For research use only. Not for use in diagnostic procedures.
This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates the utility of QuanRecovery Vials with MaxPeak High Performance Surfaces (HPS) in proteomics applications, and to investigate possible upcoming improvements compared to existing glass vial technology.
Hydrophobic peptide recovery – comparison of glass vials, polypropylene vials, and MaxPeak High Performance Surfaces.
In proteomics experiments, where sample concentration amounts may be limited but consist of hundreds to thousands of tryptically digested proteins, nanoscale chromatography coupled to high resolution mass spectrometry is usually employed to enable the highest sensitivity analysis technique. Separation of peptide species in a reversed-phase liquid chromatography experiment may involve a linear gradient of up to 120 minutes in cycle time duration, and a typical analyte set could consist of several tens of samples. With samples usually analyzed in triplicate to assess reproducibility of results, samples may therefore be present in the autosampler for several days or even weeks. These samples are typically prepared in total recovery glass vials (p/n: 186005669CV) and, while degradation effects may be reduced for samples of great complexity, there is always a possibility of this occurring and maybe affecting the accuracy of the peptide quantification results for samples injected towards the end of an analysis. In this note, we describe the use of Waters polypropylene QuanRecovery Vials with MaxPeak High Performance Surfaces (p/n: 186009186, 186009242, and 176004434) and investigate possible benefits for proteomics analyses.
