
The complexity of trastuzumab, and biotherapeutic proteins in general, require a multifaceted and flexible approach for their characterization and quantification. Highly sensitive direct quantitation at the intact level was achieved using a Waters Vion IMS QTof and UNIFI.
The present data suggests that HRMS is capable of providing highly sensitive intact level quantification of biotherapeutic proteins. Data derived from the intact level analysis will complement those from ligand binding assays or LC-MS/MS based surrogate peptide approaches for a complete understanding of the proteins in question.
Biotherapeutics have emerged as one of the major sources of innovative medicines in drug discovery and development. As these biotherapeutic agents have complex structure and extremely high molecular weight, their detection and quantification can be challenging especially for the bioanalysis community that traditionally focuses on small molecules. The most common approach for quantifying these biotherapeutics is ligand-binding based assays complimented by LC-MS/MS analysis of surrogate peptides post protein digestion and sample cleanup. More recently, direct intact level protein quantification using LC-MS has gained traction as it offers direct measurement of proteins, including isoforms, and simpler sample preparation.
This application note describes a method for highly sensitive intact level quantification of a 150 kDa therapeutic protein, trastuzumab, in BSA solution capable of achieving an LLOQ of 3 ng/mL (30 pg load on column). The development and evaluation of peak based quantification is described in detail, from acquistion through processing. It provides detailed instruction for data processing options using UNIFI (using the Quantify Assay 2D Tof analysis method), including the impact on quantification of using different mass tolerance windows and numbers of peaks for integration of signal. Quantification attributes including: linearity, dynamic range, accuracy, limits of detection, and quantification are reported.
The analytical LC-MS experiments were performed on a Waters ACQUITY UPLC I-Class System and a Vion IMS QTof Mass Spectrometer. UNIFI Scientific Information System was used for data acquisition and processing.
|
LC system: |
ACQUITY UPLC I-Class |
|
Column: |
Polystyrene divinylbenzene based, 4 μm, 1500 Å, 3 x 50 mm |
|
Column temp.: |
80 °C |
|
Sample temp.: |
10 °C |
|
Injection volume: |
10 μL |
|
Flow rate: |
0.2 mL/min |
|
Mobile phase A: |
Water with 0.1% formic acid |
|
Mobile phase B: |
Acetonitrile with 0.1% formic acid |
|
Time (min) |
Flow rate (mL/min) |
Composition A (%) |
Composition B (%) |
Curve |
|---|---|---|---|---|
|
0 |
0.2 |
98 |
2 |
6 |
|
1 |
0.2 |
98 |
2 |
6 |
|
2 |
0.2 |
75 |
25 |
6 |
|
7 |
0.2 |
50 |
50 |
6 |
|
7.5 |
0.2 |
10 |
90 |
6 |
|
8 |
0.2 |
98 |
2 |
6 |
|
8.5 |
0.2 |
10 |
90 |
6 |
|
9 |
0.2 |
98 |
2 |
6 |
|
11 |
0.2 |
98 |
2 |
6 |
|
MS system: |
Vion IMS QTof |
|
Ionization mode: |
ESI+ |
|
Acquisition mass range: |
500–4000 m/z |
|
Experiment: |
MS full scan, sensitivity mode |
|
Capillary voltage: |
2.75 kV |
|
Cone voltage: |
150 V |
|
Cone gas flow: |
0 L/h |
|
Source temp.: |
150 °C |
|
Source offset: |
150 V |
|
Desolvation gas temp.: |
600 °C |
|
Desolvation gas flow: |
800 L/h |
|
Scan time: |
1 s |
Trastuzumab solution at 20 mg/mL in formulation media (source: Roche) was serially diluted using 0.1% formic acid (FA) containing 0.1 mg/mL BSA aqueous solution to a concentration range of 0.003 µg/mL (3 ng/mL) to 100 µg/mL. Polypropylene vials were used (p/n: 186005219) for the dilution.
