We present a fully integrated HILIC UPLC FLR detection workflow that is carried out within a comprehensive compliant-ready platform that integrates informatics and instrument control.
This platform enables laboratories to perform routine biotherapeutic glycan analysis with greater speed, accuracy, and consistency than by using disparate laboratory processes.
One of the challenges of managing the routine use of analytics in biopharmaceutical laboratories is the number of approaches used to characterize a biomolecule – and the complexity of controlling instruments and processing data collected from different structural levels.
The Waters Biopharmaceutical System Solution with UNIFI integrates high-resolution biotherapeutic analyses with bioinformatics that are designed to support routine workflows used in the development process. The system enables researchers to acquire, process, report, and share mass spectrometry and chromatography data including intact mass analysis of proteins, peptide mapping, and glycan profiling – with a single software platform that can also manage multiple systems in a networked workgroup.
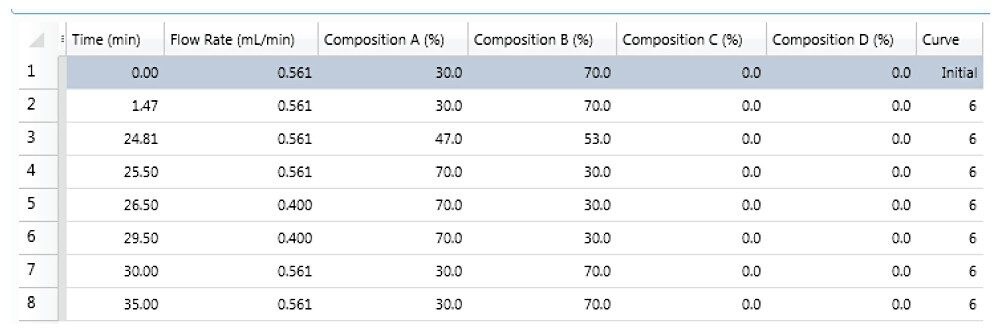
In this application note, we illustrate a dedicated workflow for the acquisition, processing, and automated reporting of data from fluorescent-labeled (such as 2-aminobenzamide, or 2-AB) released glycan samples. Sample data was acquired using a HILIC UPLC separation and fluorescent (FLR) detection methodology. Normalized retention time for the various glycans was achieved using a calibrated RapiFluor-MS Dextran Calibration Ladder (p/n 186007982).
Data acquisition, processing, reporting and management were achieved under control of the UNIFI Scientific Information System. The HILIC UPLC FLR System used for glycan data acquisition was part of a compliant-ready UNIFI workgroup, consisting of two UPLC-QTof MS systems and multiple UPLC optical detection (FLR and UV) systems. This workgroup configuration enabled centralized instrument control, processing, and data integration on a common server for the methods, data, and reports acquired from the networked systems. We demonstrate that such an integrated laboratory and data workflow provides exceptional efficiencies for routine released glycan profiling of biotherapeutics.
|
System: |
ACQUITY UPLC H-Class System |
|
Column: |
ACQUITY UPLC Glycan BEH Amide Column, 130Å, 1.7 μm, 2.1 mm x 150 mm (p/n 186004742) |
|
Column temp.: |
40 °C |
|
Mobile phase A: |
50 mM ammonium formate (pH 4.4) |
|
Mobile phase B: |
Acetonitrile |
UNIFI Scientific Information System
The 2-AB Dextran Calibration Ladder (p/n 186006841) and the GlycoWorks Reductive Amination Single Use Sample Preparation Kit (p/n 176003119) are glycan standards available from the Waters Corporation. The dextran ladder is used to calibrate and normalize labeled glycan retention times for exceptional day-to-day, system-to-system, and lab-lab reproducibility. The retention time for polyglucose 4–12 peaks were used to produce a fifth order polynomial calibration curve (Glucose Unit or GU vs. Retention Time). All analyte peaks are reported and searched using this calibrated GU value.
The majority of the therapeutic proteins are glycosylated, and the attached glycans have significant impact on the efficacy and safety of the biotherapeutic. The International Conference on Harmonization Guideline Q6B requires the analysis of carbohydrate content, structural profiles, and characterization of the glycosylation site(s) within the polypeptide chain(s).
The most widely adopted analytical workflow for routine N-linked glycan characterization involves labeling the enzymatically released glycans with a fluorescent tag (typically 2-aminobenzamide, or 2-AB), resolving the labeled glycans by hydrophilic interaction liquid chromatography (HILIC UPLC), and detecting the labeled glycan peaks with a fluorescence detector. The assignment of glycan peaks during routine analysis is fundamentally based on matching their retention time to established values. For non-routine analysis, glycosidase arrays or MS analysis are employed to give tentative assignments or resolve ambiguous peak assignments.
