
In this application note, the performance of the new pesticide screening solution was assessed via the analysis an independent proficiency test sample.
The Pesticide Screening Application Solution with UNIFI has been designed to confidently report the presence and absence of pesticide residues at Maximum Residue Limits (MRLs).The solution streamlines data analysis with customizable workflows leading to the rapid analysis of complex matrices using a scientific library functionality with flexible reporting templates.
The Pesticide Screening Application Solution with UNIFI has been designed to confidently report the presence and absence of pesticide residues at Maximum Residue Limits (MRLs). The solution streamlines data analysis with customizable workflows leading to the rapid analysis of complex matrices using a scientific library functionality with flexible reporting templates.
Current trends indicate that in excess of 500 pesticides are routinely used on a global basis. In addition, countries throughout the world have different regulations concerning licensing and MRLs. With increasing global trade, there is a requirement for multi-analyte screening strategies capable of efficiently detecting residue violations in the interest of consumer safety. Recent advances in MS and separations technology have demonstrated a strong potential to improve the analysis of complex samples that often result from the extraction of food samples. For example, these include advances in MS sensitivity and increased peak capacity where UPLC technologies are deployed. This study shows the performance of a new benchtop QTof-based platform incorporating StepWave and QuanTof technologies combined with UNIFI. UNIFI is a revolutionary scientific information system specifically designed for non-targeted accurate mass screening applications that enables scientists to test and report more samples for an increasing number of residues and contaminants. UNIFI captures high resolution, accurate mass LC-MS data, and performs rapid processing for confirmation and quantification of residues in a single software environment.
The benefits of full spectra acquisition with the specificity provided by accurate mass measurement are well understood, and include the ability to perform historical data review as well as non-targeted screening experiments. StepWave technology provides enhanced sensitivity with excellent linear dynamic range, as required for routine quantitative pesticide screening. Such enhancements in Tof hardware technology have improved the precision and accuracy of generated data, facilitating the creation and utilization of more intelligent data processing software packages.
In this application note, the performance of the new pesticide screening solution was assessed via the analysis an independent proficiency test sample. Comparison between median calculated concentrations and Z-Scores reported for the sample were used to determine the performance of the new approach to pesticide screening.
|
System: |
ACQUITY UPLC I-Class |
|
Run time: |
15 min |
|
Column: |
ACQUITY UPLC BEH C18 2.1 x 100 mm, 1.7 μm |
|
Column temp.: |
45 °C |
|
Mobile phase A: |
10 mM ammonium acetate dissolved in water |
|
Mobile phase B: |
10 mM ammonium acetate dissolved in methanol |
|
Flow rate: |
0.45 mL/min |
|
Injection volume: |
5 μL |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
|
|
1 |
Initial |
0.45 |
98 |
2 |
|
2 |
0.25 |
0.45 |
98 |
2 |
|
3 |
12.25 |
0.45 |
1 |
99 |
|
4 |
13 |
0.45 |
1 |
99 |
|
5 |
13.1 |
0.45 |
98 |
2 |
|
6 |
15 |
0.45 |
98 |
2 |
|
MSE conditions |
|
|---|---|
|
Mass Spectrometer: |
Xevo G2-S QTof |
|
Ionization mode: |
ESI+ |
|
Capillary voltage: |
1 kV |
|
Sample cone: |
20 V |
|
Source temp.: |
120 °C |
|
Desolvation temp.: |
550 °C |
|
Desolvation gas: |
1000 L/H |
|
Reference mass: |
Leucine enkephalin [M+H]+ = 556.2766 |
|
Acquisition range: |
m/z 50 to 1200 |
|
Acquisition rate: |
4 spectra/s |
|
Collision energy low energy: |
4 eV |
|
Collision energy ramp high energy: |
10 to 45 eV |
Sample extracts, matrix matched calibrants, and quality control samples generated for an EU-RL proficiency test (FV-13 2011) were generously supplied by WIVISP, Brussels, Belgium. A mixed pesticide standard containing 135 pesticides was prepared at six different concentrations. Bulk blank extract was prepared according to a standard liquid-liquid extraction protocol for mandarin. Following the solvent extraction step, the extract was aliquoted and spiked using 50 μL of the appropriate spiking solution to give concentrations equivalent to 1, 5, 10, 20, 50, and 100 μg kg-1 bracketing the EU “blanket” MRL of 10 μg kg-1 for pesticide residues. The spiked solvent extracts were diluted 10-fold in water, prior to analysis. The resulting final solution concentrations were 0.1, 0.5, 1, 2, 5, and 10 ng mL-1.
