
In this application note, we explore the feasibility of using the ACQUITY UPLC I-Class System in combination with ion mobility mass spectrometry (IMS-MS) as a new strategic approach for the screening of pesticide residues in food.
With increasing global trade there is a requirement for multi-analyte screening strategies capable of efficiently detecting residue violations in order to protect consumers. Countries have different regulations concerning licensing and Maximum Residue Limits (MRLs). It is essential that food commodities are screened efficiently and with confidence. Maintaining efficiency for pesticide residues analysis in food has become an increasingly challenging task, considering the increasing number of compounds to be monitored at low concentrations with generic extraction procedures. The analytical challenge to rapidly and efficiently identify target compounds is exacerbated because the target analytes can occur with a large number of co-extracted matrix components.
Enhancements in data processing software packages have enabled multiple screening parameters such as retention time, precursor, and fragment accurate mass to be built into scientific libraries. Data can be screened with tolerances that are compliant with the SANCO/12571/2013 guidance document.1 Advances in mass spectrometry technology have provided sensitivity increases; however this is the case for both matrix and target ions. The challenge reverts to removing false detections through careful optimization of software screening parameters, while ensuring that when dealing with the impact of such complex matrices false negative identifications do not occur.
Even with enhanced informatics-based solutions, it is still common for the resultant data to be interrogated manually. Manual data interrogation is labor intensive and time consuming. Consider the accepted mass accurracy tolerance of 5 ppm – this is not the value initially applied in many laboratories. The actual screening tolerances applied are realistically <10 ppm. Hence laboratories automatically screen using wider tolerance parameters than the guidelines suggest and they need to manually interrogate data to ensure that no false negative detections have occurred. Wider tolerances produce larger numbers of false detections which impact laboratory efficiency. In addition, as previously described, retention times can also vary and be affected by matrix shifts, column loading, as well as system setup.2 In order to efficiently deploy high resolution accurate mass spectrometry for routine contaminant screening, a workflow that removes the need for manual interrogation is essential.
At MRL levels and below, it can be a challenge to obtain isotope fits, fragment ions, and ion ratios for all pesticides. This is often due to matrix suppression, but it may also be due to poor ionization efficiencies of some compounds, and the tendency of certain compounds to form adducts. If sodium adducts form, it is unlikely that characteristic fragmentation will be obtained. Sodium and potassium are ubiquitous and originate from the matrix or from the glassware used. Pesticides that contain carboxyl or carbonyl ether or ester groups are likely to bind alkali metal ions such as sodium or potassium. They form very stable species and usually do not fragment. This may have the consequence of removing a potential identification point (in the form of fragment ions) if the protonated form is not also present to provide the opportunity to detect fragment ions.
A strategy where further information could be provided for more confident identifications would be advantageous. This additional information could work in conjunction with accurate mass and fragment ions, when retention time shifts occur, when a monoisotopic peak has been identified, or adducting has prevented fragmentation data being obtained. At the MRL, confidence in screening assays could also be improved if the matrix and analyte could be routinely deconvoluted. This would allow better visibility of the fragmentation information generated and remove the matrix interferences from low level isotope distributions. This strategy can be employed using the combination of UPLC separations and orthogonal ion mobility with accurate mass MS, where very specific retention time aligned and mobility aligned data is produced.
