High Resolution Characterization of Lipid Nanoparticles Using the Xevo™ Charge Detection Mass Spectrometry (CDMS) Instrument - Single Particle Mass Analysis of Intact LNP-mRNA Formulations
Rebecca J. D’Esposito, Ying Qing Yu, Michelle Chen
Waters Corporation, United States
Published on March 17, 2026
Abstract
Lipid nanoparticles (LNPs) underpin the success of mRNA therapeutics, yet their polydispersity, diverse composition, and large size require highly sensitive analytical techniques that can measure high molecular weight to fully characterize them.1 Charge detection mass spectrometry (CDMS) overcomes many of these limitations by directly measuring the charge (z) and mass over charge (m/z) of intact particles in the megadalton range, enabling resolution of discrete particle populations and detailed mass distributions at single particle resolution.2
In this work, the Xevo CDMS Instrument is used to analyze intact LNP mRNA vaccine formulations, uncovering multimodal population profiles and mass variability not observable with ensemble based techniques. These findings highlight the potential of Xevo CDMS Instrument as a powerful approach for evaluating LNP critical quality attributes (CQAs) throughout development and manufacturing.
Benefits
- CDMS measures every particle directly, delivering true single particle masses without deconvolution or charge state assumptions and accurately capturing particle‑to‑particle variability.
- CDMS delivers unmatched clarity, exposing subpopulations that ensemble methods obscure and revealing the full landscape of LNP heterogeneity and loading differences.
- CDMS delivers actionable insight from early formulation through process development and quality comparisons, enabled by high resolution mass readouts and rapid data acquisition.
Introduction
LNPs support the successful delivery of mRNA therapeutics by protecting nucleic acid payloads, improving cellular entry, and enabling controlled pharmacokinetic profiles.1 These particles consist of ionizable and helper lipids, cholesterol, and PEG lipids (Figure 1). The interplay of these components results in a highly diverse set of particle structures with a broad range of sizes and payloads.
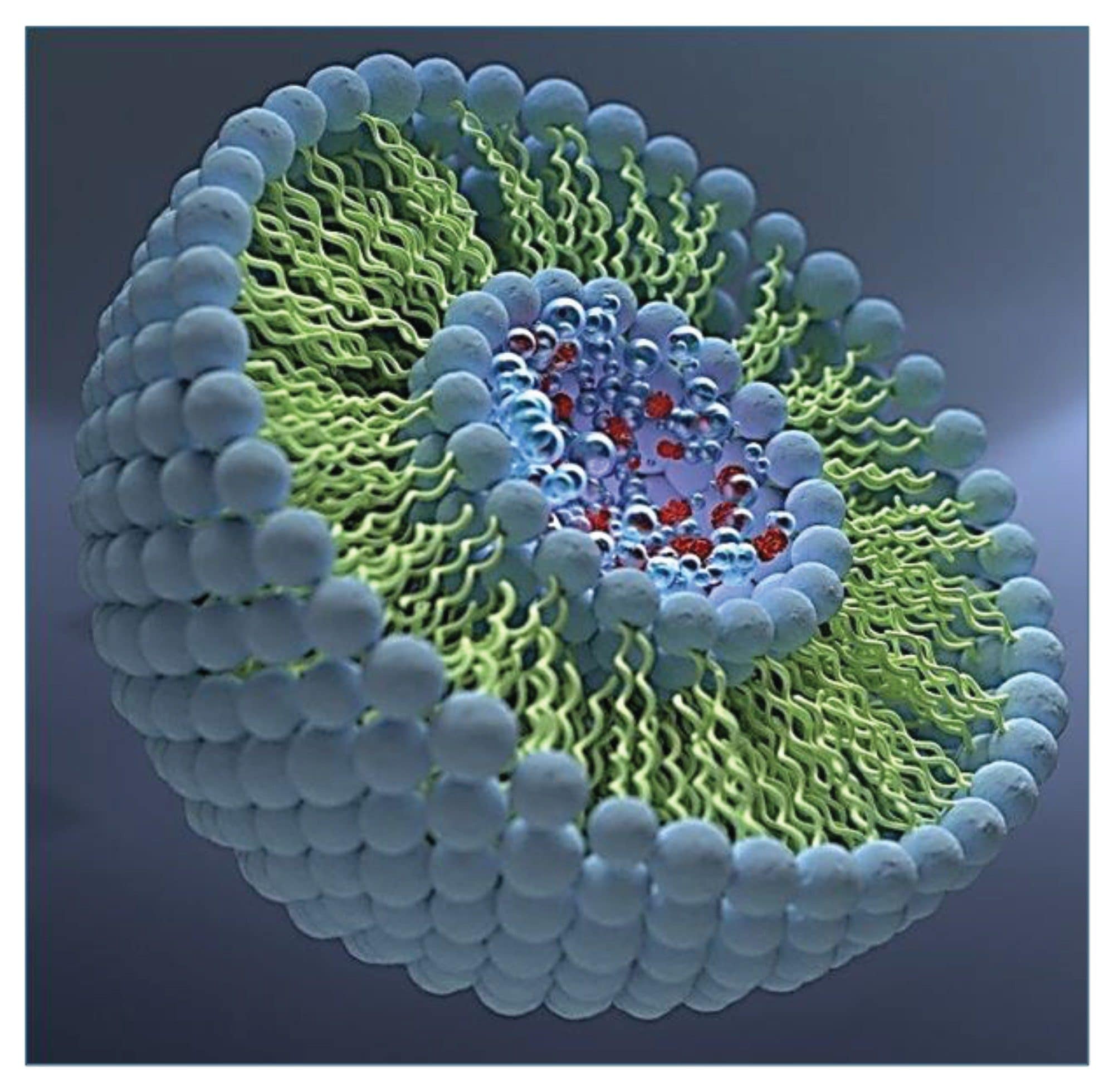
Accurate characterization of this complexity is essential for understanding LNP formulation behavior and ensuring product consistency.1 A broad range of analytical techniques is used to interrogate LNPs, each contributing complementary information. These techniques include light scattering approaches (DLS, SEC‑MALS, FFF‑MALS), microscopy methods such as cryo‑EM and AFM, capillary electrophoresis, analytical ultracentrifugation, and other orthogonal tools.1,5 While