Liraglutide Native SEC-MALS Characterization Using Empower™ Chromatography Data System (CDS)
Udayabagya Halim
Waters Corporation, United States
Published on March 17, 2026
Abstract
In this application note, the use of native SEC-MALS, configured with a DAWN™ MALS photometer and an Optilab™ dRI Detector, for accurate molecular weight determination of low-molecular-weight peptides is demonstrated. The method highlights the ability to characterize peptides under native conditions with high sensitivity and precision. All data acquisition and analysis were performed using Empower CDS 3.10.0 Software, providing a fully integrated workflow for chromatographic control, MALS and dRI or TUV signal processing, and automated reporting.
Benefits
Non-destructive, calibration-free approach to determine absolute molecular weight.
Effective separation of higher-order oligomers without exclusion effects.
Accurate quantitation to avoid the bias introduced by UV detection for large particles.
Introduction
Size exclusion chromatography (SEC) is a widely used technique for analyzing peptide and protein size distributions and oligomeric states. SEC can be performed under denaturing conditions, which disrupt non-covalent interactions to reveal monomeric species, or under native conditions, which preserve the molecule’s natural conformation and non-covalent assemblies. While denaturing SEC provides insight into primary structure, native SEC is indispensable for studying aggregation, oligomerization, and higher-order structures as they exist in final drug product formulations.
Despite maintaining the final drug product with traditional native SEC methods, this technique often relies on calibration standards for molecular weight determination to distinguish the nature of oligomeric state that introduces error. A calibration-based SEC method lacks absolute accuracy for complex higher order structures or oligomers. This inherent bias be overcome by combining SEC with multi-angle light scattering (MALS) and differential refractive index (dRI) detection under native conditions for a non-destructive, calibration-free approach to determine absolute molecular weight and assess aggregation without altering the sample’s native state. MALS operates seamlessly under physiological and formulation-relevant conditions, including buffers that may contain non-volatile salts. This makes it a strong complement to techniques such as mass spectrometry, which often favor volatile buffer systems for optimal performance.
Experimental
Methods
|
Parameter |
Details |
|
System: |
Arc™ Premier HPLC System with quaternary solvent and sample manager |
|
Column: |
XBridge™ Premier Protein SEC Column, 250 Å pore size, 2.5 µm, 7.8 mm × 300 mm |
|
Mobile phase: |
10 mM Na2HPO4, pH 8.1 |
|
Flow rate: |
0.5 mL/min |
|
Detection: |
Arc Premier 2998 PDA UV Detector, DAWN MALS photometer, Optilab™ dRI Detector |
|
Software: |
Empower 3.10.0 CDS with advanced detector license option for MALS analysis |
|
dn/dc: |
0.185 mL/g for all samples under native conditions |
|
UV extinction coefficient: |
6990 M⁻¹ cm⁻¹ at 280 nm (1.86 (mg/mL)⁻¹ cm⁻¹) for liraglutide |
Accurate characterization of peptide oligomers and aggregates begins with careful selection of both the SEC column and detection strategy. Peptides are prone to forming oligomers, larger aggregates, and even fibrils that can exceed 20–50 nm in size.1,2 Under native conditions, columns with larger pore sizes are therefore required to effectively resolve oligomeric species, separate higher-order aggregates, and avoid size-exclusion effects.
In this study, an XBridge Premier Protein SEC Column with a 250 Å pore size was selected as to accommodate the higher-order oligomers present in liraglutide under native conditions. This pore size enables effective separation of oligomeric species while maintaining adequate recovery of large assemblies.
For concentration detection, the Optilab dRI Detector with dn/dc input of 0.1850 mL/g was used; traditional UV is another option for a concentration detector, but RI was used for two primary reasons. First, dRI detection provides flexibility across peptide development workflows, as it enables quantitation of peptides that lack aromatic residues and therefore exhibit weak or no UV absorbance. Second, prior work has shown that aged liraglutide drug product contains particulate species in the 20–30 nm size range that can interfere with UV-based concentration measurements.3 While UV detectors are commonly used for simple proteins and peptides, particles larger than ~20–30 nm can scatter light in addition to absorbing it, leading to overestimation of aggregate content and biased quantitation. The Optilab dRI Detector operates at a longer wavelength than alternative optical detectors and is less susceptible to light-scattering artifacts than UV detection, enabling more accurate determination of mass fractions.