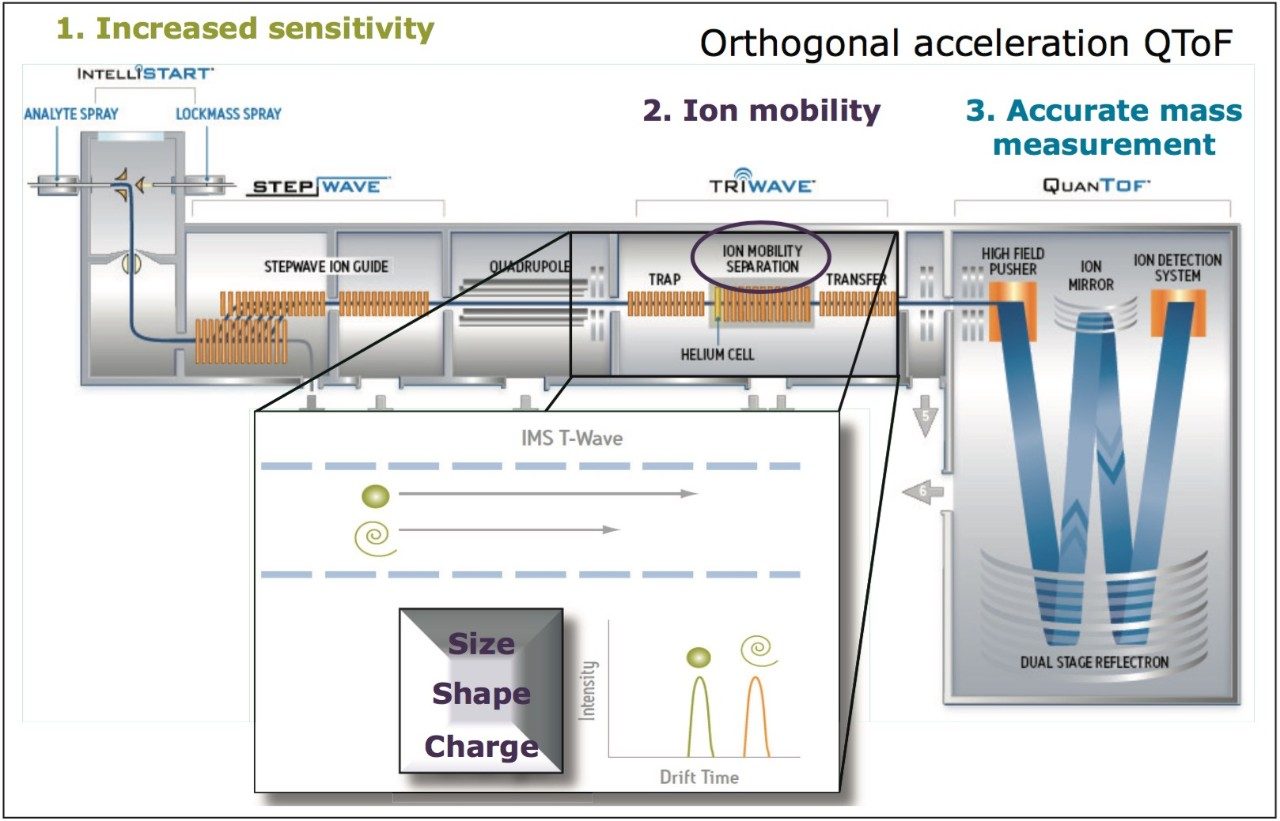
In this application note, we explore the feasibility of using the ACQUITY UPLC I-Class System in combination with ion mobility mass spectrometry (IMS-MS) as a new strategic approach for the screening of pesticide residues in food. This technique offers some unique advantages for profiling complex matrices. Ion mobility provides an orthogonal separation to UPLC, where compounds can be differentiated based on size, shape, and charge. Collision cross section (CCS) values can be routinely generated to provide additional identifying information. CCS information requires fewer stringent screening parameters, and offers the confidence of avoiding false negative identifications. The orthogonal separation allows cleaned up precursor and fragment ion spectra to be obtained. In addition, both precursor ion and fragment ion information can be acquired in a single injection on all components, with the fastest UPLC compatible ion mobility duty cycle available (10.8 ms).
The study undertaken uses a Waters SYNAPT G2 HDMS platform, for which a schematic representation and illustration of the mechanism of ion mobility are shown in Figure 1. CCS measurements can provide a route to specific and unambiguous identification, enabling the unequivocal distinction of target residues. The application of ion mobility to remove false positive identifications and more importantly, false negative identifications, spectral cleanup, and increased identification confidence will be discussed.
10 g of homogenized sample was extracted with 60 mL of 20-mM ammonium acetate in methanol using an Ultra-Turrax device. The crude extract was filtered and diluted up to 100 mL with 5-mM ammonium acetate in water before injection.
An organic mandarin sample was used to produce a matrix matched calibration curve and a European Proficiency Test FV-13 sample was analyzed (European Commission proficiency tests for pesticide residues in fruits and vegetables. FV-13 Mandarin Homogenate, 2011).
