This is an Application Brief and does not contain a detailed Experimental section.
In this study HDMS Compare Software together with ion mobility mass spectrometry (IMS) has been evaluated as a comparative analytical approach to highlight and detect differences in batches of D-α-tocopheryl polyethylene glycol 1000 succinate (TPGS) produced by a number of suppliers.
HDMS Compare Software provides a rapid and powerful comparative approach highlighting differences in complex samples.
TPGS is an amphiphilic polymer frequently used in drug formulations as a water-soluble alternative of vitamin E. As a non-ionic surfactant, TPGS has drawn special attention in the drug industry as an emulsifier, stabilizer, absorption/permeation enhancer, and a drug solubilizer approved by the FDA. The structure is shown in Figure 1. It is crucial that the material is of suitable and consistent quality so bio-availability is not compromised.
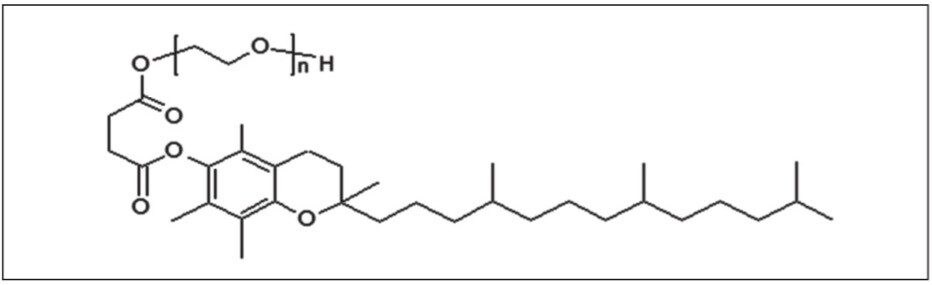
Different analytical techniques have been used to analyse TPGS, such as MALDI/MS, HPLC, SEC, and FTICR/MS. Due to the complex nature of the data, rapid comparison between samples is difficult.
IMS can be used to enhance the analysis of complex mixtures. It separates ionic species based on their charge and rotationally averaged collision cross-section (shape and size). The combination of ion mobility with high resolution MS significantly enhances routine MS and tandem MS workflows, which can aid structural elucidation.
Data processing can often create bottlenecks due to the high amount of data generated. The high definition mass spectrometry (HDMS) data sets generated using ion mobility MS are often complex and, until recently, difficult to compare two samples. HDMS Compare Software is a powerful tool, which is dedicated to IMS data and assists in the detection of differences in specific analytes by automated data comparisons and reviews pairs of HDMS data sets.1
In this study, a SYNAPT G2-S ion mobility MS and HDMS Compare Software have been evaluated as a comparative analytical approach to highlight and detect differences in three different batches of TPGS produced by a number of different suppliers.