50 fmol/µL samples of a four-protein tryptic digest (MassPREP Digestion Standard Mix 1 [p/n: 186002865]), were prepared and transferred to LCMS Certified Total Recovery Glass Vials (p/n: 186005669CV) and QuanRecovery Vials (p/n: 186009186). The proteins within Mix 1 are enolase, ADH, glycogen phosphorylase B, and BSA; and preparation and transfer of samples was performed immediately (<1 min) prior to first injection from that vial.
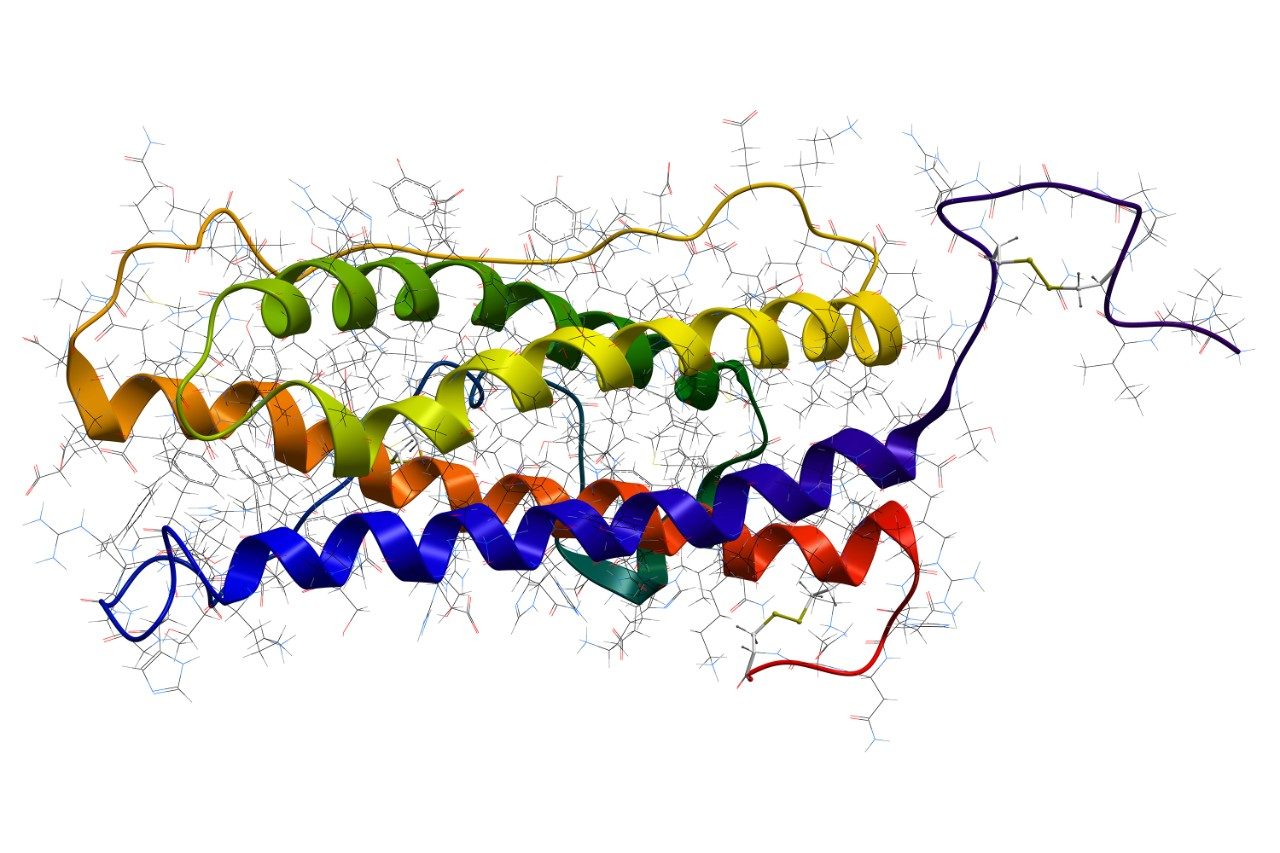
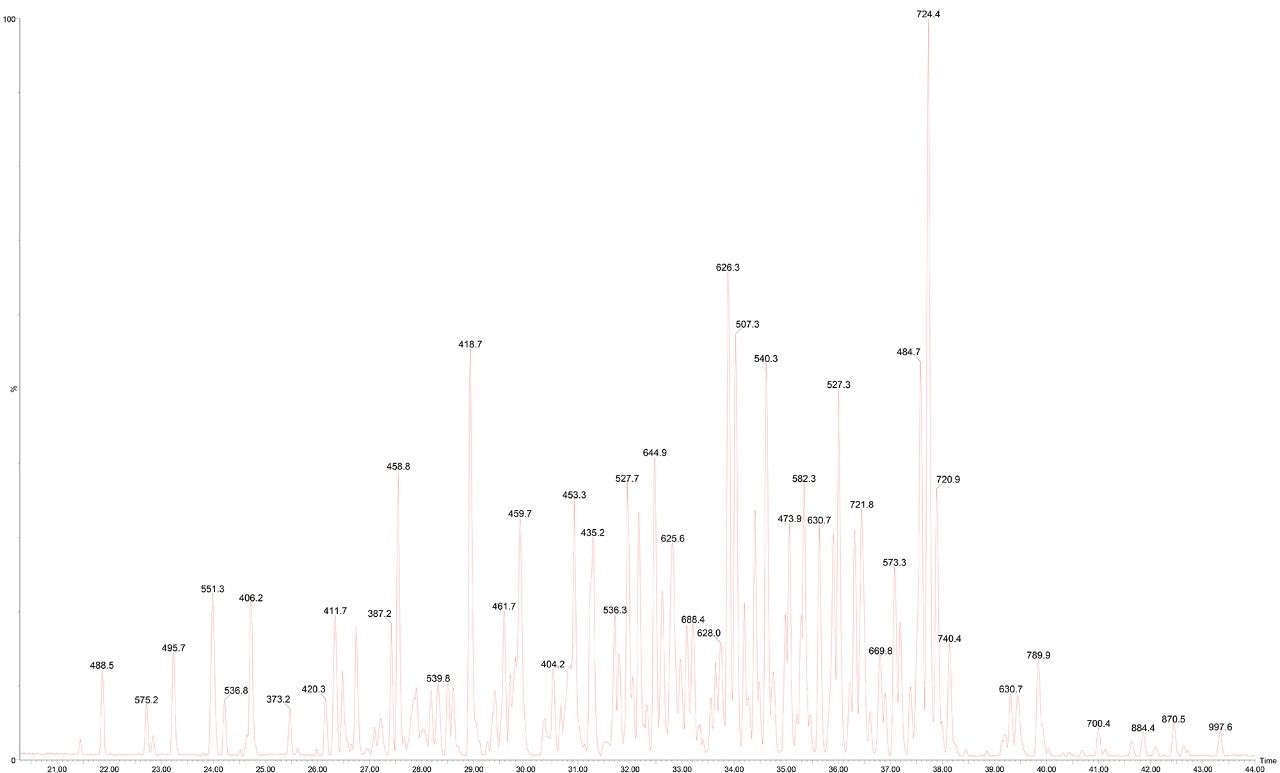
An ACQUITY UPLC M-Class System was configured in trapping mode, using a nanoEase M/Z Symmetry C18, 180 µm x 20 mm Trap Column (p/n: 186008821) and a nanoEase M/Z HSS T3, 1.8 µm, 75 x 250 mm Column (p/n: 186008818). The analytical separation was a reversed-phase gradient of 30 