|
System: |
ACQUITY UPLC I-Class |
|
Column: |
ACQUITY UPLC BEH C18 2.1 x 100 mm, 1.7 μm |
|
Column temp.: |
45 °C |
|
Flow rate: |
0.45 mL/min |
|
Mobile phase A: |
Water (0.1% formic acid) |
|
Mobile phase B: |
Acetonitrile (0.1% formic acid) |
|
Injection volume: |
5 μL |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
|---|---|---|---|
|
0.00 |
0.45 |
98.0 |
2.0 |
|
0.25 |
0.45 |
98.0 |
2.0 |
|
12.25 |
0.45 |
1.0 |
99.0 |
|
13.00 |
0.45 |
1.0 |
99.0 |
|
13.01 |
0.45 |
98.0 |
2.0 |
|
13.00 |
0.45 |
98.0 |
2.0 |
|
17.00 |
0.45 |
98.0 |
2.0 |
|
System: |
SYNAPT G2-S |
|
Ionization mode: |
ESI+ |
|
Desolvation temp.: |
550 °C |
|
Mass range: |
50 to 1200 Da |
|
Acquisition rate: |
5 spectra/sec |
|
Capillary voltage: |
1 kV |
|
Cone voltage: |
20 V |
|
Drift gas: |
N2 |
|
Collision energy ramp: |
10 to 45 eV |
|
IMS wave velocity range: |
650 m/s |
|
IMS wave height: |
40 V |
|
IMS gas flow: |
90 mL/min |
|
IMS duty cycle: |
10.8 ms |
|
Lock mass: |
m/z 556.2766 (Leucine enkephalin) |
It has previously been demonstrated that CCS values that were generated from pesticide standards can be utilized as a confirmatory parameter to increase confidence in identification, whilst reducing false detections.2 Estimated CCS values, precursor ions, fragment ions, and retention time values were determined for pesticide standards. These values were entered into the UNIFI Scientific Library. Subsequent non-targeted data sets were acquired for pear, ginger, leek, and mandarin matrix matched calibration series and European Proficiency Test FV-13 samples. CCS values have been utilized to reduce false detects and avoid false negative identifications in the EU RL proficiency test samples and matrix matched calibrant series analyzed. The robustness of the CCS screening strategy was tested on a blank matrix in order to assess the efficiency of the workflow for removing false positives.
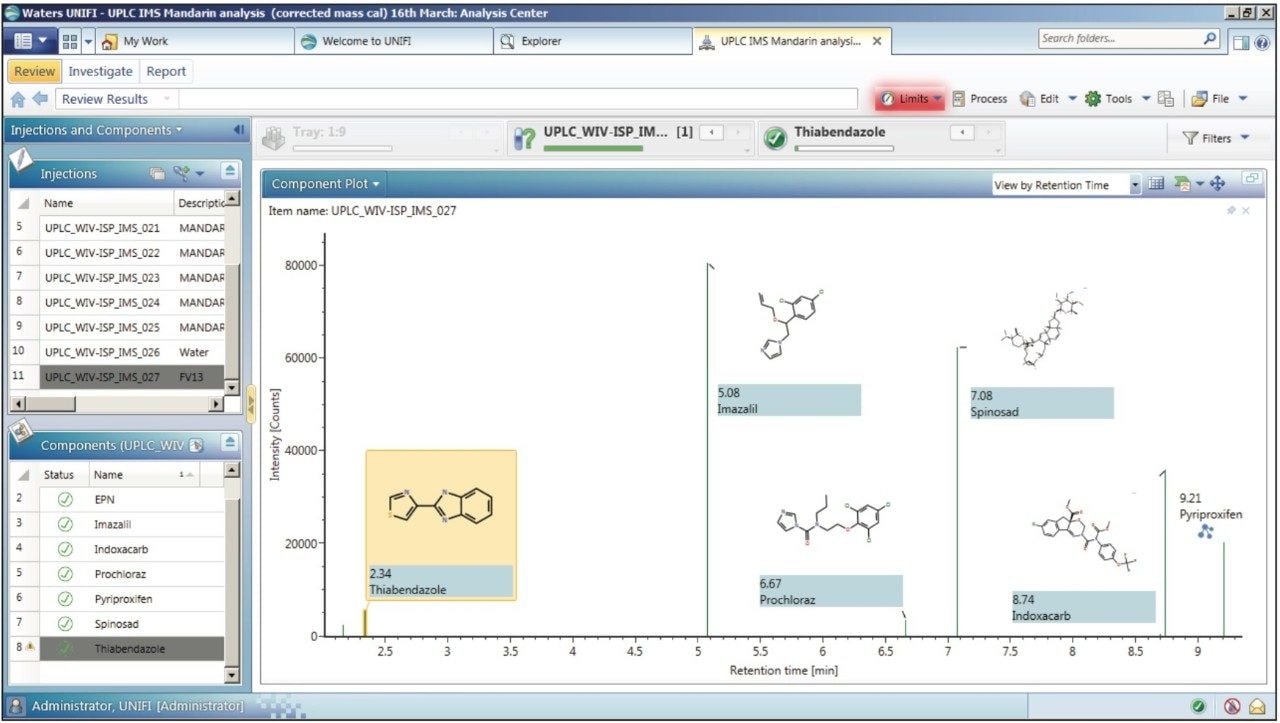
The previously described CCS screening process3 mimics the wider tolerance parameters that are applied in some laboratories for which time-consuming manual interrogation is required. The UNIFI component plot summary for the pesticide residues identified in the European Proficiency Test FV-13 sample is presented in Figure 2. The CCS screening component summary shows that 100% of the detected compounds were identified correctly and presented.