In order to best control method variation (between runs, days, instruments, scientists, and labs) glycan profiles from the HILIC separation are always calibrated and normalized against a 2-AB Dextran Calibration Ladder (glucose homopolymer). Glycan peaks in an unknown sample can be assigned a Glucose Unit (GU) value from the GU vs. Retention Time calibration curve using the dextran ladder, which is typically fitted with a fifth-order polynomial or cubic spline calibration line.1
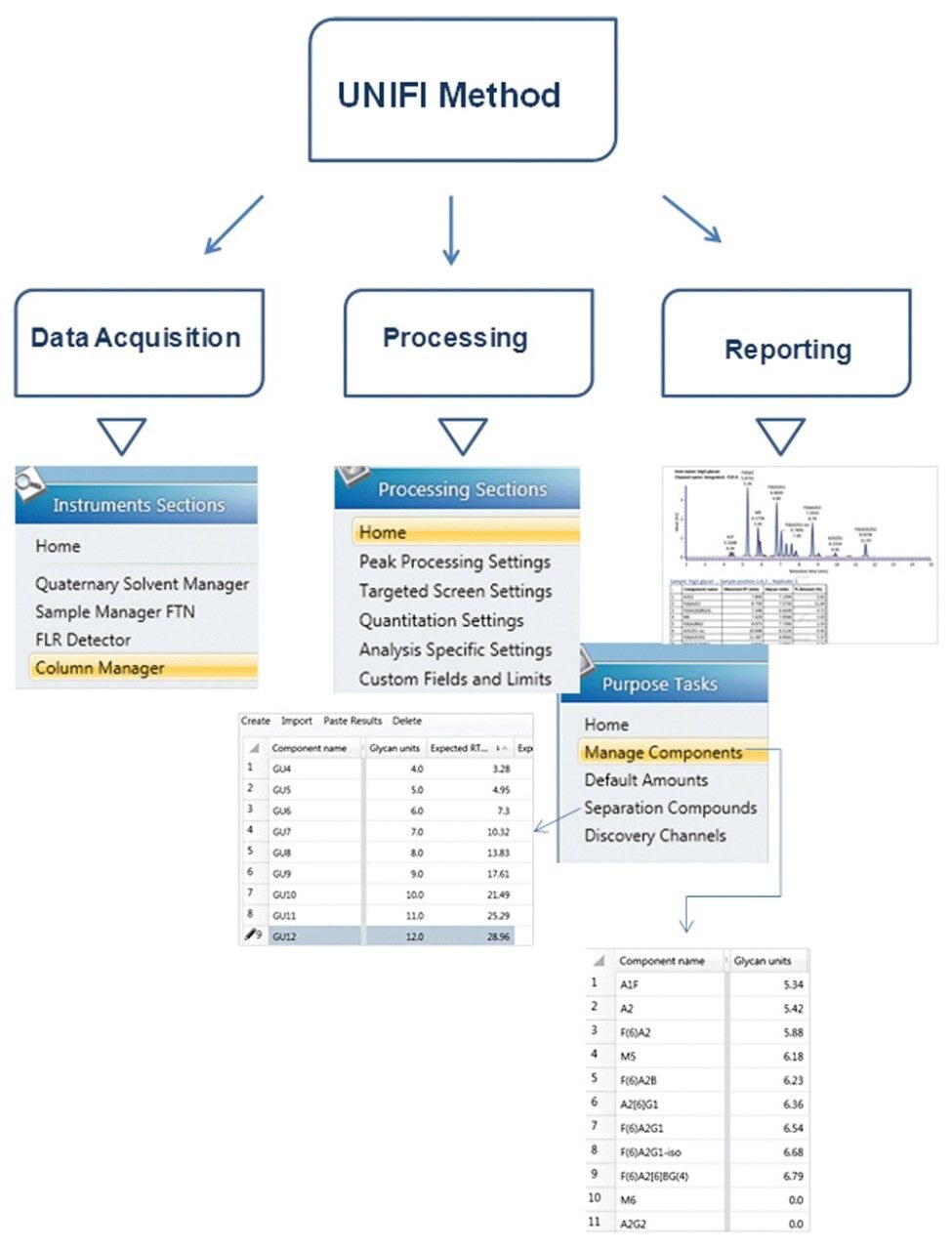
The new streamlined workflow in UNIFI Software, v. 1.6, (Figure 1) enables users to automatically collect one or more dextran ladder standard data sets, process the chromatograms, generate dextran calibration curve, and apply curves directly to unknown samples. Assigned GU values for peaks in an unknown are searched through Glycobase 3.1+, an integrated UPLC GU released glycan database, developed by Waters in collaboration with the National Institute for Bioprocessing Research and Technology (NIBRT) (https://glycobase.nibrt.ie/glycobase/browse_glycans.action).