minutes, changing the acetonitrile (+0.1% formic acid) composition from 5 to 40% over 30 minutes at a flow rate of 300 nL/min. A wash at 85% organic and column re-equilibration extended the run time to 75 minutes. Column eluent was connected to a Universal Sprayer (p/n: 205000320) fitted with a PicoTip emitter (p/n: 186003916) which was attached to a NanoLockSpray Mass Ionization Source of a SYNAPT G2-Si Mass Spectrometer operating in the MSE acquisition mode. A typical chromatogram from an injection of 0.5 µL of the sample is shown in Figure 1. Samples were then injected from each vial (polypropylene QuanRecovery or glass Total Recovery) alternatively. A total of nine injections of each, or approximately 22.5 hours.
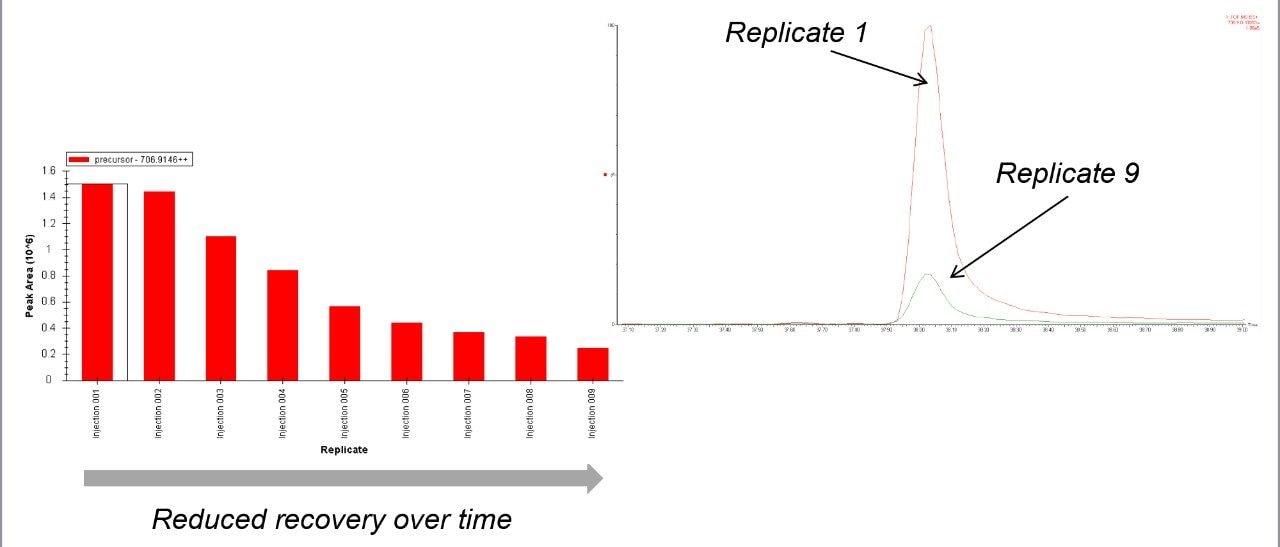
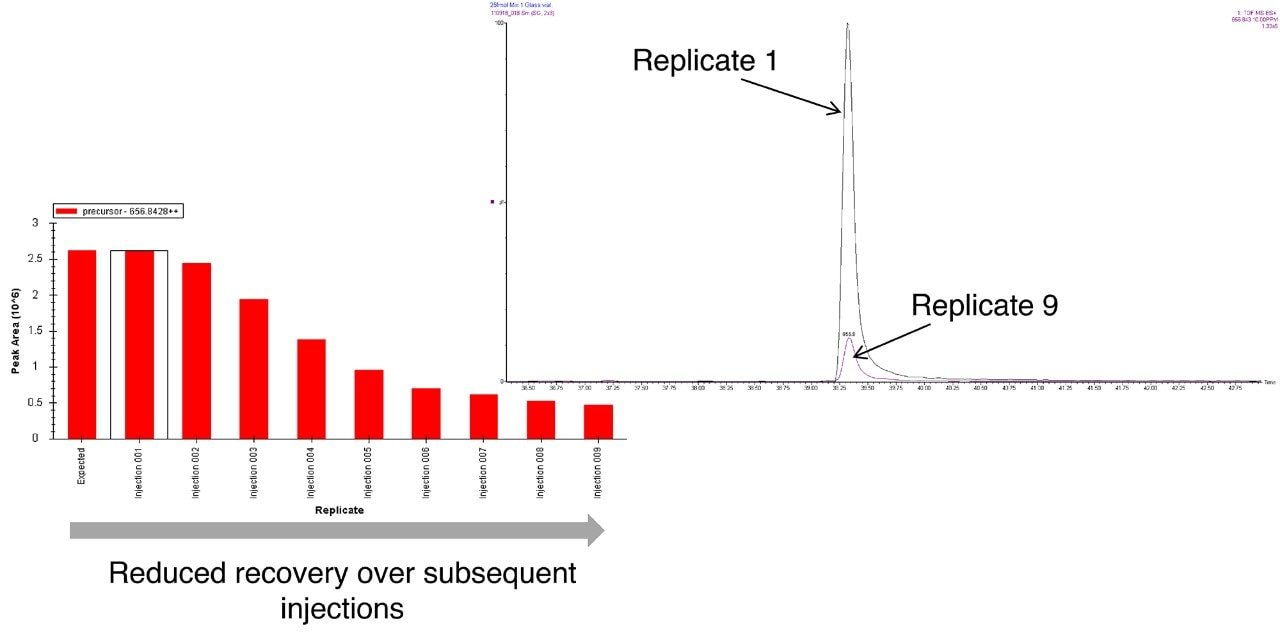
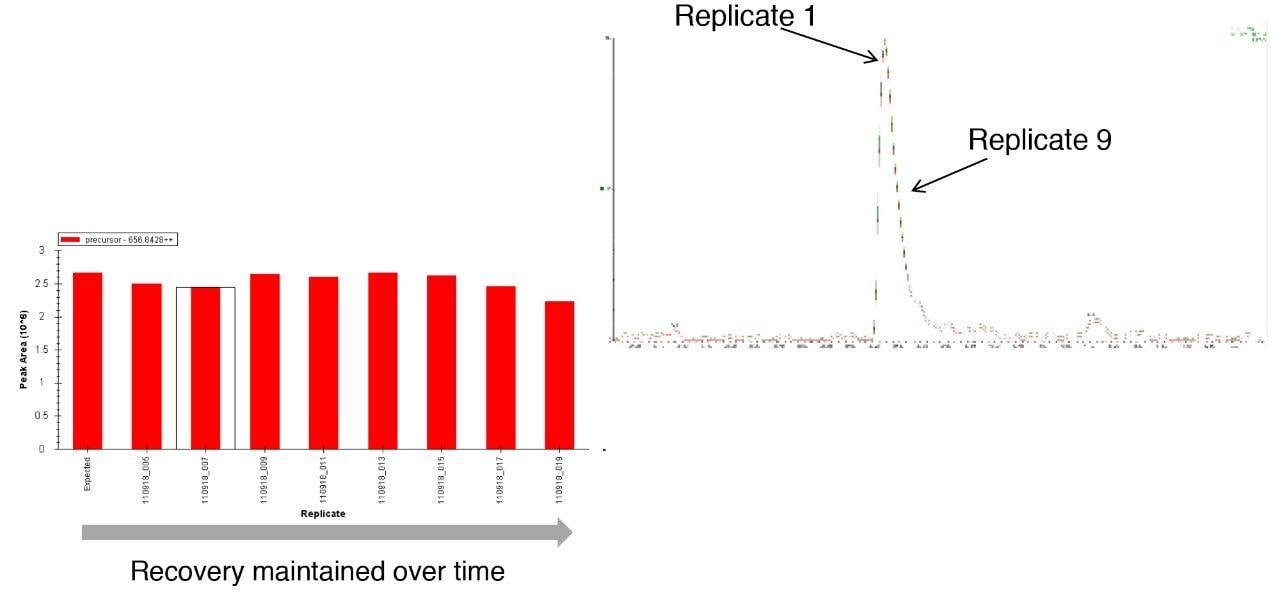