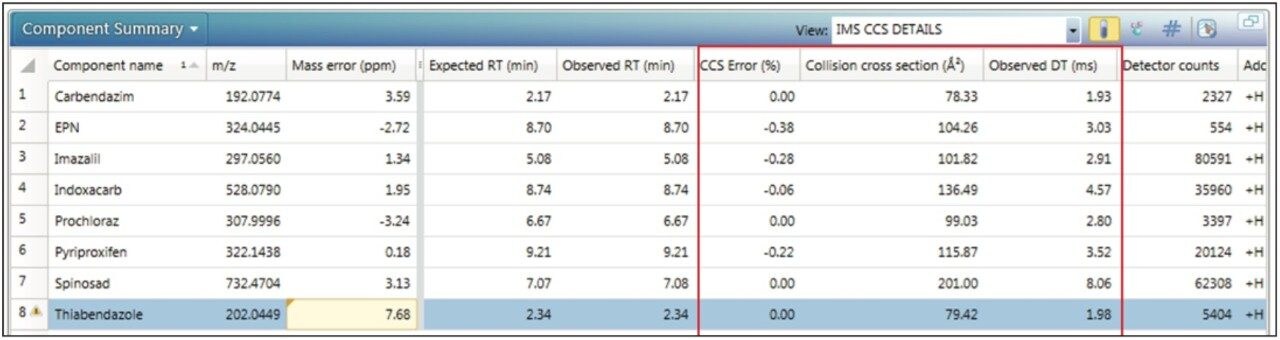
Using the functionality within UNIFI, it is possible to set tolerances that illustrate where exceedance has occurred. For the targeted screen of proficiency sample FV-13, a component summary is presented in Figure 3. Mass accuracy tolerance limits have been set to display a warning if the observed mass accuracy is within 10 ppm but >5 ppm. The expected CCS, observed CCS, and CCS error are shown in Figure 3, in addition to the conventional screening parameters of mass error (ppm) and retention time (Rt). From the results obtained it is clear that CCS provides an orthogonal measurement that gives added confidence in the identifications made. Using the CCS screening strategy as a part of a routine workflow, wider tolerance parameters can be used, which avoids false negatives without increasing false positive identifications, to improve overall screening efficiency.
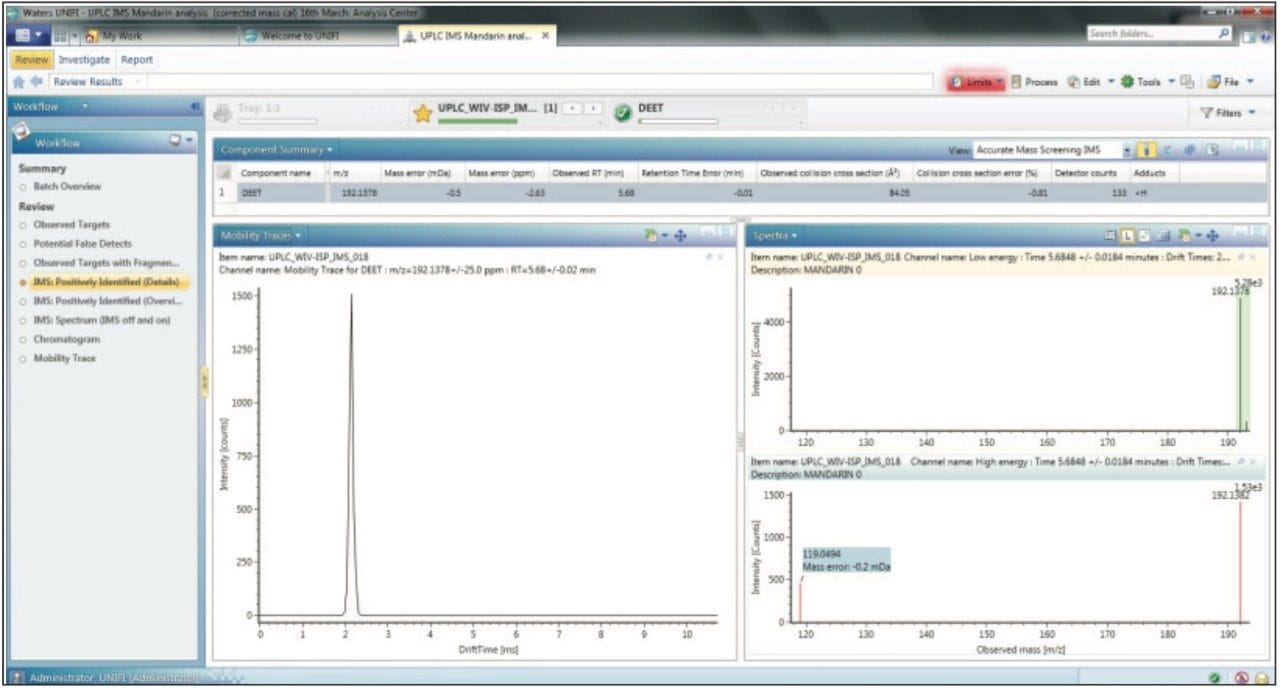
To avoid high false detection rates from an automatic screening process, different parameters and related criteria that can affect peak designation should be carefully set. Analysis of a non fortified blank matrix presents a very effective way of testing the CCS screening workflow within UNIFI. A blank mandarin sample