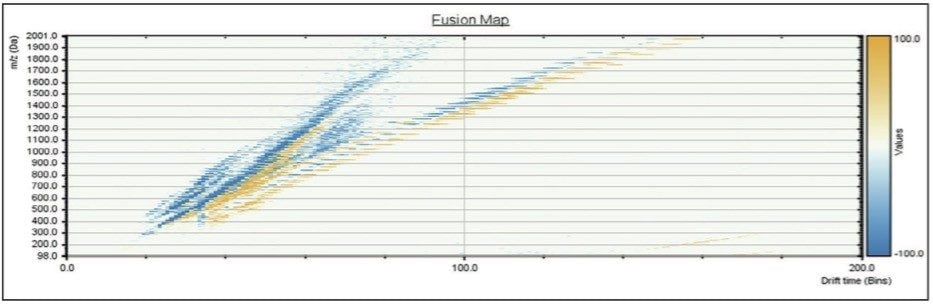
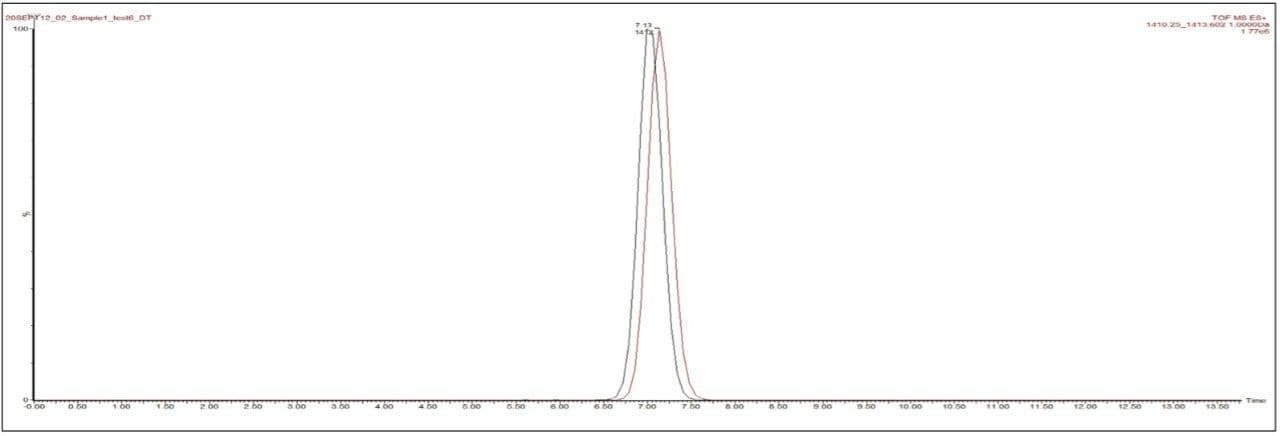
The IMS data was processed using HDMS Compare Software. Figure 2 shows the initial fusion map from the comparison of two batches of TPGS A and B. The orange regions in the mobility image indicate ions that are more significant in sample A and the blue regions indicate ions more significant in sample B. There is an option at this point to carry out further interrogation of the data within the HDMS Compare Software or export the data into MassLynx Mass Spectrometry Software. In this study the region between m/z 1200–1600 was selected and exported into MassLynx for further processing. A comparison of the mobility drift times for the 20-mer of m/z 1411 of TPGS showed that the two ions displayed different mobility drift times. The 20-mer from batch A had a reproducible drift time of 6.99 ms and batch B showed 7.13 ms, as shown in Figure 3. The different ion mobilities indicate a difference between the size and/or shape of the two 20-mers.
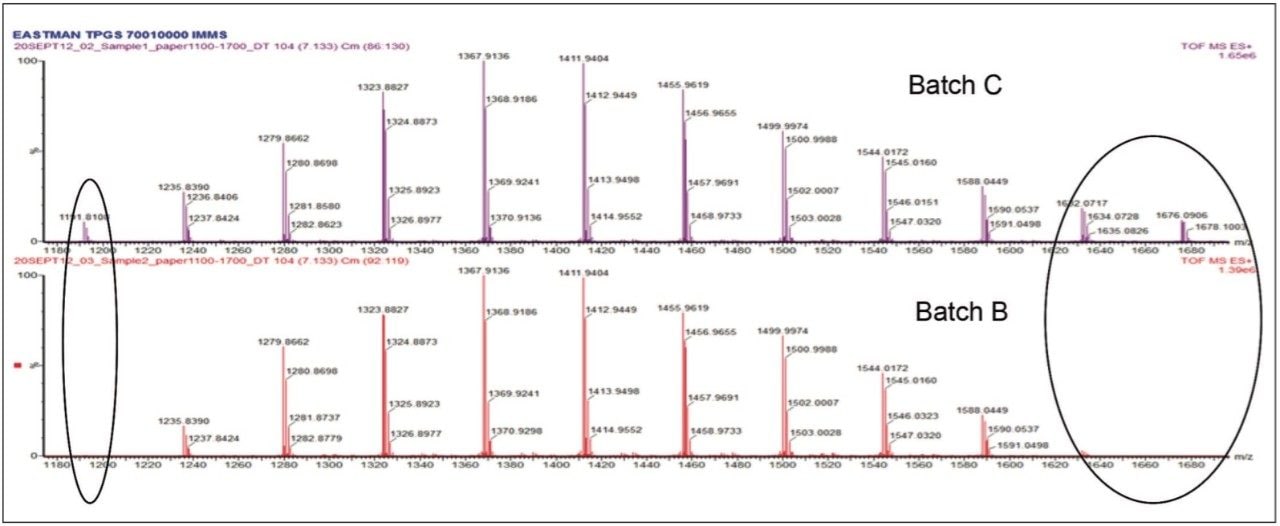
Comparison of batch B with batch C of TPGS also showed differences in the fusion map. Comparison of the mobility drift times for the 20 mer showed the two ions displaying the same drift times of 7.13 ms. The mass spectra from the region between m/z 1150 to 1700 for both batches (illustrated in Figure 4) showed that the abundance of the ions at m/z 1191, 1632, and 1676 is more significant in batch C, potentially impacting the physical properties of the TPGS.
TPGS is a complex polymer. Although analysis can be achieved by a number of techniques, rapid comparison has not been readily successful. In this study the use of ion mobility mass spectrometry and HDMS Compare Software has allowed batch to batch variations in the complex polymeric material to be readily observed. HDMS Compare Software highlights differences in batches of TPGS produced by a number of different suppliers, which allows important analytical decisions to be achieved in a more timely manner.
The authors would like to acknowledge the help of Kieran Neeson, Richard Denny and Kirsten Craven.
720005211, October 2014