were replaced in the original analysis and the data re-processed to ensure that all compounds were identified using the new library.
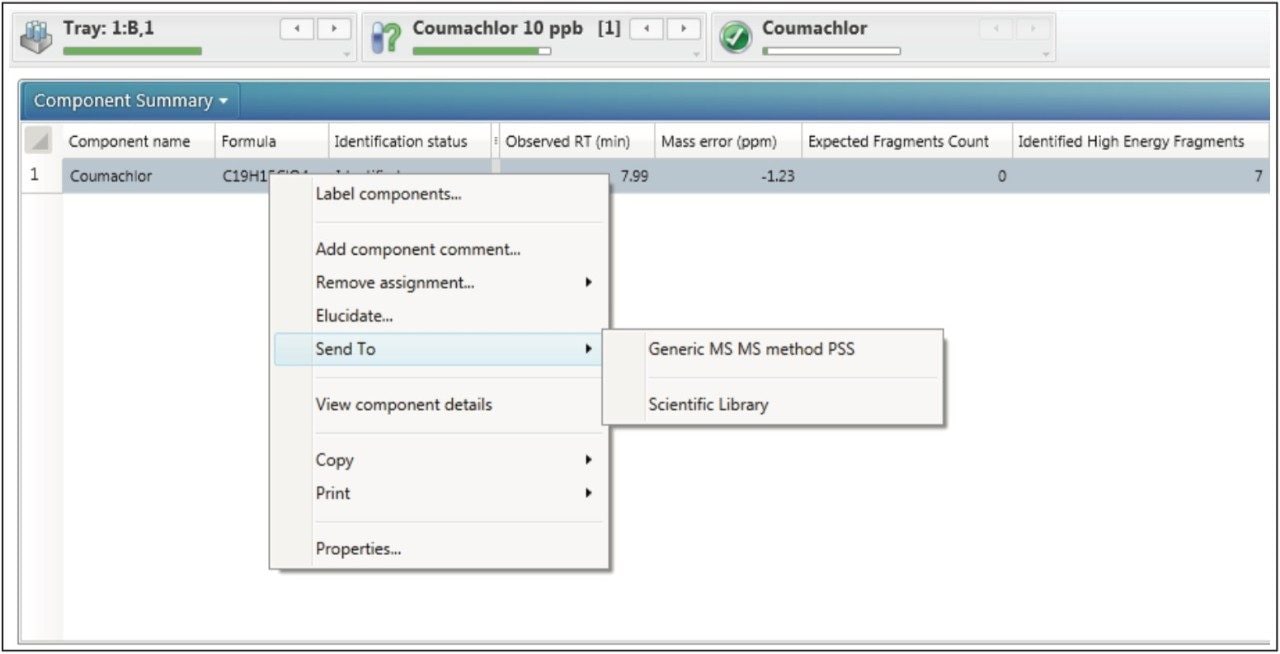
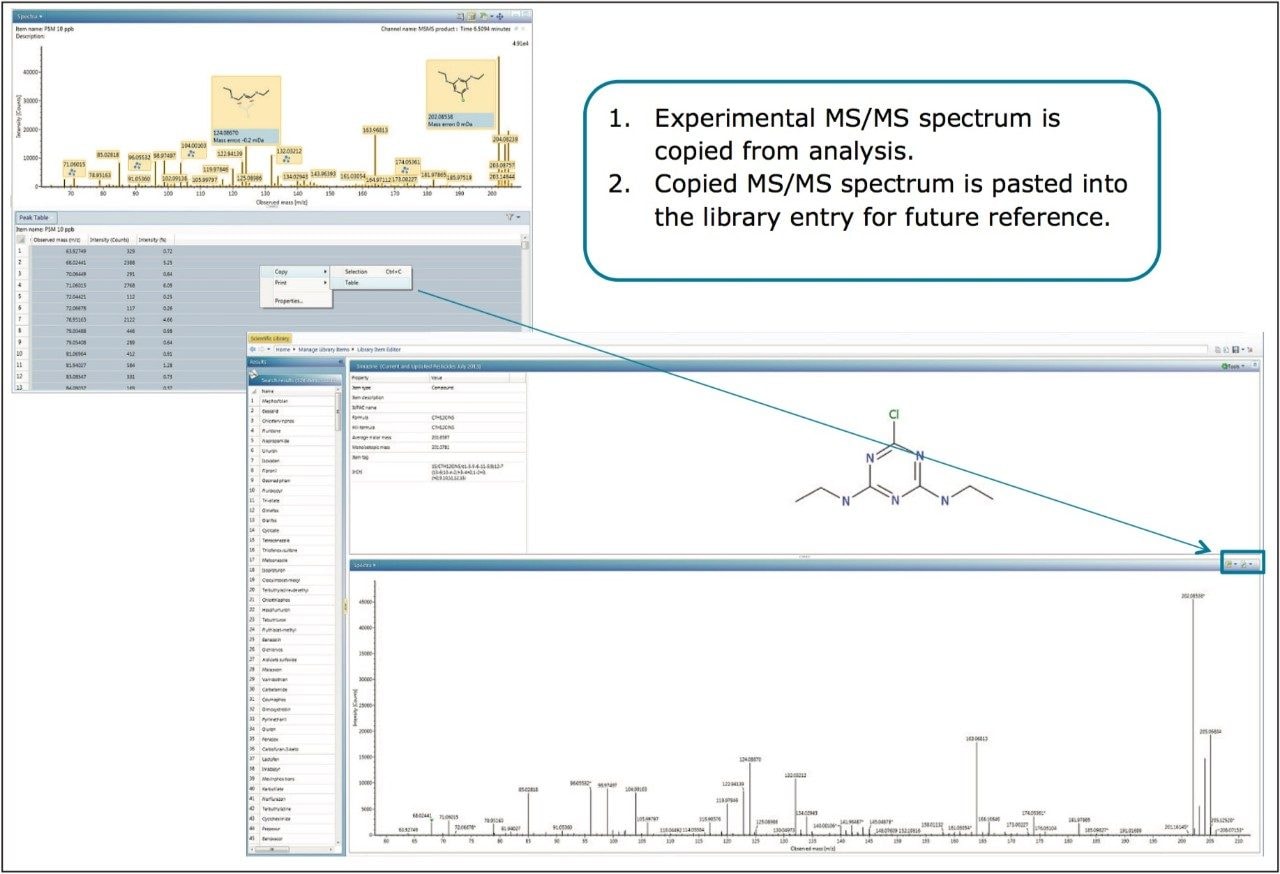
As an additional means of confirmation of the selected fragment ions, MS/MS analysis can be performed on the standards following MSE identification. In the past, separate MS/MS experiments had to be set up for each compound of interest, which was laborious and time consuming. UNIFI allows the user to directly import the compound information