was injected and subjected to automatic processing. More than 10,000 peaks were detected, but by employing the first CCS screening workflow step (20 ppm m/z, ± 0.5 min Rt, 10% CCS error), only 20 pesticides were reported. After applying a mass accuracy tolerance of <10 ppm and a CCS error of 2%, DEET was the only pesticide reported, as shown in Figure 4. DEET is commonly used as an insect repellent and it is often found as background contamination in solvents. DEET ionizes very efficiently giving a high response at very low concentration, and it is often detected at concentrations of <1 ppb in methanol used for sample preparation. Therefore, this finding is accurate and cannot be considered as an erroneous result. In addition to a matched Rt and mass measure measurement error of -2.63 ppm, a CCS error of -0.81% was obtained. The CCS screening parameter settings proved to be very effective, providing a very high level of confidence and no false detects for the non-spiked sample. Consequently we were even more confident about any positive detections that were observed, even where only the monoisotopic mass had been detected.
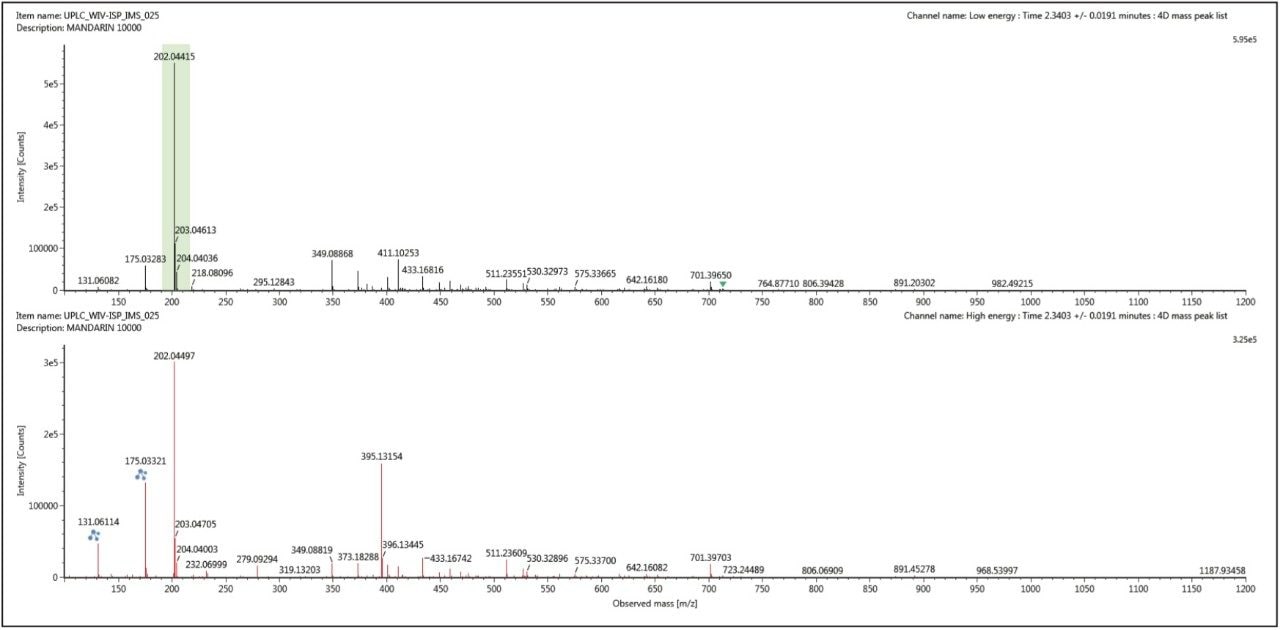
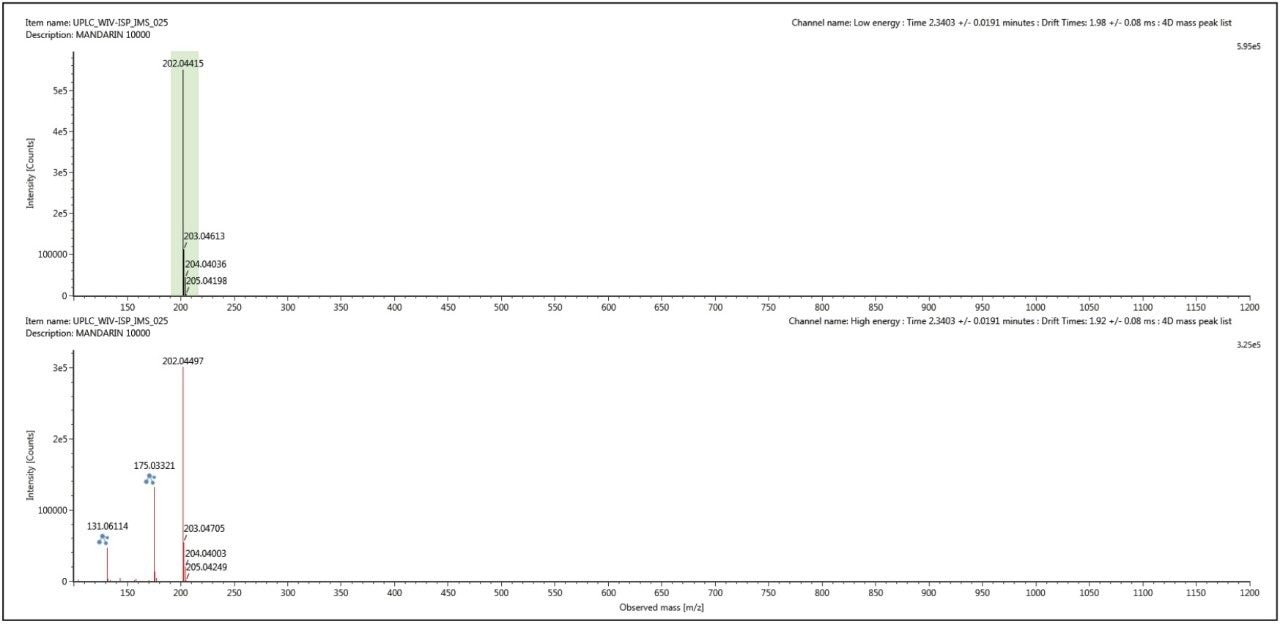
The conventional retention time aligned precursor and fragment ion spectra for thiabendazole (m/z 202) at retention time 2.34 mins, shown in Figure 5, was generated using UNIFI’s 3D peak detection algorithm, which produce cleaner spectra compared to other peak detection algorithms. However this spectra can be cleaned up even further when ion mobility acquisition has been performed. Using the ion mobility Resolve functionality, the multi-component spectrum was completely cleaned up using UNIFI’s 4D peak detection algorithm. The precursor ion and fragmentation spectra were retention time aligned and ion mobility aligned. The ion mobility cleaned up spectra for thiabendazole is presented in Figure 6. This is an extremely useful analytical tool, since the analyst will need to elucidate and confirm the identified pesticide species. The resolution provided by ion mobility elicits specific precursor and fragmentation ion information for the targeted species identified. This highly specific data was acquired for all components (in this case >10,000) when the analysis is performed on the SYNAPT G2 HDMS ion mobility platform – unlike Field Asymmetric Waveform Ion Mobility Spectrometry (FAIMS), which requires defined specific targets and the duty cycle limits spectral cleanup to just a few components.
720005080, June 2014