This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates the benefits of UPLC separations in HILIC mode for the glycans’ linkage analysis of the released 2-AB labeled N-linked glycan pool of fetuin, sequentially digested by exoglycosidase array.
In glycan analysis, UPLC delivers a clear advantage for resolving coeluting positional isomers, which are often difficult to separate with conventional HPLC.
Many biotherapeutics are glycosylated. Glycoprotein characterization includes glycan profiling, which is an important step in the process of development and production of biopharmaceutical proteins.
The released glycan pool is of a great complexity due to branching and linkage isomers. Such structural heterogeneity of glycans is attributed to their biological role and requires thorough characterization for glycobiology applications and regulatory purposes.
At the same time, glycan complexity often hinders complete structural elucidation by conventional HPLC methods since many species remain unresolved. Entire oligosaccharide characterization is incomplete if structural or linkage isomers cannot be closely monitored in order to maintain consistency and high quality during therapeutic manufacturing.
UltraPerformance Liquid Chromatography (UPLC) in hydrophilic interaction chromatography (HILIC) separation mode has become a routine and widely-recognized technique for rapid, efficient, sensitive, and reproducible analysis of 2-aminobenzamide (2-AB) labeled glycans. UPLC Glycan columns packed with sub-2-μm particle sorbent offer significant improvements in separation power and allow enhanced resolution of complex glycans.
To compare resolving capabilities of HPLC and UPLC systems, a polysaccharide ladder of the released bovine fetuin N-glycan was created by a sequential digestion using enzyme array (Figure 1). Each monosaccharide was released based on the enzyme specificity (Figure 2).
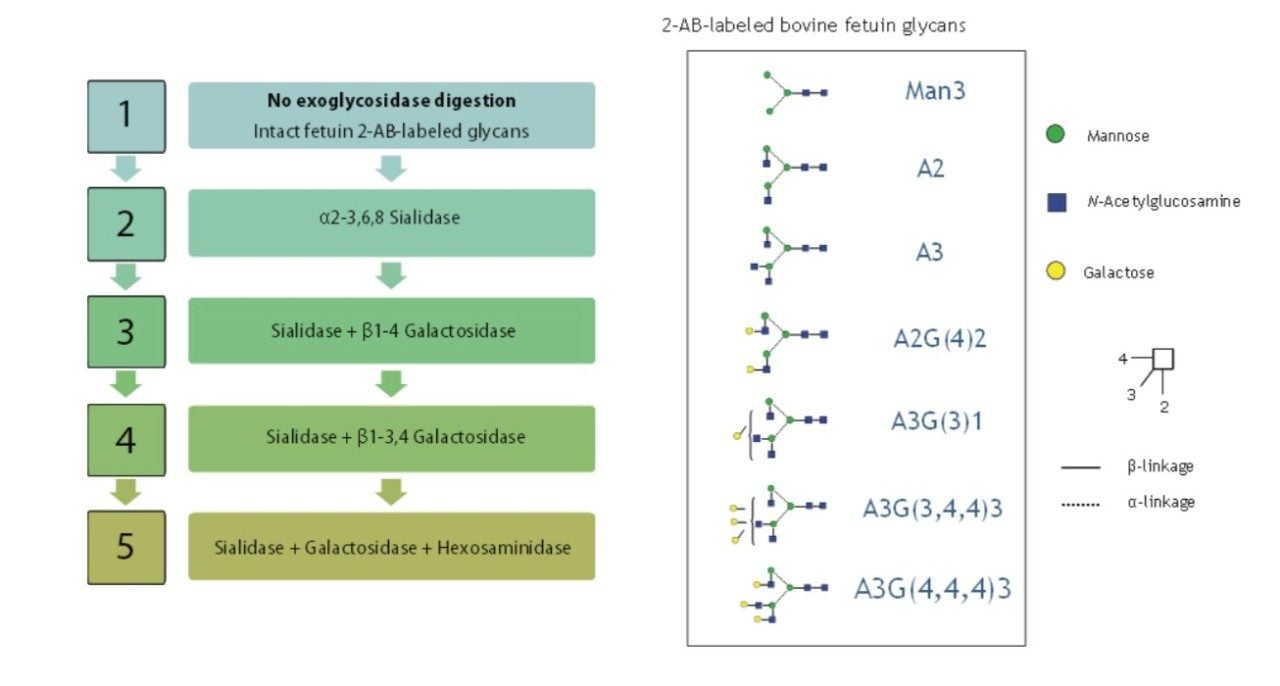