and detection results from the MSE analysis by clicking Send to MS/MS. With a generic MS/MS method open, compound specific information from the MSE analysis are sent to the MS/MS analysis by right clicking the identification, then select the Send To (Figure 7). As with the MSE analysis, system performance was first verified using the Pesticide Screening Mix to ensure acceptable retention times and mass accuracies. From these acquisitions, MS/MS spectra of compounds can be added to the scientific library for future reference (Figure 8).
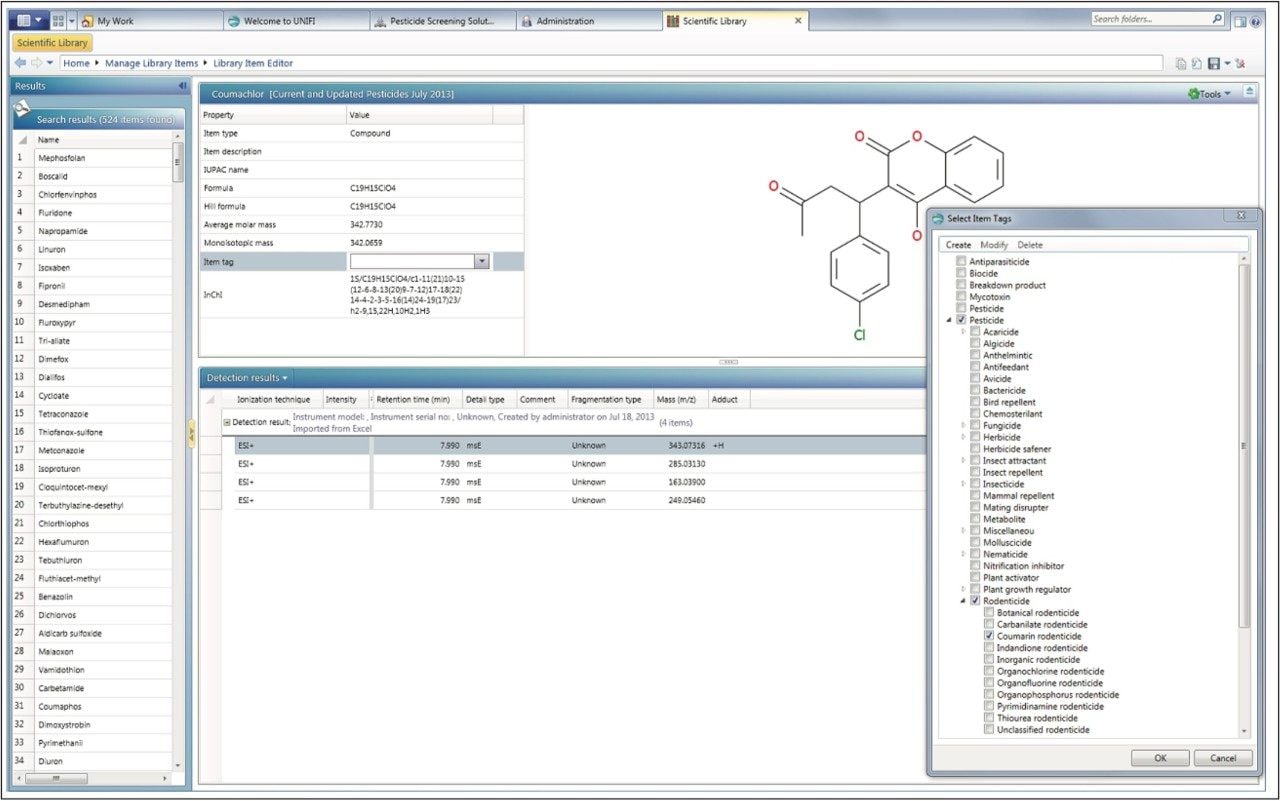
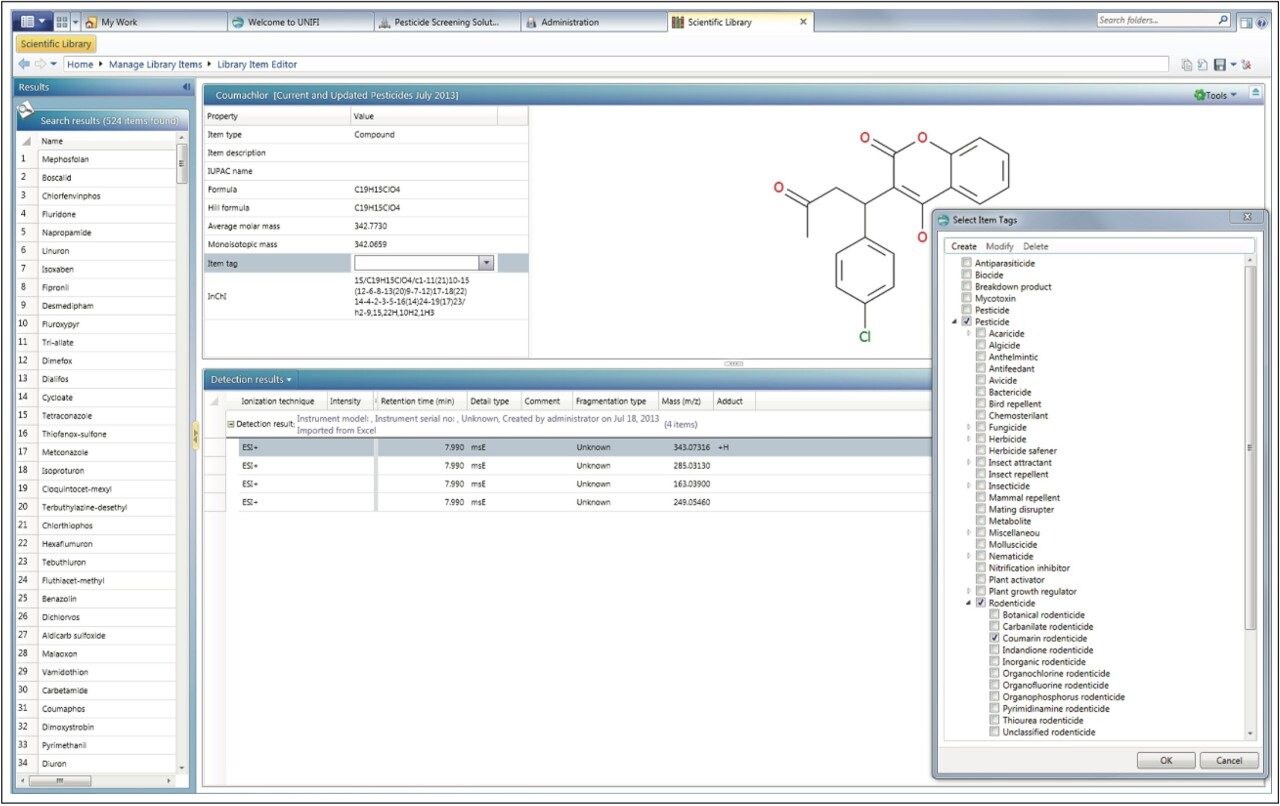
In order to maximize the available information and facilitate future searches the UNIFI Scientific Library can associate user-definable tags, references, or other documents with library entries. In the example presented here, individual pesticides were tagged with their pesticide class and sub class, if available (Figure 9). References were added and categorized (Figure 10). Any edits can be made through the Manage Library Items section of the scientific library.
720004927, January 2014