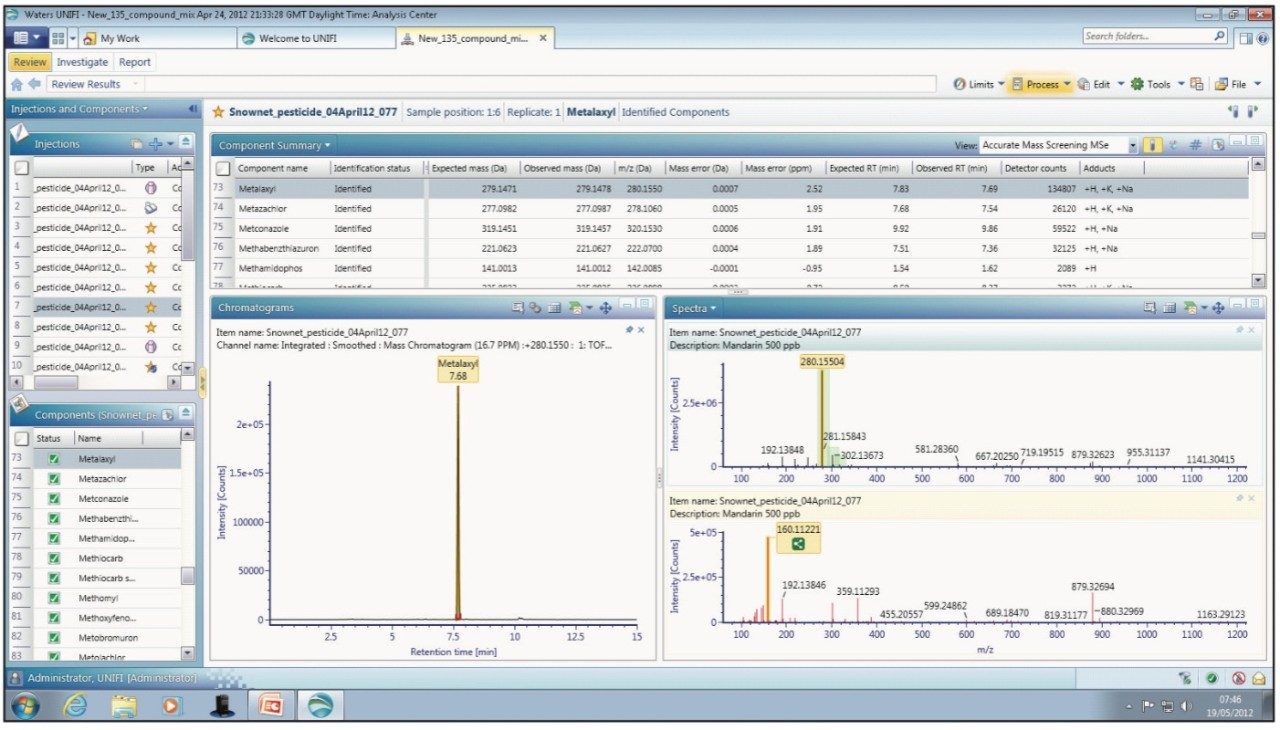
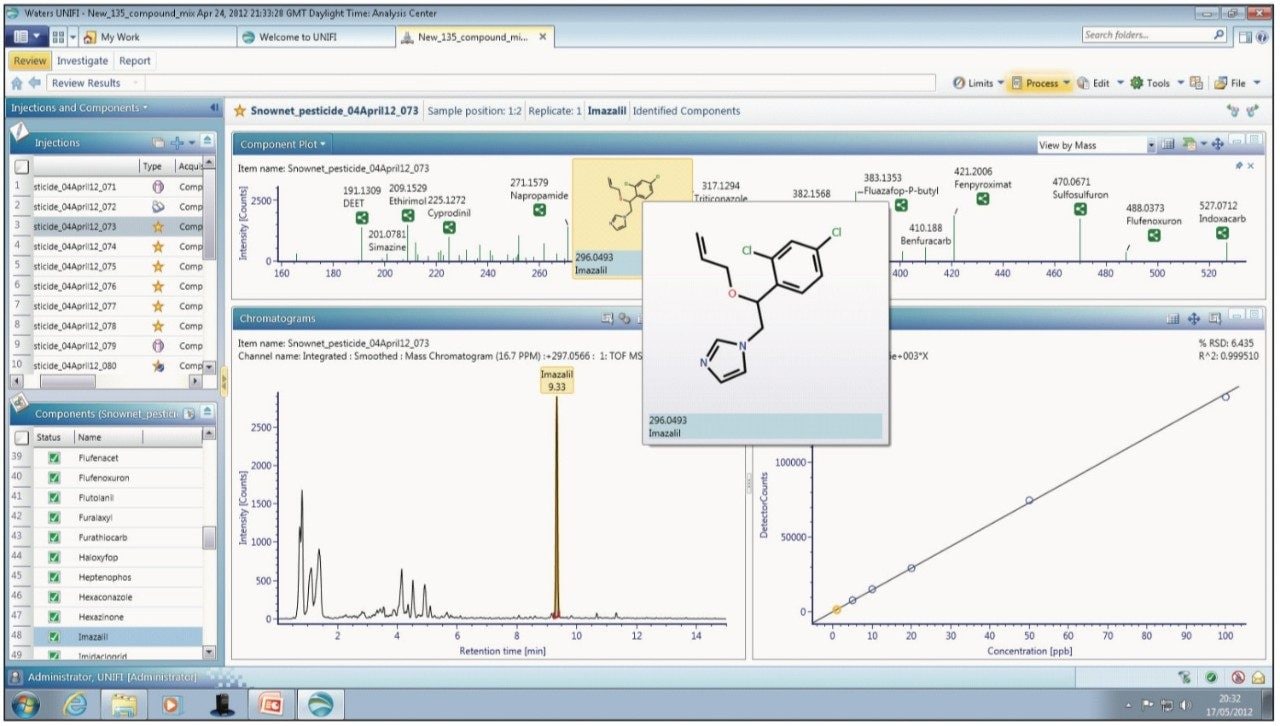
Within UNIFI, pesticides are screened using the scientific library, which allows identification based on the accurate mass of the parent, including common adducts like Na+, K+, and NH4, retention time, the presence of fragment ions, as well as isotopic patterns and intensities. The ability to identify pesticides using multiple compound criteria enables the analyst to be confident in making an identification, while speeding up the review process by allowing user-defined filters and workflow steps to be applied. The use of workflow steps in conjunction with user-defined filters provides an efficient approach to review both qualitative and quantitative analysis in a single software environment. Figure 1 shows an example view that can be configured by the user. In this case, the extracted ion chromatogram of the selected pesticide from the current injection is displayed along with the low energy (precursor) and high energy (fragment) spectra.
