
This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates to develop a method for the analysis and quantification of aminoglycoside residues in bovine milk by ion-pair liquid chromatography using the ACQUITY UPLC System coupled with the Xevo TQ-S Mass Spectrometer.
Aminoglycosides (AGs) are a group of broadspectrum of antibiotics that have bactericidal activity against some aerobic gram-positive and gram-negative organisms. Their structures characteristically contain two or more amino sugars linked by glycosidic bonds to an aminocyclitol component. Stringent maximum residue levels (MRLs) for AGs in various animal-origin foods have been issued by the European Union (EU), United States, Japan, and other countries. Therefore, there is a need to develop sensitive and reliable analytical methods to monitor trace-levels of AG residues in complex matrices.
There are several challenges faced by analysts performing AG analysis – difficulties that arise from extraction, clean-up, and separation, to detection of AG residues. Some of the AGs have several pKa values, complicating the development of a generic sample preparation method. These compounds exhibit strong hydrophilic behavior, with a tendency to undergo strong binding to matrix proteins. AGs are also poorly retained on reversed phase columns.
Liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) is becoming an emerging technique in the analysis of AG antibiotic residues, due to its high sensitivity, and because it doesn’t require the lengthy derivatization reactions in GC or GC-MS analyses.
In this technology brief, a multi-residue UPLC-MS/MS method was developed for the routine analysis of 13 aminoglycoside residues in bovine milk.
Separation of aminoglycoside antibiotic residues was performed using a Waters ACQUITY UPLC System, on an ACQUITY UPLC HSS PFP Column with ion-pair chromatography. Volatile heptafluoroabutyric acid (HFBA) was used as an ion-pairing reagent. HFBA is compatible with mass spectrometry and enabled strong retention of AG residues on the HSS PFP Column. Trichloroacetic acid (TCA) was used to extract AG residues from bovine milk, and further cleanup was performed using Oasis SPE Cartridges. The final extract was reconstituted in 20 mM HFBA prior to the UPLC-MS/ MS analysis.
The Xevo TQ-S Mass Spectrometer was used in ESI positive mode for multiple reaction monitoring (MRM) and RADAR full scan acquisition. RADAR provides continuous monitoring of the matrix background to ensure a robust and reliable method.
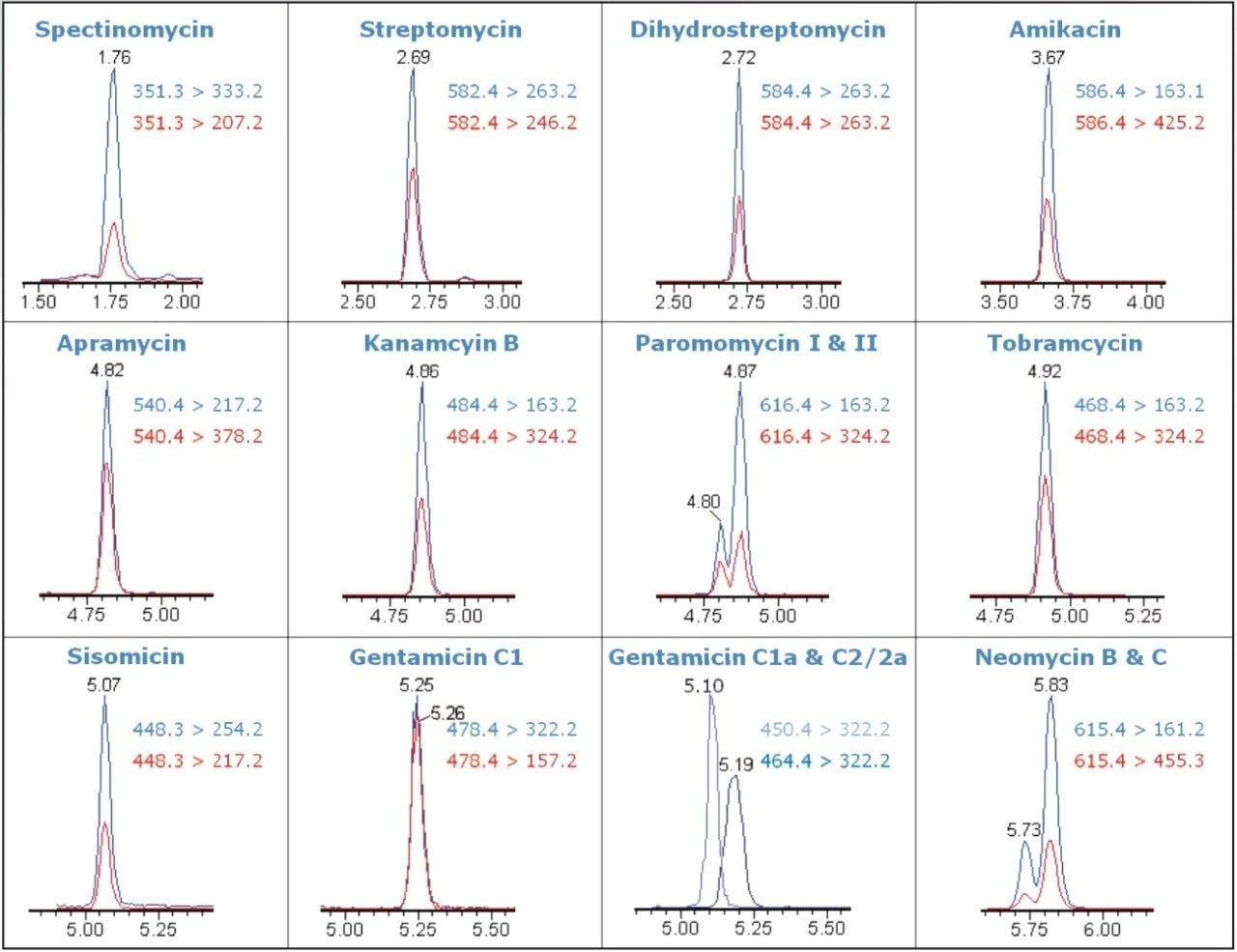
Figure 1 shows the MRM chromatograms for aminoglycoside residues spiked into bovine milk at 10 μg/kg. Quantitative (blue trace) and confirmatory (red trace) transitions were both detected at this level, which is at least 10x below the EU MRL levels (0.1 to 0.5 mg/kg). The ultra-high sensitivity of Xevo TQ-S enabled quantification and confirmation of AGs below the required regulatory levels, even in the presence of signal suppressing ion pairing reagents. Neomycin and paramomycin exist as isomeric components (neomycin B and C, paramomycin I and II respectively), and using this method, these isomers were partially separated, as shown in Figure 1.