each technique provides valuable insight, they also carry inherent limitations: many rely on ensemble‑averaged measurements, require model‑dependent assumptions, or probe only a single component of the particle (lipids, RNA, or morphology).1,5 Critically, none of these conventional methods measure the intact mass of individual particles or resolve particle‑to‑particle heterogeneity across the full distribution. A recent study from the Mitchell Group, which utilized multiple ensemble methods, further underscores that even well‑established LNP systems exhibit substantial nano‑ to mesoscale heterogeneity that remains obscured when relying solely on ensemble techniques.5 These limitations highlight the need for truly particle‑resolved analytics, and CDMS meets this need by directly revealing the multimodal and compositional landscape hidden within LNP formulations.
CDMS enables direct, simultaneous measurement of the m/z and charge of each ion,2 allowing accurate mass determination of intact LNPs on a particle‑by‑particle basis. This single‑particle capability provides analytical resolution that cannot be achieved with traditional ensemble solution‑phase techniques.2 In a recent study, Miller et al. used a research‑grade CDMS platform to analyze thousands of empty and mRNA‑loaded LNPs, demonstrating strong agreement between CDMS‑derived particle sizes and cryo‑TEM measurements.4 These findings confirm that LNPs remain structurally stable under CDMS vacuum conditions and highlight the technique’s suitability for intact‑particle characterization.4 Together, these results establish CDMS as a rapid, label‑free analytical approach for assessing LNP mass, size distribution, and structural stability, providing a powerful complement to existing biophysical methods.4
Here, the Xevo CDMS Instrument is applied to characterize LNP mass distributions and population heterogeneity. The results illustrate how single particle mass analysis can enhance understanding of formulation quality.
Experimental
COMIRNATY® (Pfizer-BioNTech) COVID-19 vaccine (2025-2026 formula) was stored and handled following the manufacturers’ guidelines over the duration of the experiments. The samples were buffer exchanged into 20 mM ammonium acetate solution (pH 7.4) using Slide-A-Lyzer™ MINI Dialysis Devices (Thermo Fisher Scientific®) for 15 minutes at 4 °C before CDMS analysis. Ions were generated in positive ion mode using nano-electrospray ionization (nESI) and mass analysis was performed using the ELIT-based Xevo CDMS Instrument (Figure 2). All data were acquired within the waters_connect™ Informatics Platform (version 4.2.0) using the CDMS Toolkit application and processed within the same application.