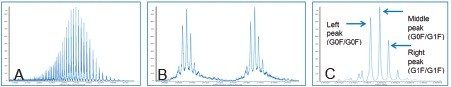
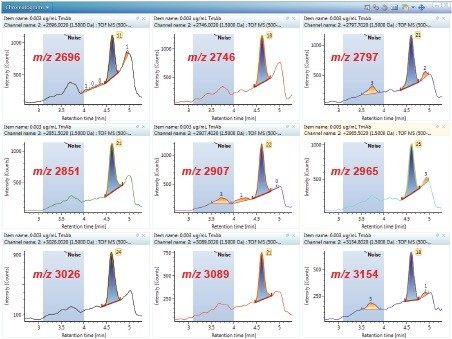
Trastuzumab, sold under the brand name Herceptin, is a monoclonal antibody, which was approved by the FDA in 1998 for the treatment of breast cancer (HER2 receptor positive). It has a molecular weight of around 150 kDa. Under typical ESI+ MS conditions, trastuzumab exhibits a mass/charge envelope distribution typical for biotherupeutic proteins (Figure 1A). For a 100 µg/mL solution, the multiply charged species has an m/z of 2000–3500 with a plateau at around 52+ to 55+ charge states, centered around 2600 m/z at concentration. Within each charge state, there are multiple peaks representing the distribution of various glycosylation forms (Figure 1B). The average mass of the molecule could be obtained through MaxEnt1 deconvolution as shown in Figure 1C. The deconvoluted spectrum is also useful for structural confirmation.
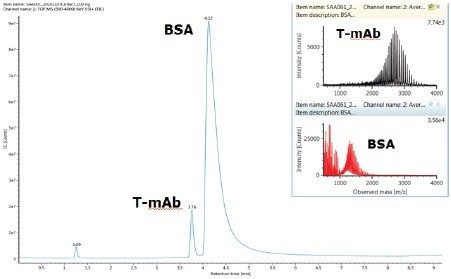
Under the present LC conditions, trastuzumab eluted as a symmetrical peak at 4.6 min and is baseline separated from the BSA peak (Figure 2). MS spectra of the sample show there is minimal mass overlap with the charge envelop for BSA between m/z 1000–2000 and trastuzumab between m/z 2000–3500. This baseline LC separation and minimal mass overlap suggest the matrix interference from BSA in trastuzumab quantification is likely to be minimal and the quantification results are a good measure of the protein quantity.
Serially diluted solutions of trastuzumab in 0.1% formic acid containing 0.1 mg/mL BSA were analyzed under full scan ESI+ data acquisition conditions. Sample at each concentration was injected in triplicate and quantification assessed using bioanalytical criteria for biologics (deviation <25% and S/N at LLOQ = 10). The effects of several processing parameters on quantification outcomes were evaluated. Specifically, quantification was carried out using mass tolerance windows ranging from 0.1 to 2.0 Da, and a wide 1000 Da window. In addition, the effects of summing varying numbers of peaks (1 to 42 peaks or summation of the entire protein envelope) were studied and are described in detail in the section that follows.
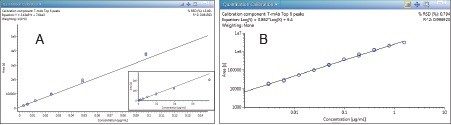
Overall, the best quantification was derived based on a 0.5–1.0 Da mass tolerance window. The 3 ng/mL LLOQ was achieved with a S/N ratio greater than 10. The calibration curve was fitted with linear regression using 1/X2 weighing. The extracted ion chromatograms (XICs) showed a concentration dependent response and the observed linear range was from 3 ng/mL to 780 ng/mL or 2.4 orders of linear dynamic range with R2 = 0.98 (Figure 3A). Table 1 shows the calibration table for the three replicate injections at each concentration. The data shows that linear deviation at the LLOQ using a “sum of multiple peaks based method” – whether it is six peaks, nine peaks, 14 peaks, or 14 charge states, is <25%. When quantification was derived based on a single peak or a wide 1000 Da mass tolerance window, the data showed outliers with deviation >25% at the LLOQ. When outliers were excluded, the same LLOQ of 3 ng/mL was obtained with the dataset meeting the method validation criteria. When the data was plotted in log-log scale, the calibration range extended from 3 ng/mL to 1560 ng/mL or 2.7 orders of magnitude with R2 = 0.997 (Figure 3B).

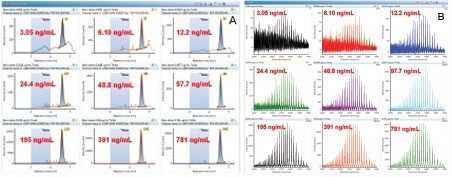
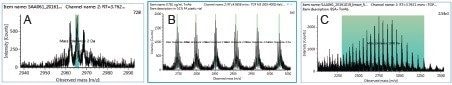
Representative XICs for the m/z 2907 peak (3–780 ng/mL) are shown in Figure 4. Mass spectra for each concentration provided positive confirmation (Figure 4). When multiple peaks are summed for quantification purposes, a total peak area is reported in the results table. Individual chromatograms for each m/z of the multiple peaks are displayed in the chromatogram window allowing the user to inspect and manually adjust the peak integration when desired. Figure 5 is an example for the sum of nine top GF0/GF1 peaks at the LLOQ concentration (3 ng/mL) using a 0.5 Da mass tolerance window. For these nine peaks, the calculated S/N values ranged from 11 at m/z = 2696 to 22 at m/z = 2907, all meeting the LLOQ criteria of S/N >10. At this concentration range, mass shift of the charge states are minimal, multiple peaks used for quantification provided adequate coverage of most abundant charge states at each concentration.