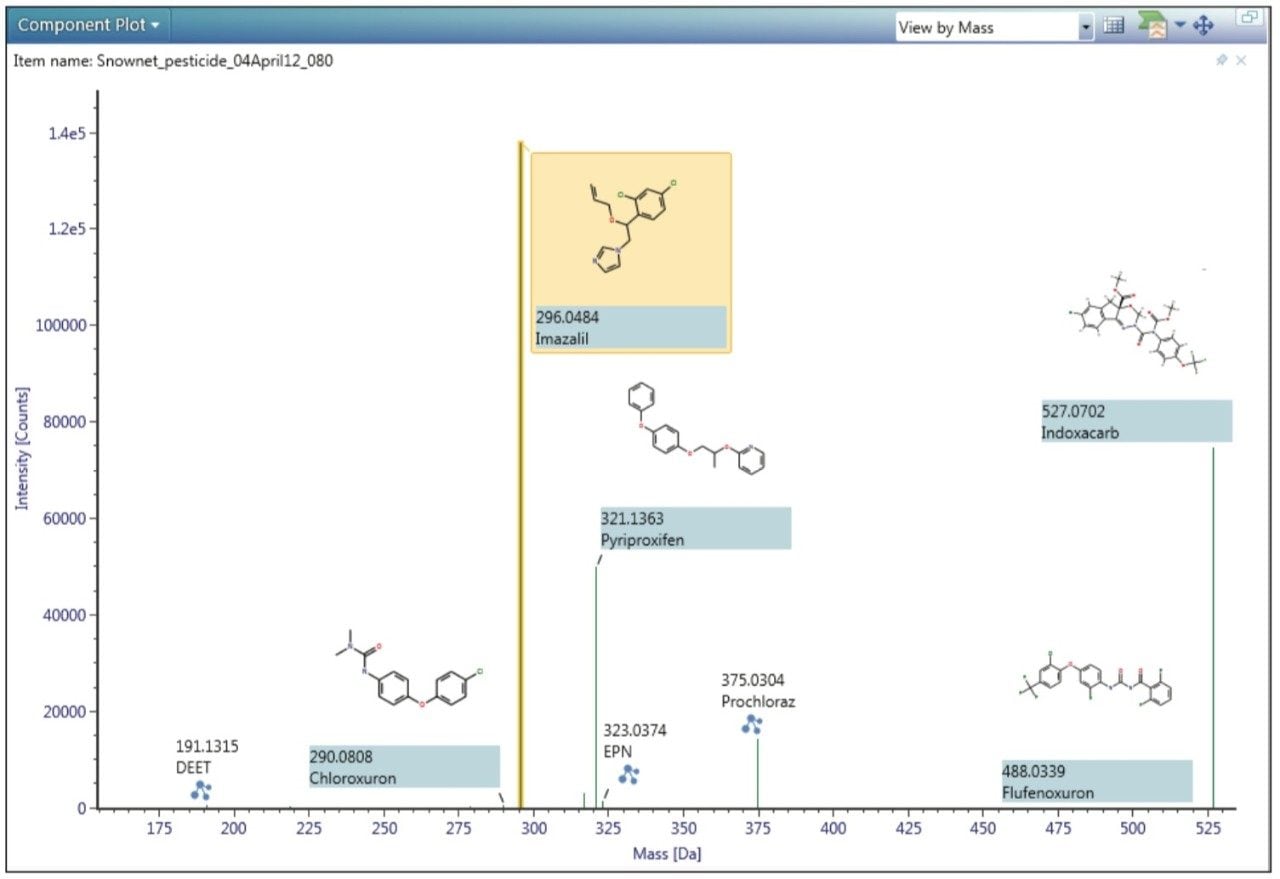
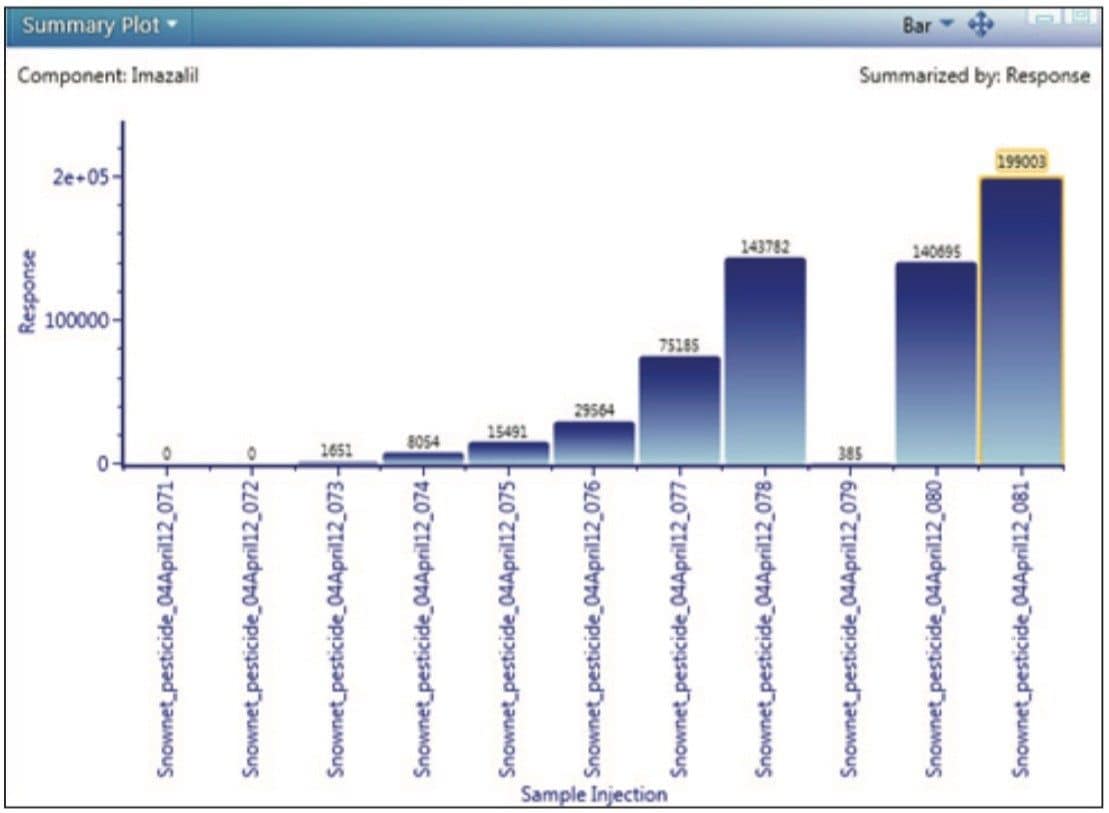
In addition, a summary table, which is also user-configurable, is included to show further information (such as mass error, adducts identified, retention time, and isotope match) to assist with confirmation of the analyte identity. The pesticides identified in a sample can also be quickly viewed in the component plot along with their expected masses and structures, as shown in Figure 2. Another useful feature of UNIFI is the summary plot, which is shown in the example view in Figure 3. This plot allows the user to view the concentration of pesticides across the sample set, enabling rapid detection of any pesticides present in blanks or samples, and determining if the concentration varies between the injections.
The user-defined filters enable selection of matches that meet specific criteria. In this way, a user can choose how to review the data in a step-wise fashion. Those pesticides that are identified with high confidence (as defined by the user) can be reviewed rapidly. If there are matches that fail defined criteria, they can be filtered out to be reviewed with additional information. The criterion that has failed the acceptable limit can be flagged to be easily recognizable. Example criteria that could be flagged include an accurate mass shift or questionable isotope ratios. In addition to screening for the presence or absence of pesticides, quantification is also configurable within the same view. This allows the analyst to filter those pesticides that are above an action concentration (for example, 0.5 MRL), thereby improving laboratory efficiency by ensuring that noncompliant samples are accurately detected and reported.