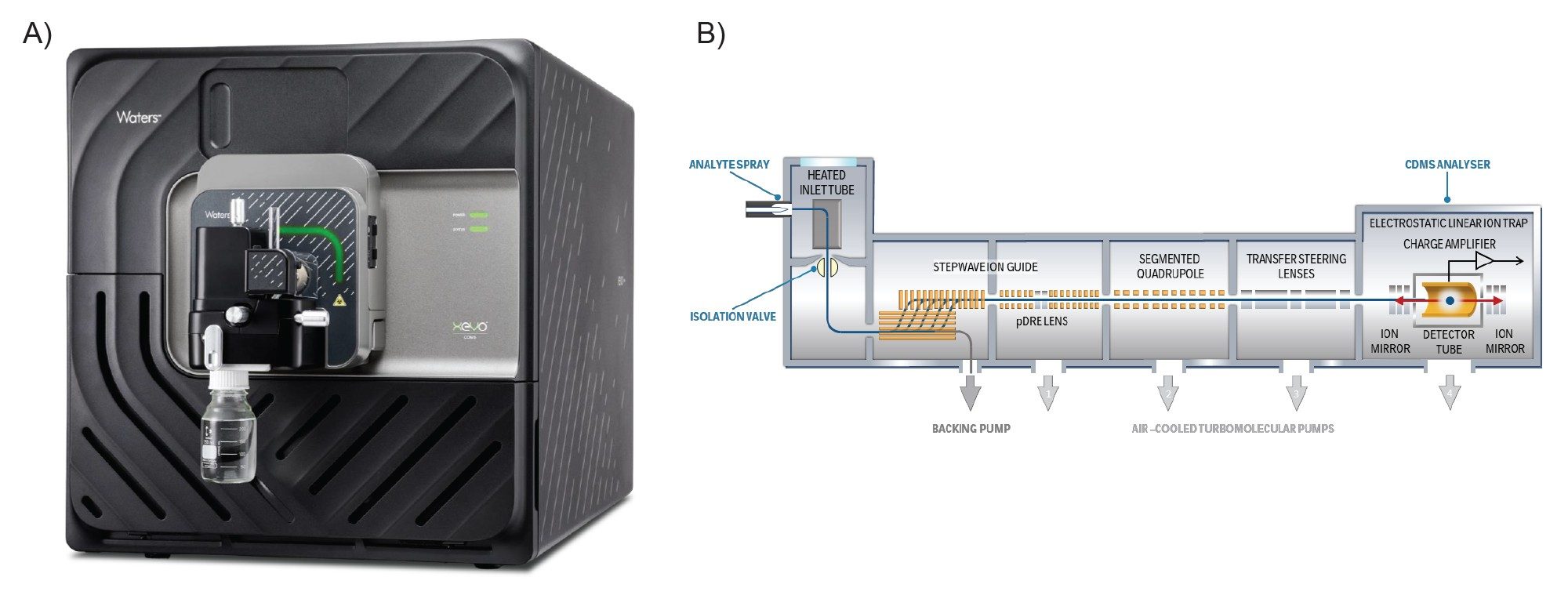
MS Conditions
|
MS system: |
Xevo CDMS Instrument |
|
Acquisition mode: |
Full Scan |
|
Ionization mode: |
Positive |
|
Capillary voltage: |
1.4–1.8 kV |
|
Cone voltage: |
40 |
|
Trapping duration: |
100 ms |
|
Acquisition duration: |
60 minutes |
|
MS software: |
waters_connect Software ver. 4.2.0 |
Results and Discussion
Single Particle Mass Analysis of LNPs by CDMS
CDMS measures both m/z and charge for individual ions, enabling direct calculation of intact particle mass across the megadalton regime. As a result, CDMS reports true single particle mass distributions, rather than ensemble averages, and can resolve overlapping sub‑populations that confound traditional methods when heterogeneity is high. Practically, CDMS enables (i) mass histograms that reveal modal structure and polydispersity, (ii) population level summaries such as mass mean/median and distribution width, and (iii) comparisons across lots or process conditions without deconvolution, even when charge states overlap. These capabilities have been demonstrated broadly for heterogeneous, high‑mass bioparticles and nanoparticles, extending well into the MDa to GDa range.