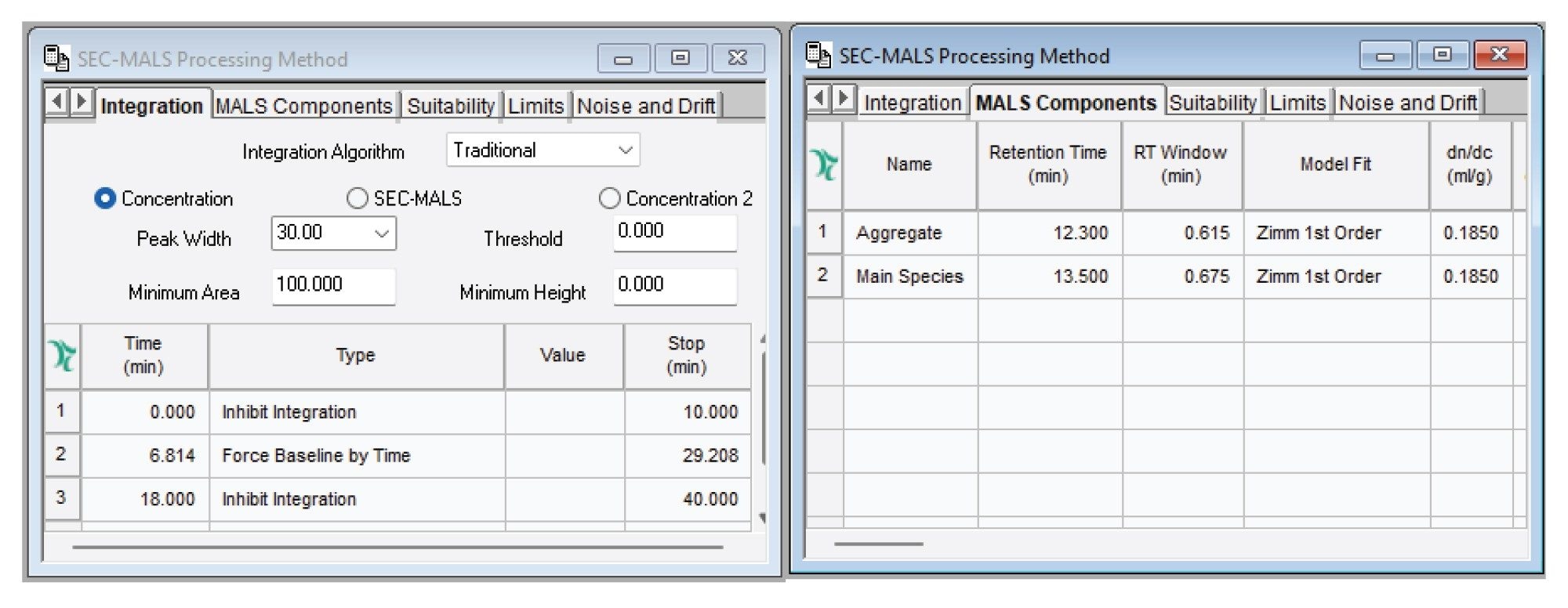
Data analysis was performed using Empower CDS 3.10.0 with the traditional algorithm for automatic peak detection and integration. This approach provides improved reproducibility of recovery and percentage mass calculations while simplifying method implementation, as it does not rely on fixed elution times and therefore reduces method development effort.
Although not part of this technical brief, the method for denaturing SEC-MALS analysis is described in a previously published application note.3 Whereas native SEC-MALS provides information on oligomerization and aggregation, denatured SEC-MALS using organic solvents separates GLP-1 RA molecules into individual monomers, providing information on covalently and more strongly bonded aggregates.
Results and Discussion
Native SEC-MALS analysis was performed on liraglutide drug substance (DS) at a concentration of 6 mg/mL under non-denaturing conditions. This concentration was selected to reflect the typical formulation of liraglutide drug product.3 The chromatogram exhibited two distinct peaks: a primary peak eluting at approximately 13 minutes and a secondary peak corresponding to higher-order oligomeric species.