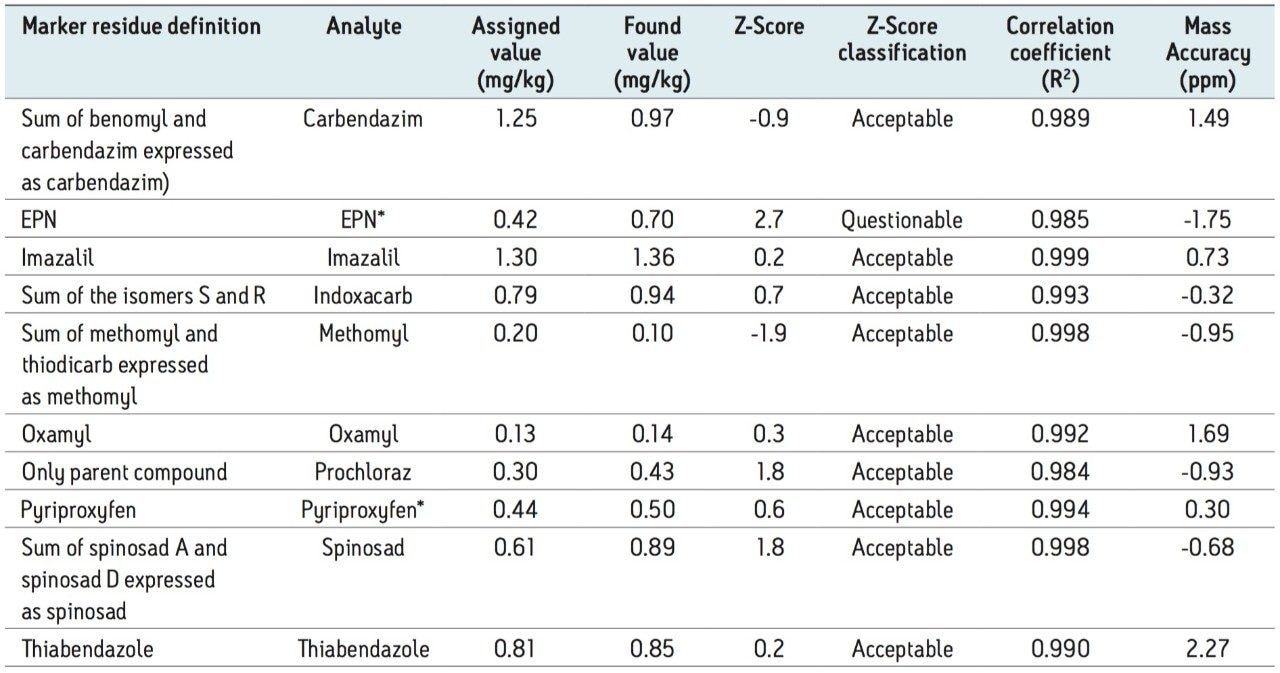
The mandarin extract was screened for the presence of pesticide residues using a target list of LC amenable pesticides. For each of the pesticide residues identified, calculated concentrations were determined using external standard calibration. The results where compared to the published median values obtained from approximately 150 laboratories who participated in the EU-RL proficiency test (FV-13 2011). All of the eight LC amenable analytes present in the sample were correctly identified and quantified using the new pesticide screening solution, as shown in Table 1. In addition, two GC amenable compounds were also detected at the concentrations present in the sample. The Z-Score values for these analytes were also calculated (according to the same formula used by the proficiency test organizers). Nine out of the 10 analytes detected had Z-Score values less than 2.0 and were classified as “Acceptable.” For one of the analytes, EPN, most commonly analyzed using GC, the Z-Score value was 2.7 and classified as “Questionable.” For all 10 analytes detected, the mass accuracy (for both precursor and fragment ions) was less than 3 ppm (exceeding the SANCO 12495 requirement of 5 ppm). The quantitative performance of the instrument, in the presence of a complex matrix, was excellent, as demonstrated by the correlation coefficient values.
Analyte specific information that can be used for confirmation purposes, such as elemental composition and fragmentation pathway, can be routinely generated from the accurate mass spectral data acquired within a single injection. Processing simultaneously acquired accurate mass precursor and fragment ion data through the new scientific information system has enabled the pesticide residues to be confirmed with confidence and very low false positive rates.
The calibration series generated for imazalil is shown in Figure 4. Correlation coefficients of the order of ≥R2=0.99 and residuals of <5% were routinely obtained in matrix extract. The component summary, component details, and component plots are also illustrated, providing easily accessible information to further enhance confidence in the results and accelerate the review process.
Enhancements in both hardware and software technology have contributed to the development of a robust Tof-based solution for pesticide residue screening. These have contributed to the development of the Waters Pesticide Screening Application Solution that allows the routine detection, identification, and quantification of pesticides in complex matrices at MRL concentrations. In this study, residuals of <5% and mass accuracy in the order of 2 ppm were obtained over the dynamic range of the study. A good correlation to EU-RL proficiency test sample (FV-13 2011)was seen for the LC amenable pesticides, as demonstrated by the Z-Score values (≤2). Structural elucidation information for all of the identified pesticides was generated by collecting simultaneous acquisition of accurate mass precursor and accurate mass fragment ions. Processing simultaneously acquired accurate mass precursor and fragment ion data through a new scientific information system enabled pesticide residues to be confirmed with confidence, minimized false positive rates, and accelerated the review process to ultimately increase laboratory throughput.
720004491, November 2012