For LNPs, CDMS is informative because particle mass reflects the combined contributions of lipid composition, nucleic acid loading, and associated solvent. The technique therefore provides a direct measurement of encapsulation heterogeneity and can identify minor heavy or light sub‑fractions that may be obscured in ensemble analyses. These capabilities make CDMS a valuable tool for distinguishing structural and compositional differences that influence overall LNP quality.
Case Study: COMIRNATY (Pfizer BioNTech) LNP Formulation
To illustrate how CDMS informs on marketed LNPs, COMIRNATY (Pfizer-BioNTech COVID-19 mRNA vaccine) as a reference system was analyzed. COMIRNATY’s LNPs comprise an ionizable lipid (ALC‑0315), helper lipid, cholesterol, and a PEG‑lipid, and they are known to present structural and compositional heterogeneity at the particle level. Prior studies using microscopy and DLS have reported soft, compliant particles, with granular internal cores and variable lamellar features, consistent with a distribution of particle architectures rather than a single, uniform structure.
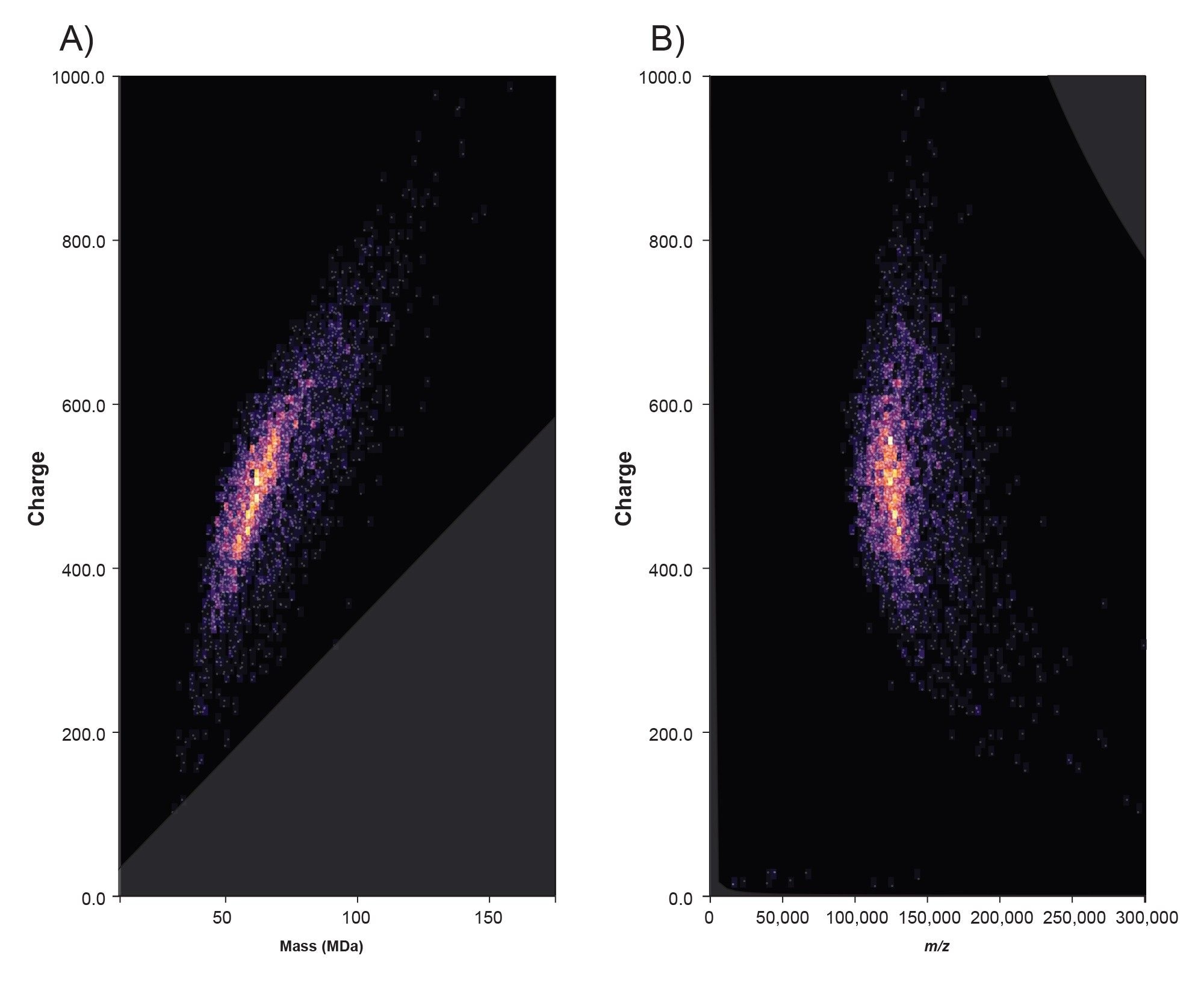
CDMS analysis of COMIRNATY produced a comprehensive single‑particle view of the formulation. The 2D plot of mass versus charge (Figure 3A) shows well‑defined ion populations spanning a continuous charge range, indicating stable ion detection across the formulation’s heterogeneity. The complementary charge versus m/z plot (Figure 3B) separates ions that share