The presence of multiple peaks in the full scan spectrum, for large proteins, shown in Figure 1, raises a general question about how best to perform quantification. A single peak or multiple peaks may be combined to yield a trace that is quantifiable. In the case of multiple peaks, there can be many charge states and/or isoforms. Additionally, the mass spectrum peak of an intact protein is wider than typically seen for small molecules and peptides and leads to another variable that may be optimized (mass tolerance window). For example, Figure 6a shows that for an XIC using a 2 Da mass tolerance window (2965.3 +/- 1 Da), the entire peak (isoform) is integrated. Reducing the mass tolerance window may lead to enhanced selectivity albeit with a reduced peak area. When the mass tolerance window is too narrow, low peak coverage could lead to low sensitivity. Therefore, balancing selectivity and peak coverage maximizes sensitivity and reproducibility (ultimately balancing S and background N to yield the maximum S/N).
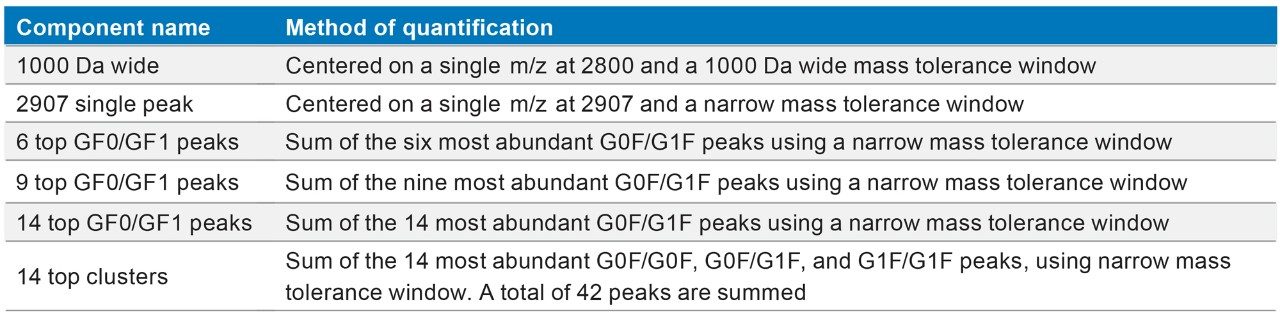
To study the effect of peak selection and summation, and mass tolerance window on quantification, the full scan data was processed using one of seven different ways of peak summation according to Table 2. With the exception of a 1000 Da wide window, for each peak list selection, the mass tolerance window was also varied (0.1 Da, 0.5 Da, 1 Da, 1.5 Da, and 2 Da). Figure 6 shows examples of mass spectra used for, single peak, top six peaks, and 1000 Da wide mass window data processing. The green highlighted regions display the mass tolerance used. Instructions on how to enter the peak list and corresponding m/z, global, and component specific mass tolerance values are described in detail in Appendix I.
Results showed that using an increasing number of peaks or mass tolerance window led to an increase in peak area as one would expect, but the increase did not directly translate to better data quality or an increased sensitivity. Close examination of the calibration table shown in Table 1 indicates that at concentrations >12 ng/mL all methods of quantification using a fixed 0.5 Da mass tolerance window produced acceptable %accuracy. It was near or at the LLOQ that different methods of peak selection had a direct effect on data quality. To quantify this effect, the readily available %RSD reproducibility of peak area from three replicate injections at the LLOQ was used. A higher %RSD indicates poor reproducibility where some data could appear as an outlier in the calibration table. Figure 7 is a summary of %RSD of all peak list selections and mass tolerance windows.