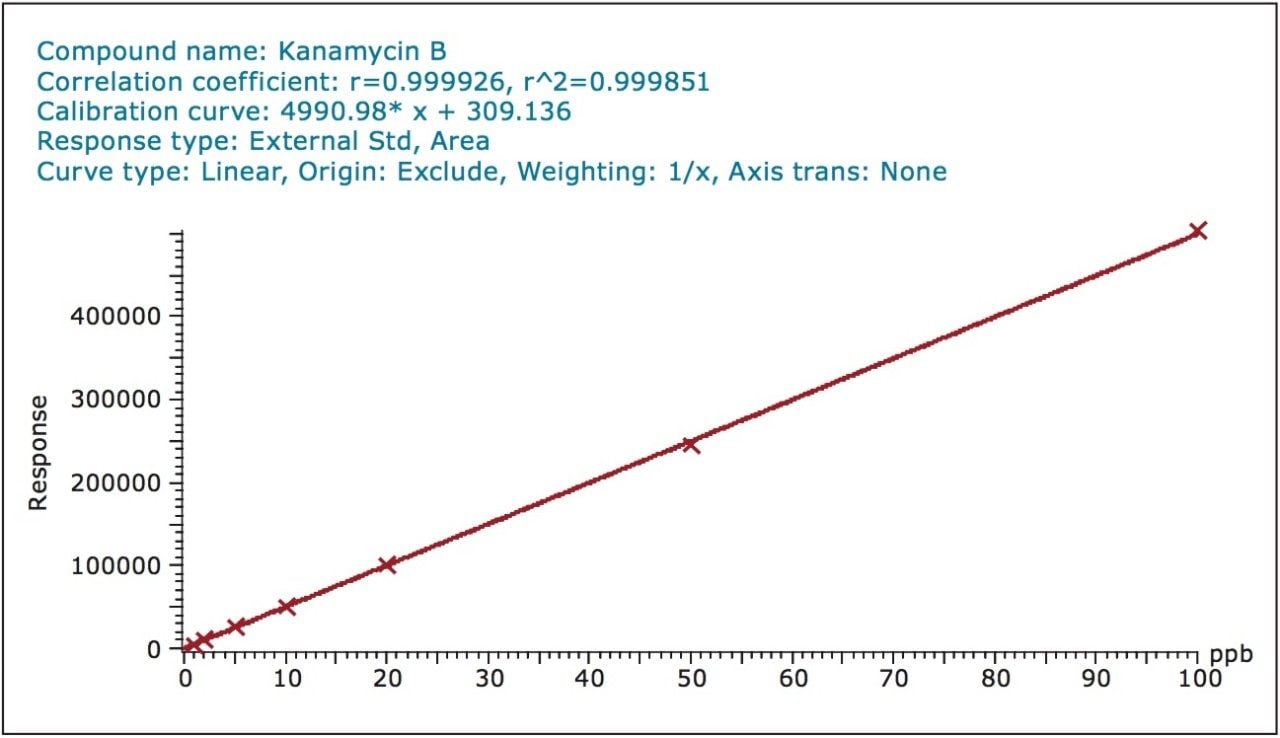
Matrix-matched standard calibration curves were prepared over the concentration range from 1 to 100 ug/kg. Excellent linearities (r2 > 0.999) were achieved for all compounds. An example of the calibration curve of an aminoglycoside residue (Kanamycin B) in bovine milk matrix is shown in Figure 2.
A multi-residue method for separation and analysis of aminoglycoside antibiotics in bovine milk using ion-pairing liquid chromatography on an ACQUITY UPLC PFP Column has been developed. The high-sensitivity of the ACQUITY UPLC System coupled with the Xevo TQ-S Mass Spectrometer enabled the detection, quantification, and confirmation of aminoglycoside residues in bovine milk to below the regulatory concentrations. The use of RADAR Technology allowed the matrix background to be monitored while providing MRM data for quantification – enabling informative decisions to be made during method development, ensuring a robust and reliable method.
720004330, May 2012