similar m/z but differ in charge, which helps disentangle overlapping features that typically obscure ensemble spectra. Taken together with the univariate histograms (Figures 4 A, B, C), these data reveal a peak‑structured mass distribution, indicating several coexisting LNP sub‑populations with different loading and/or packing. The widespread in the CDMS m/z and z profiles for COMIRNATY LNPs reflects real-world heterogeneity in LNP size, mRNA payload (0–100+ copies per particle), and charge states from polydisperse manufacturing. Importantly, the full distribution, including low‑abundance shoulders, is retained without deconvolution, supporting unbiased comparisons across lots or processing conditions.
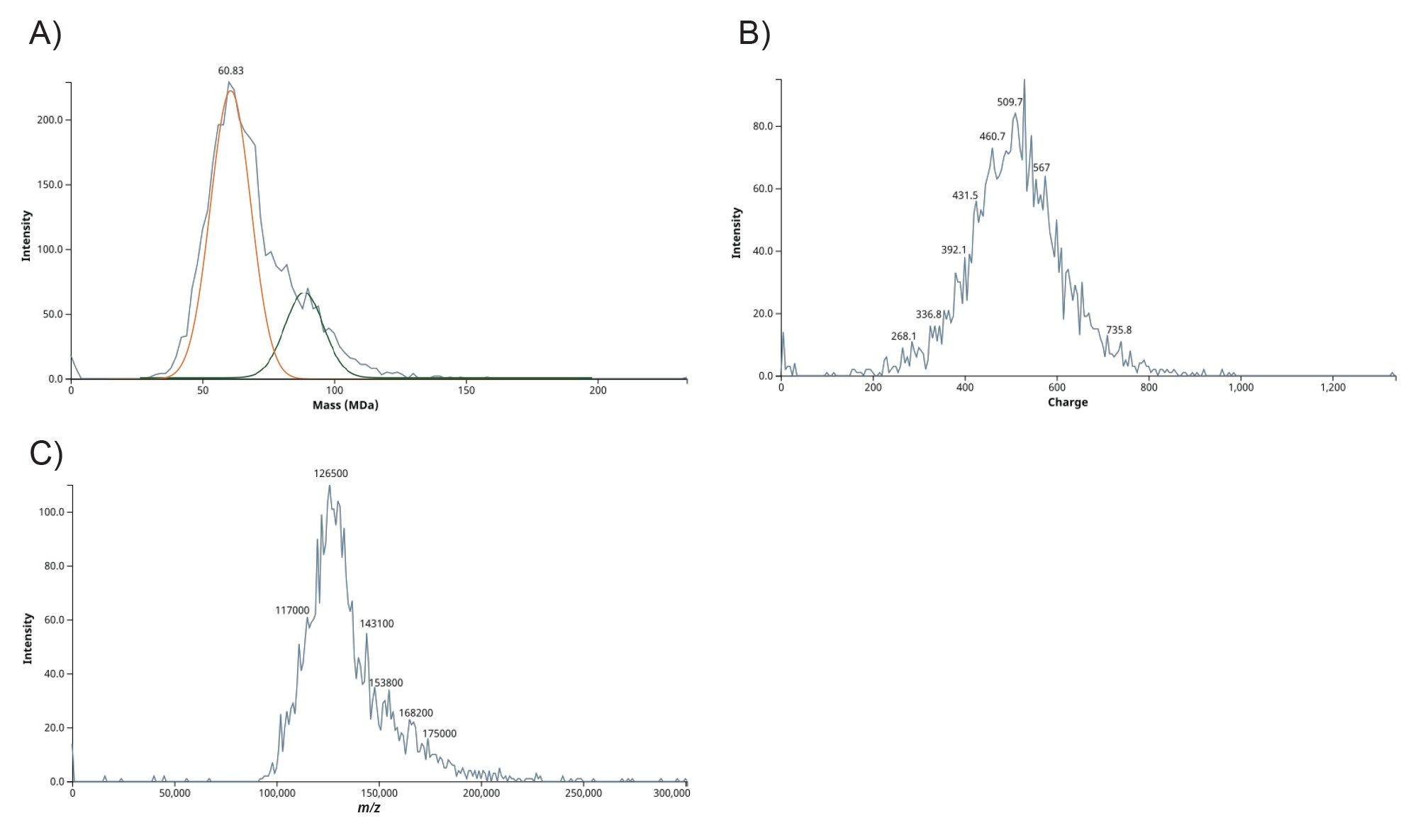
Conclusion
This work presents the first LNP–mRNA data generated on a commercial Xevo CDMS Platform, demonstrating its ability to directly measure single‑particle masses and resolve the intrinsic heterogeneity of LNP formulations. By reporting intact particle mass without deconvolution or charge‑state assumptions, CDMS distinguishes coexisting subpopulations and preserves low‑abundance features that are routinely obscured in ensemble‑based measurements. Applied to COMIRNATY, CDMS produces well‑resolved, peak‑structured mass profiles that reflect meaningful differences in particle loading and composition, enabling deeper insight into formulation behavior, process development, and lot‑to‑lot comparability. Ultimately, Xevo CDMS Instrument provides the high resolution insight necessary to streamline development and strengthen LNP program outcomes.