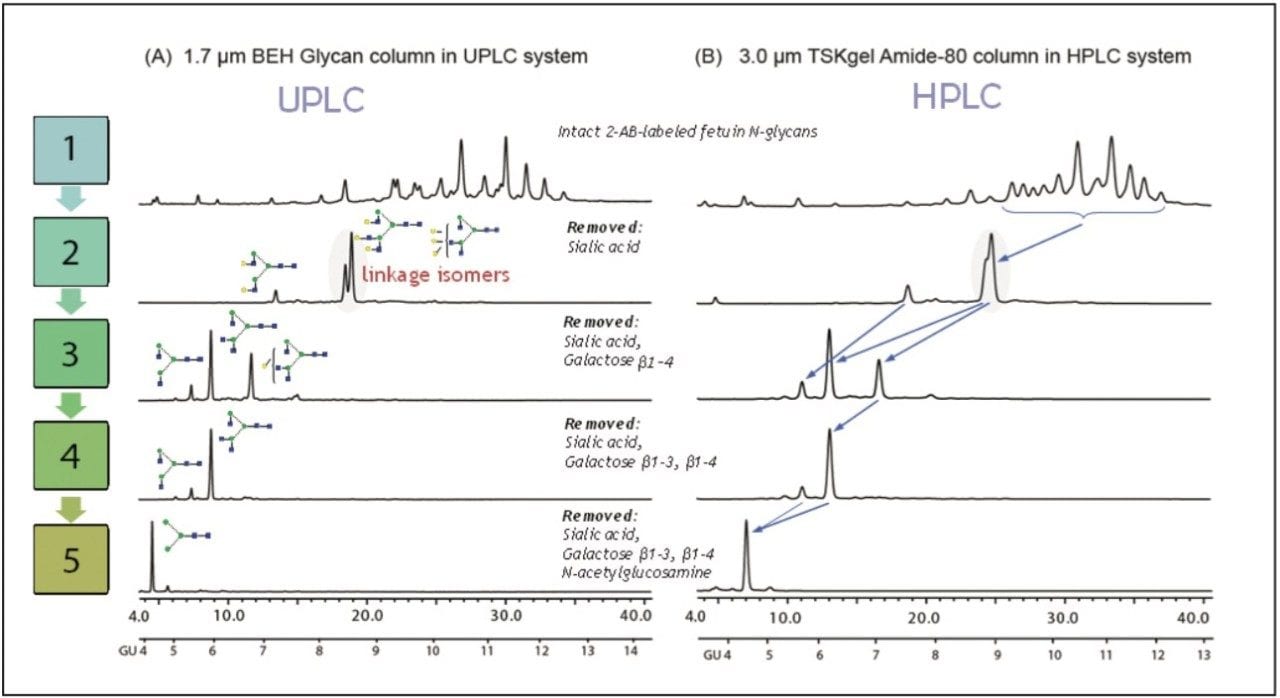
LC experiments were run on a Waters ACQUITY UPLC System with fluorescence detection (FLR) and an ACQUITY UPLC BEH Glycan Column (2.1 x 150 mm, 1.7 μm). The Waters Alliance 2695 HPLC System with a TOSOH Bioscience TSKgel Amide-80 Column (2.0 x 150 mm, 3.0 μm) was used as the default HPLC method. The structures of the digested glycans were confirmed by glucose units (GU) values and specificity of the applied enzyme.
The top chromatograms represent the undigested total N-glycan pool, and the remaining chromatograms correspond to the sequential digestion of the N-glycan pool with the exoglycosidase enzymes indicated.
The first enzyme applied was Arthrobacter Ureafaciens sialidase, which releases sialic acids. This step yields two linkage isomers A3G(3,4,4)3 and A3G(4,4,4)3, which are hard to resolve under HPLC conditions.
As highlighted on the chromatogram, these 2-AB labeled isomers were separated using the 1.7-μm ACQUITY UPLC BEH Glycan Column on the UPLC system, but coeluted on the 3.0-μm HPLC column. Such an improvement in resolution is due to the higher peak capacity of sub-2-μm particle packing.
Method transfer between HPLC and UPLC is straightforward, since selectivity of neutral and siaylated glycans in both columns are comparable under the conditions used for this study.
Side-by-side comparisons of UPLC and HPLC methods of the fetuin N-glycan digest, obtained by exoglycosidase array, illustrates the suitability of UPLC separations for glycan identification. The results demonstrate UPLC’s clear advantage for resolving coeluting positional isomers, which are often difficult to separate with a conventional HPLC column and system.
This highly efficient and fast UPLC separation of glycans meets regulatory obligations as a part of complete glycoprotein characterization, and conveniently offers users a seamless HPLC-to-UPLC method transfer.
720003865, January 2011