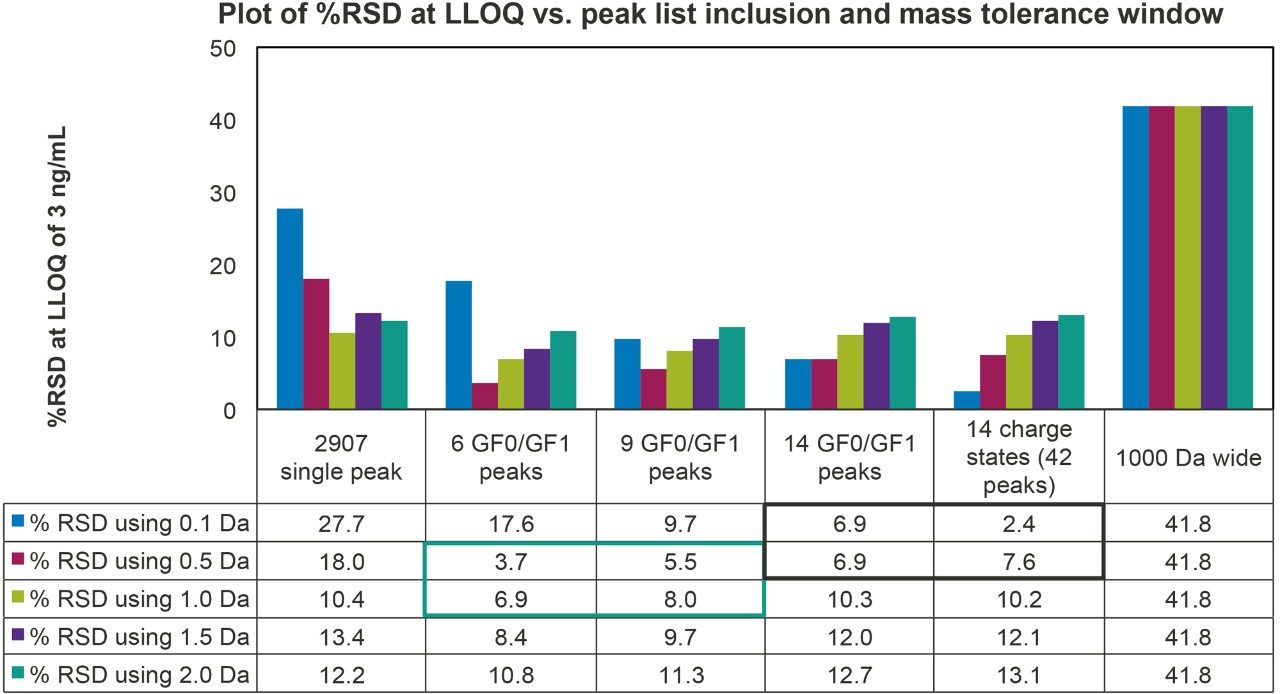
Data showed when the sum of multiple peaks was used as the basis for quantification, in general, increasing the mass tolerance window from 0.5 Da to 2.0 Da led to an increase in %RSD. The best data quality was obtained by summing six or nine top peaks and using a 0.5–1.0 Da mass tolerance window (the green bracket region in Figure 7). There was no outlier and the S/N ratio of all individual traces was >10. Using a narrow 0.1 Da mass tolerance window, the %RSD was generally higher. Although %RSD from the sum of 42 peaks using the 0.1 Da window was low, further inspection of individual XIC traces indicated poor data quality with S/N for many of the traces <3. The use of a 0.1 Da window is thus not recommended. The use of a single m/z peak or a wide 1000 Da mass tolerance window is also not recommended due to poor data quality as reflected by high %RSD. Although other data processing methods result in similar or lower %RSDs, they did not meet the >10 S/N ratio requirement at the LLOQ (the black bracket region in Figure 7). To summarize, the present analysis suggests that quantification based on summation of multiple peaks and using a mass tolerance window between 0.5–1.5 Da produced the best quantification results especially at the LLOQ level.
It should be noted that the present study applies to the specific sample/matrix used and is restricted to the limited number of permeations of peaks and mass tolerance windows investigated. For new samples and new matrix, especially those where potential matrix interferences are expected, users are encouraged to explore additional permeations for obtaining the best quantification outcome. How to set up processing parameters is illustrated in Appendix I.
The complexity of trastuzumab, and biotherapeutic proteins in general, require a multifaceted and flexible approach for their characterization and quantification. Highly sensitive direct quantitation at the intact level was achieved using a Waters Vion IMS QTof and UNIFI. Using trastuzumab, a 150 kDa mAb protein, the LLOQ in the presence of 0.1 mg/mL BSA is 3 ng/mL (or 30 pg load on column) with linear dynamic range of 2.4 orders of magnitude. Thorough examination of the quantification method, using different peak selection and mass tolerance windows, suggests that a “sum of multiple m/z peaks” based approach provides better quantification results at the LLOQ than a single m/z peak or a very wide 1000 Da mass window based approach. Above LLOQ different methods of quantification produced similar results. Overall, the present data suggests that HRMS is capable of providing highly sensitive intact level quantification of biotherapeutic proteins. Data derived from the intact level analysis will complement those from ligand binding assays or LC/MS/MS based surrogate peptide approaches for a complete understanding of the proteins in question.