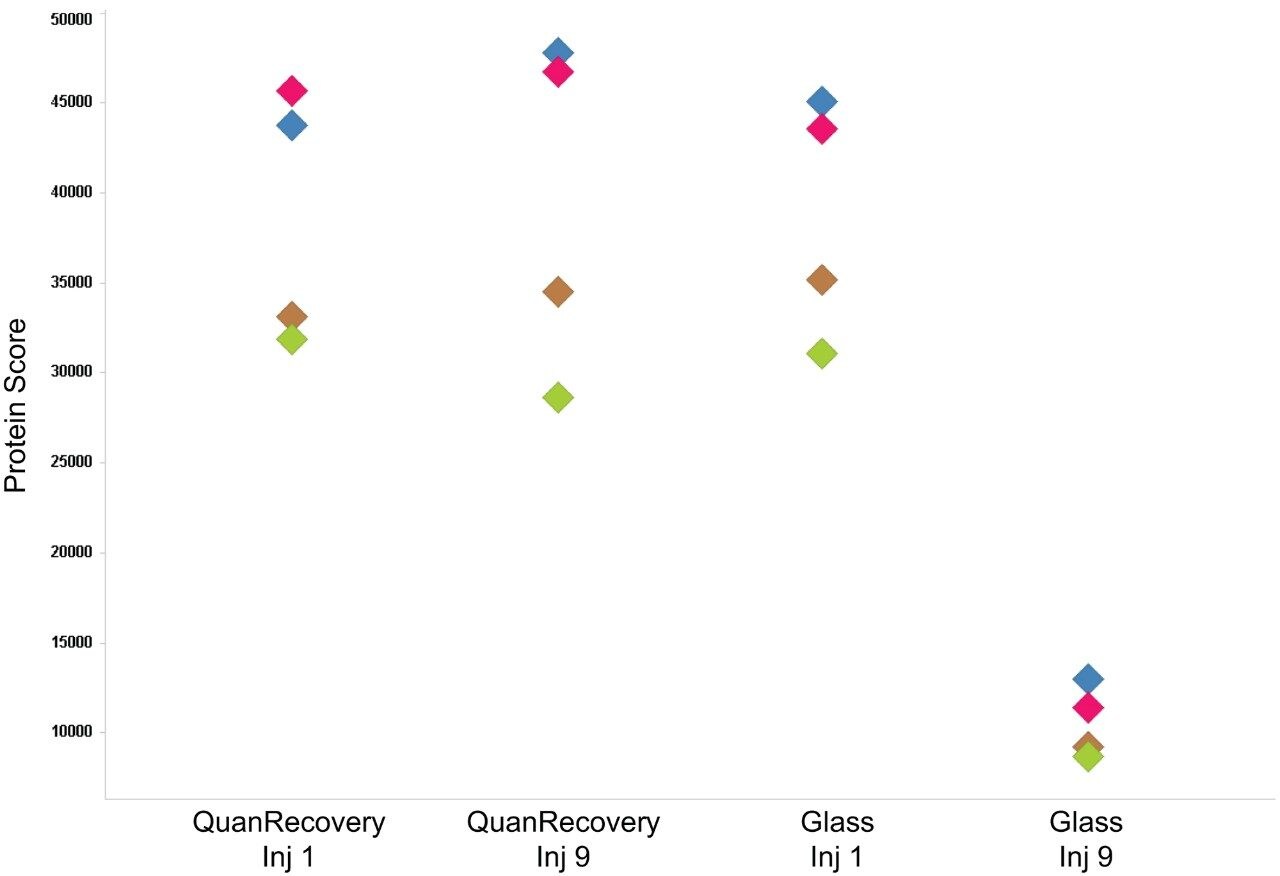
To access individual peptide peak area responses over the experimental time, data were processed using Skyline (University of Washington). For this protein mixture, several peptide peak areas were shown to drop for the sample in the glass vials when compared with the sample from the QuanRecovery Vials (Figure 2 [A, B, C, and D]). This reduction in peak areas tends to be more pronounced for hydrophobic peptides. Another important aspect is the effect on the overall protein identification, and this is shown in Figure 3 where the data has been processed using Waters ProteinLynx Global Server. While the individual protein scores are maintained when injections are from the QuanRecovery Vials, there is a clear reduction by the ninth injection from the glass vials.
