This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates that any appropriate calibrant can be used for determining collision cross section (CCS) measurements on a SYNAPT HDMS System, providing a completely flexible approach for polymer characterization.
The SYNAPT HDMS with travelling wave ion mobility spectrometry (T-Wave IMS) capabilities provides a straightforward approach to CCS calibration and measurement.
Polymeric materials are becoming increasingly sophisticated to meet the demanding requirements of new industrial applications. In addition to a polymer’s bulk physical properties, it has been predicted that 3D shape may have functional importance in the future.1 Therefore, it is reasonable to assume that the ability to measure CCS to confirm theoretical calculations of a polymer’s 3D arrangement will gradually become more important.
Waters SYNAPT HDMS System uses T-Wave IMS to provide a simple, reliable, and rapid approach to measuring CCS values. The mobility cell within the instrument must be calibrated to carry out these measurements. A popular calibrant is polyalanine. The singly protonated ions have CCS values between 89 and 276 Å2, for molecules with between 3 and 19 repeat units. These values have been taken from a widely referenced source, Professor David Clemmer’s Group at Indiana University, USA.2 Accurate CCS results require the analyte to be within the calibration range, and the ions must have the same charge state.
Some polymeric materials are relatively large, and multiple charging is common with electrospray ionization. So, what are the options if your sample doesn’t fit within the polyalanine calibration?

DriftScope Software was created by Waters to allow visualization and interpretation of ion mobility data, and it includes the ability to automatically calculate CCS values. A known substance needs to be infused under the same instrument parameters as the sample. The software requires a “csv” file for the calibrant, containing a list of m/z and respective Å2 values, to be imported into the software to create a calibration file. This means that any calibrant can be chosen assuming it meets a few simple requirements: the CCS values are known, the range of CCS values are appropriate for the analyte, and the charge state of the ions are the same.
The Clemmer Group’s database of CCS values covers a range of peptides, proteins, and oligonucleotides with a variety of charge states. To demonstrate that any appropriate calibrant can be chosen, the following two substances from Clemmer Group’s database where purchased, polyalanine and polylysine. Each sample was run under the same conditions to allow CCS values to be measured for both compounds, using the other as a calibrant. Figure 2 shows the graph of results. The CCS values measured had errors between +3% and -2%, well within the ±5% error generally quoted for these measurements on the SYNAPT.
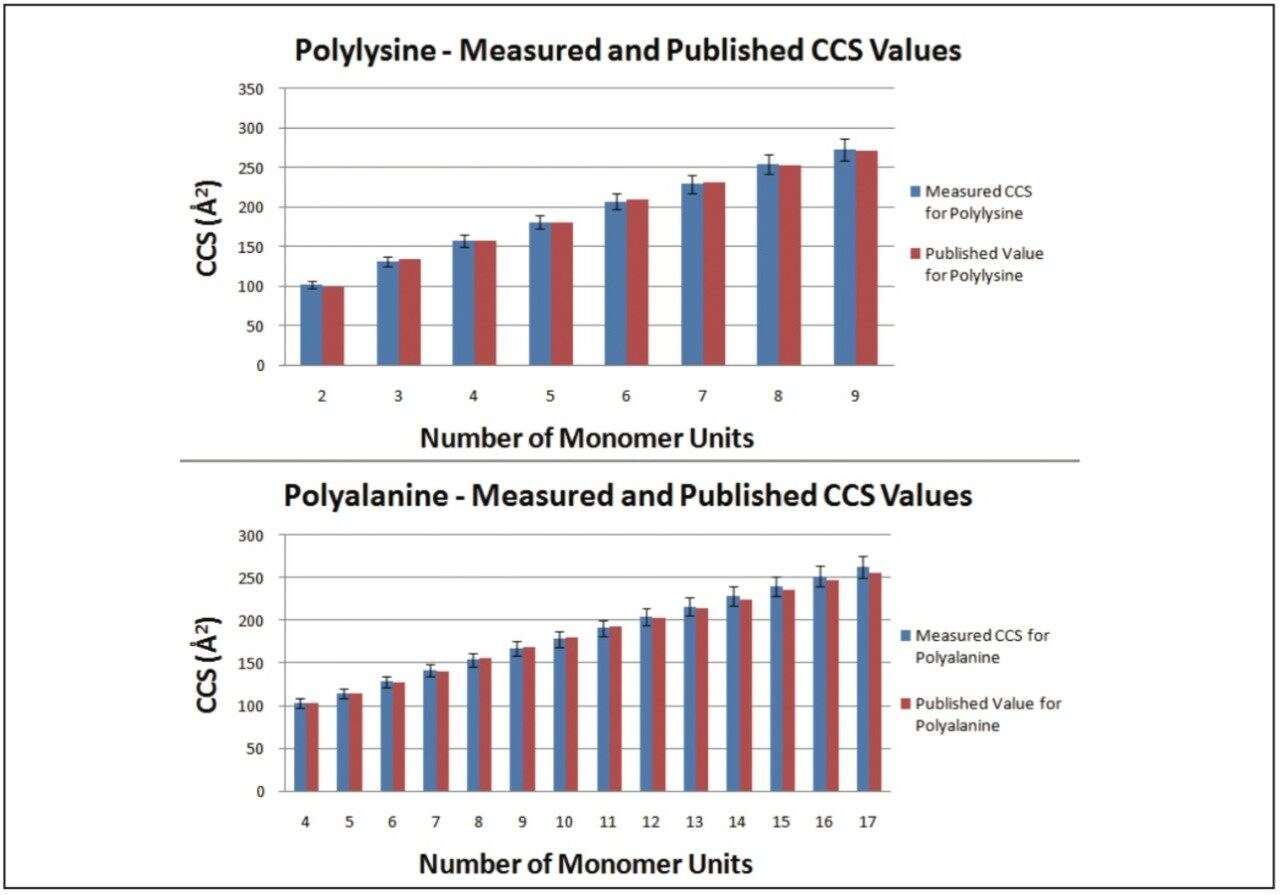
If polyalanine is not an ideal calibrant, due to the range of CCS values, an alternative can be found in published literature. Polymers, such as polyethylene glycol and poly methyl methacrylate, have been characterized in detail.3,4 CCS value for multiply charged polyalanine have also been published.5
This study has demonstrated flexibility for calibrating the SYNAPT HDMS ion mobility cell for CCS measurements. The requirements for a calibrant are limited to the following: the CCS values must be known, the calibration range is appropriate, and the charge state of the calibrant and sample are the same.
It has been predicted that, as polymers become increasingly sophisticated, their 3D arrangement will become important. Theoretical experiments are required to predict the most likely spacial arrangement of these molecules. Currently, ion mobility is the only methodology that can confirm these calculations.
720004699, May 2013