References
- Hu, C., Bai, Y., Liu, J., Wang, Y., He, Q., Zhang, X., … Liang, Z. (2024). Research progress on the quality control of mRNA vaccines. Expert Review of Vaccines, 23(1), 570–583. https://doi.org/10.1080/14760584.2024.2354251.
- Jarrold MF. Applications of charge detection mass spectrometry in Molecular Biology and Biotechnology. Chemical Reviews. 2021 Oct 12;122(8):7415–41. doi:10.1021/acs.chemrev.1c00377.
- Johann C., Lipid Nanoparticle and Liposome Characterization with FFF-MALS-DLS, WP2608, Wyatt Technology, https://wyattfiles.s3-us-west-2.amazonaws.com/literature/white-papers/WP2608-Lipid-Nanoparticle-and-Liposome-Characterization-with-FFF-MALS-DLS.pdf.
- Miller ZM, Narsineni L, Li YX, Gardner MR, Torpey JW, Williams ER. Single particle charge detection mass spectrometry enables molecular characterization of lipid nanoparticles and mRNA packaging. J Control Release. 2025 Aug 10;384:113856. doi: 10.1016/j.jconrel.2025.113856. Epub 2025 May 18. PMID: 40393530; PMCID: PMC12807447.
- Guimaraes PPG, Zhang R, Spektor R, Tan M, Chung A, Billingsley MM, El-Mayta R, Riley RS, Wang L, Wilson JM, Mitchell MJ. Ionizable lipid nanoparticles encapsulating barcoded mRNA for accelerated in vivo delivery screening. J Control Release. 2019 Dec 28;316:404-417. doi: 10.1016/j.jconrel.2019.10.028. Epub 2019 Oct 31. PMID: 31678653; PMCID: PMC7032071.
Featured Products
720009270, March 2026