This is an Application Brief and does not contain a detailed Experimental section.
Quantitative proteomics involves the identification and quantitation of proteins from biological systems. This application brief shows results from protein mixtures spiked at different concentrations into an E. coli cytosolic lysate.
Xevo G2 QTof provides quantitative measurements of proteins from complex mixtures with wide dynamic range.
Studies of differential protein expression by LC-MS are conducted in many laboratories to gain a better understanding of biological processes. For the identification of proteins, the use of time-of-flight systems is well accepted, and there are advantages in terms of mass accuracy and resolution. However, obtaining good quantitative measurements from complex mixtures with wide dynamic range challenges the entire analytical system from sample preparation to bioinformatics. The label-free LC-MSE approach avoids the cost and complexity of stable isotope labeling and provides a robust method for making quantitative measurements. In this technical brief we demonstrate the relative quantification of known standard proteins contained in a four protein mixture spiked at two different levels into a complex sample matrix.
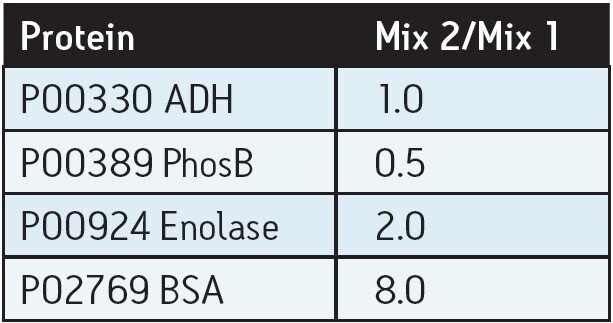
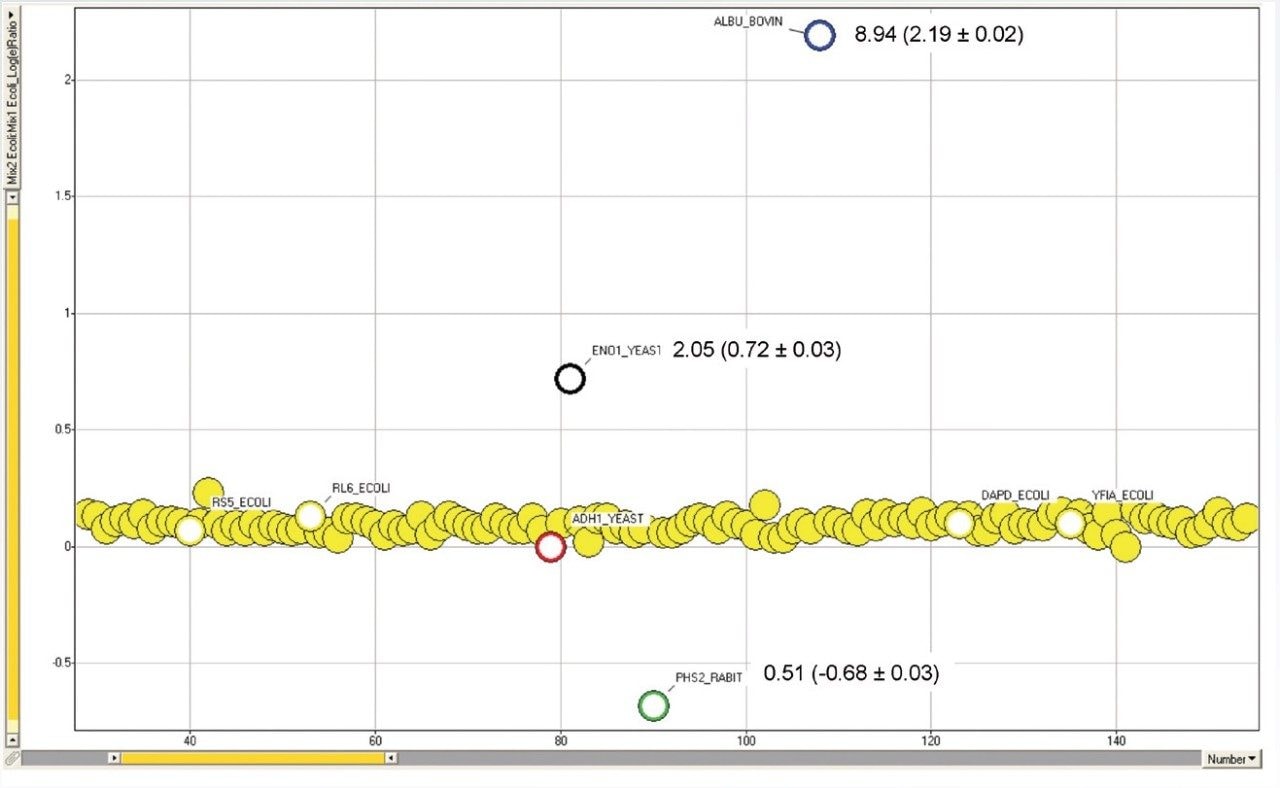
Standard protein mixtures containing the four proteins, as shown in Table 1, were spiked at an ADH concentration of 25 fmol into a 400 ng sample of digested E. coli cell lysate. A Waters nanoACQUITY UPLC System coupled with a Xevo G2 QTof were used to separate and analyze the tryptic peptides in triplicate. The Xevo G2 QTof was operated in LC-MSE mode whereby the collision energy was switched between low and elevated energy during alternate scans. Peptide precursor information was acquired in the first function (used for quantitation) and fragment ion information was acquired in the second function (used for protein identification). Using ProteinLynx Global SERVER, data were processed and searched against a non-redundant E. coli database, with the four known protein sequences added. By selecting ADH as the internal standard, relative ratios for each spiked protein were determined.
Here we have shown relative quantification of four standard proteins spiked into a complex matrix using the Xevo G2 QTof Mass Spectrometer. Good correlation was achieved between the expected and experimentally determined values.
![Spectrum at peak top (left) and chromatogram of the [M+2H]2+ ion for the VIFLENYR peptide.](/content/dam/waters/en/app-notes/2010/720003483/720003483en-f2.jpg.82.resize/img.jpg)
720003483, May 2010