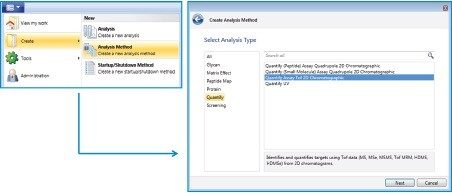
To create a new method, choose the dropdown menu on the top left corner and then choose “Create Analysis method”. A “Create Analysis Method” wizard appears. In the “Select Analysis Type” panel, choose “Quantify” and then “Quantify Assay Tof 2D Chromatographic” (Figure A). After advancing to the next panel, enter a filename and click on finish, the method file will be created.
Once the method is created, select the “Purpose” tab and “Manage Components” section. Enter component name, retention time, and expected m/z as shown in Figure B. Multiple masses can be entered by clicking on “specify mass chromatogram(s)”, as shown in the insert of Figure B. Enter individual masses in the table. After clicking the “OK” button, the entered masses will appear in the component table. The six components used to process the presented data set are displayed.
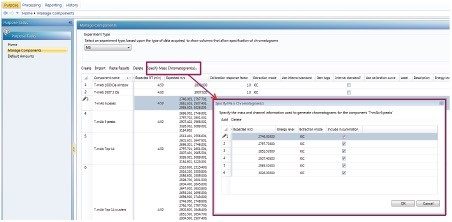
Figure B. Entering single or multiple m/z values on the manage component page. Note, in UNIFI version 1.8.2, if evaluating multiple settings while simultaneously using the same mass, the repeating masses in different components need to be slightly different. In the present case, the third decimal place of the repeating m/z value is varied to ensure they are different and displayed correctly.
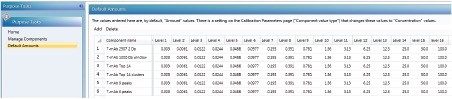
The standard concentrations are entered on the “Default Amounts” tab. Each component has its corresponding concentration, which is named by the header: Level 1...Level 2...This header is called in the sample list.
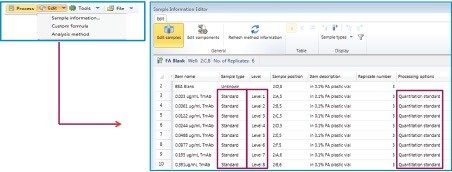
In order to perform calibration of the sample set, make sure “Sample type”, “Level”, and “Processing Options” are filled. After acquisition, the sample list can be modified by choosing “Edit, sample information” at the top right of the screen as shown in Figure D.
The mass tolerance can be entered as a default value which applies for all components in the method, or as a custom value applying to a specific component only. Navigate to the Processing Tab and select Mass Tolerance in the analysis method under Processing Settings. The default mass tolerance value is entered in the method on the “Extraction Settings” panel as shown in Figure E and will apply to all components entered in the “Purpose, Manage Components” section. In this example, 2.0 Da is entered as the default mass tolerance. Component specific tolerances may be entered by clicking “Specify components…” which opens the “Specify components” panel. Highlight the component of interest and click the “add > >” button to move the component to the “Selected components” panel on the right and click “OK”. You may now enter a specific mass tolerance for this component. In this case, a 1000 Da mass tolerance value is entered for component “T-mAb 1000 Da Window” (Figure E). All other components can be edited in this manner, or will continue to use the default value if not changed.
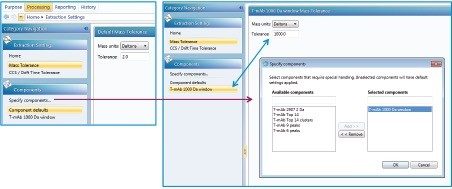
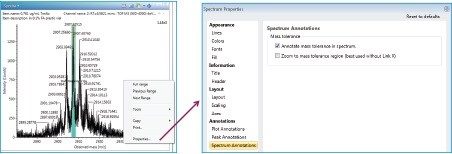
Once processed, the mass tolerance may be highlighted in the spectrum (as a green box). To view the tolerance window, right click in the spectrum screen and choose “Properties...”. The “Spectrum Properties” window will appear. In the “Spectrum Annotations” window check the box “Annotate mass tolerance in spectrum”.
720006222, March 2018