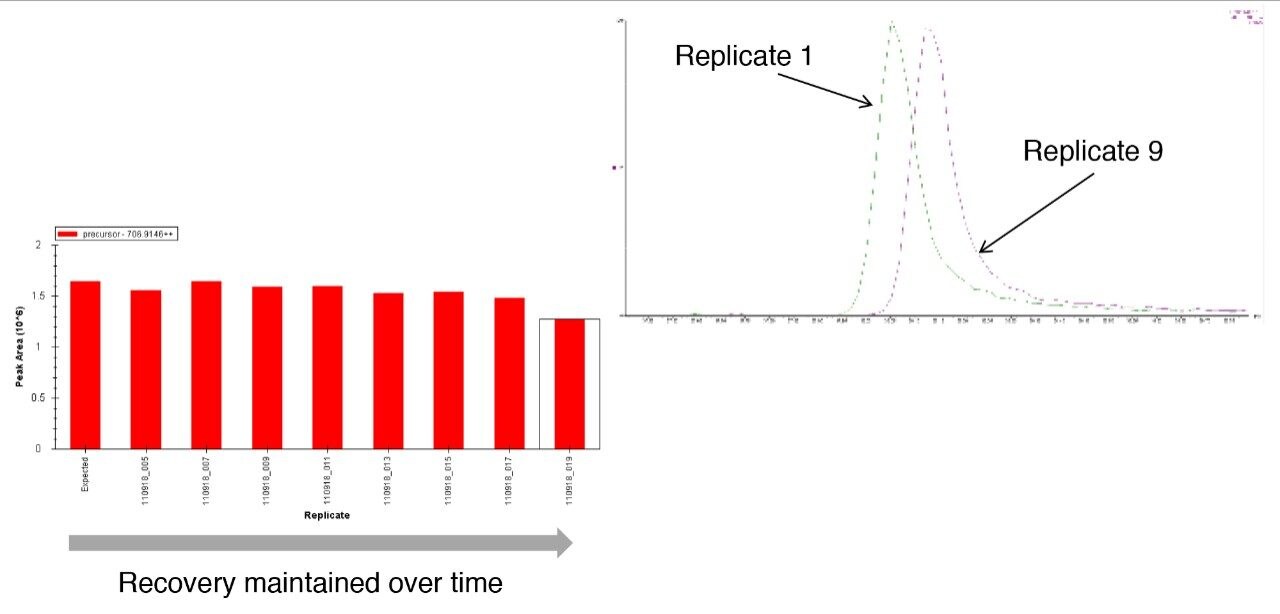
This technical brief has demonstrated the advantages of using QuanRecovery Vials with MaxPeak HPS in a relatively simple proteomics experiment. Reduction of sample degradation, ascertained by observing peak areas of individual peptides over a 22-hour period, were observed particularly for more hydrophobic peptides. This leads to overall protein identification scores being maintained over this period, which could be important for experiments where quantitation results are desired.
720006619, July 2019