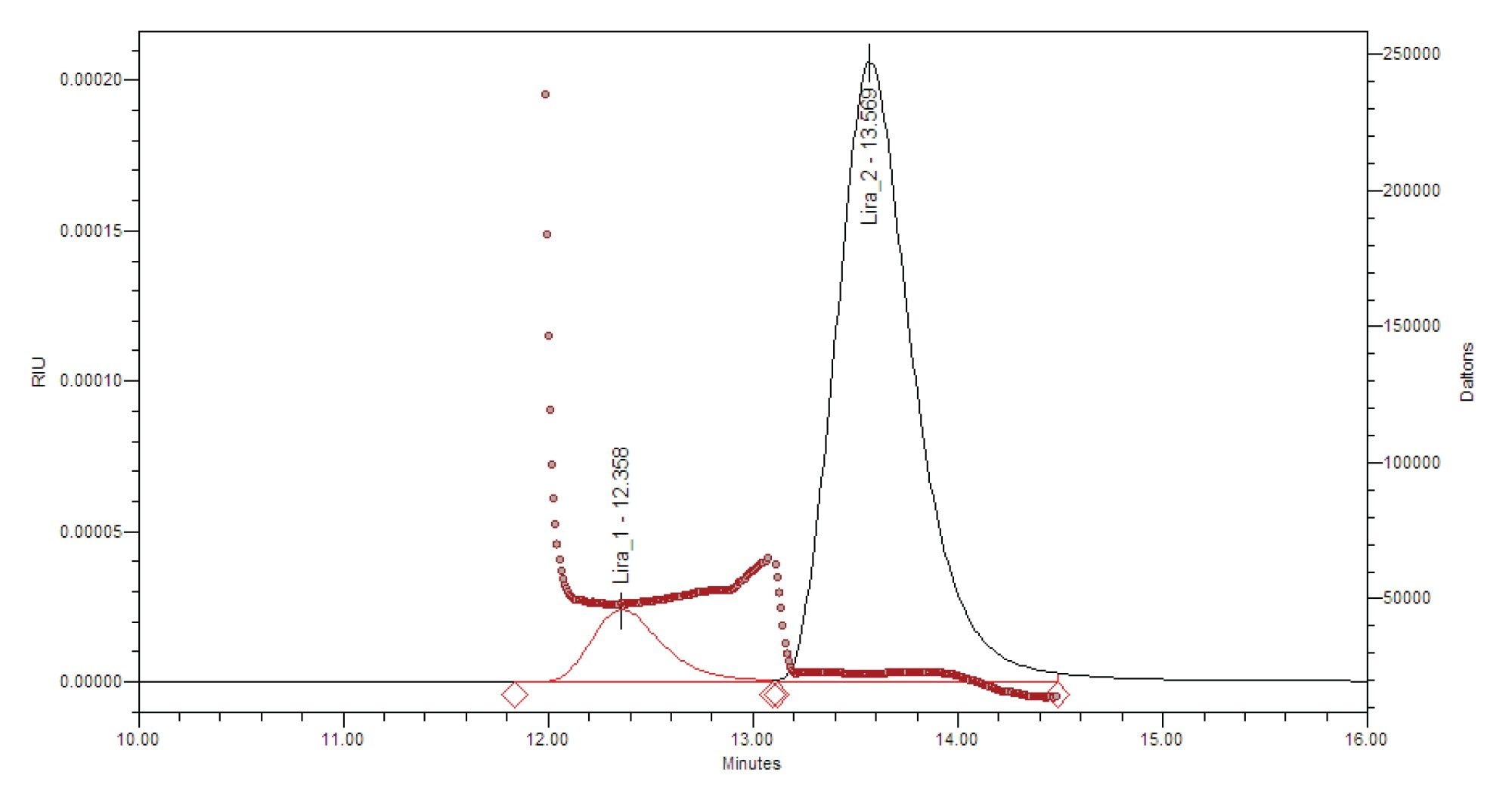
MALS analysis showed that the primary species accounted for 90.7% of the total mass, with a percent CV of 0.4%, and a weight-average molar mass (Mw)of 22.4 kDa with a %CV of 0.7% based on triplicate injections. This molar mass is consistent with a hexamer, given the theoretical monomer mass of 3.75 kDa. The secondary peak exhibited a molar mass of approximately 50 kDa, consistent with a 13-mer species. No peak corresponding to the monomeric form was detected under these native conditions, which is in agreement with previously published reports.3,4
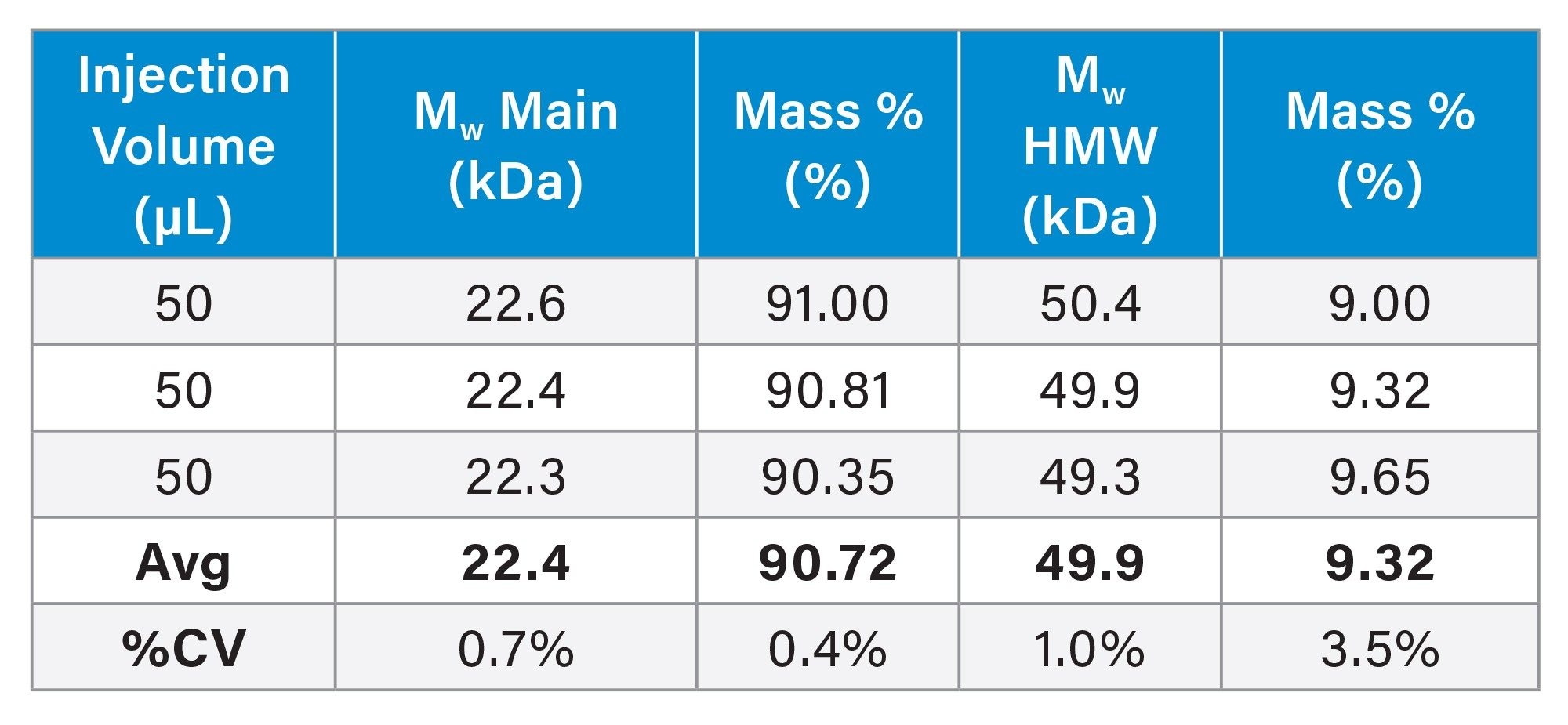
Conclusion
Native SEC-MALS analysis is essential for identifying the true oligomeric state of peptides and detecting larger aggregates under formulation-relevant conditions. Unlike denaturing workflows, native analysis requires careful selection of both columns and detectors. In this study, SEC-MALS using a DAWN MALS Detector provided absolute molecular weight determination, while the XBridge Premier Protein SEC Column (250 Å) enabled effective separation of higher-order oligomers without exclusion effects. For concentration measurement, the Optilab™ dRI Detector was employed to avoid the bias introduced by UV detection for large particles, ensuring accurate quantitation. Data processing was seamlessly performed within Empower CDS 3.10.0, providing an integrated and compliant workflow for peptide characterization.
References
- Brichtová, Eva Přáda, et al. "Glucagon-like peptide 1 aggregates into low-molecular-weight oligomers off-pathway to fibrillation." Biophysical journal 122.12 (2023): 2475-2488.
- Přáda Brichtová, Eva, et al. "Effect of Lipidation on the Structure, Oligomerization, and Aggregation of Glucagon-like Peptide 1." Bioconjugate Chemistry 36.3 (2025): 401-414.
- Zhang, Xujun, et al. “Identification of GLP-1 analog oligomeric states using SEC-MALS.” Application Note, Wyatt Technology Corporation
- Bothe, Jameson R., et al. "Peptide oligomerization memory effects and their impact on the physical stability of the GLP-1 agonist liraglutide." Molecular pharmaceutics 16.5 (2019): 2153-2161.
720009263, March 2026