Targeted component search is enabled within a more information-rich integration of Glycobase in the next version of UNIFI Software, v. 1.6.1. The integrated Glycobase database will reference hundreds of glycan GU entries along with supporting structural, mass, and exoglycosidase digestion pathway information.
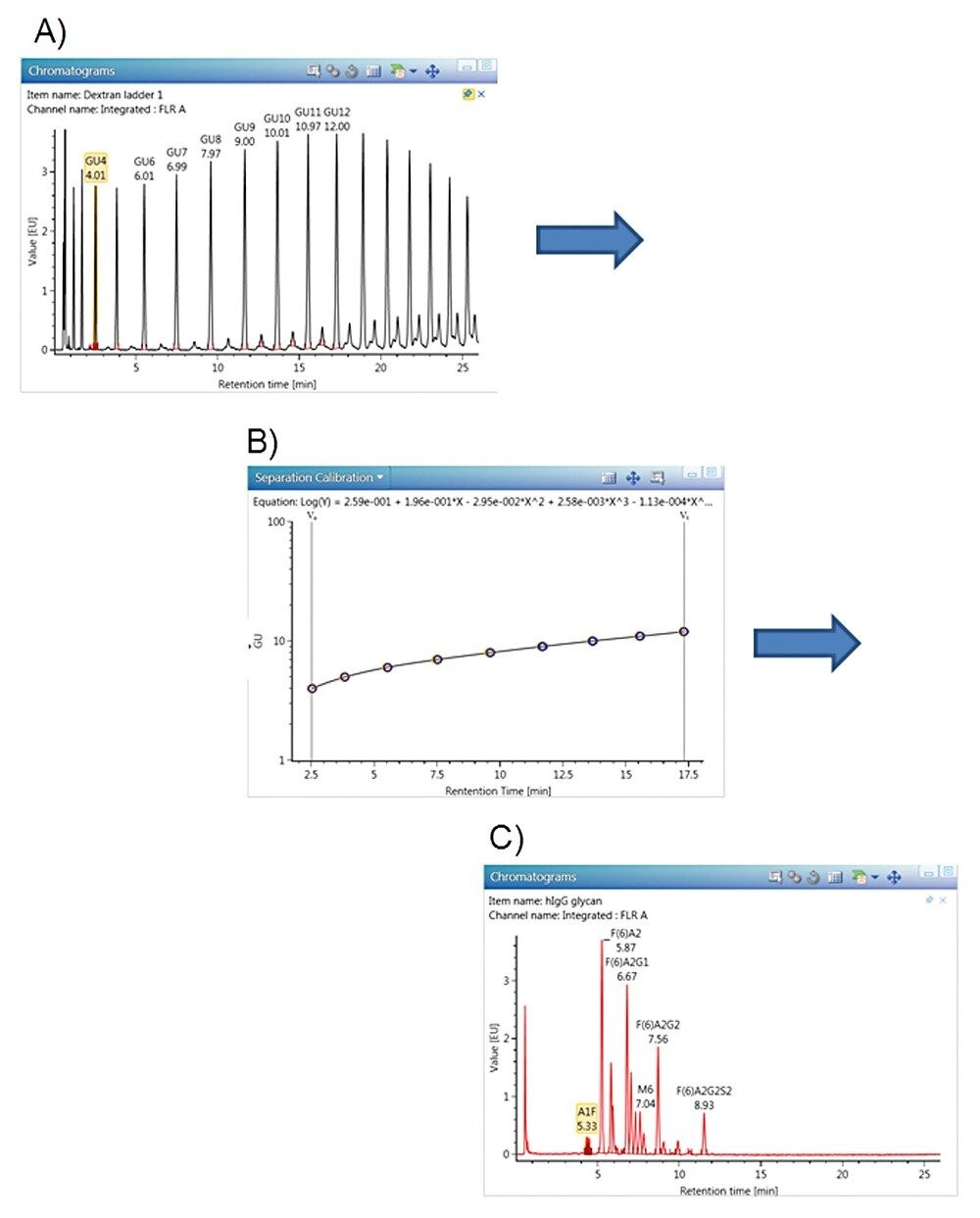
Figure 2 depicts the key elements of producing an automated and holistic glycan acquisition, processing, and reporting method within UNIFI. The instrument settings define the instrument modules required for the analysis, and define method parameters for HILIC UPLC separation and fluorescence detection. The processing setting defines peak integration and dextran ladder calibration settings. A comprehensive report can be generated automatically following acquisition and processing using one or more report templates that can be readily modified. Modifications to the processing and reporting settings within the method are possible post-acquisition, and are audit trailed for simplified documentation and compliance purposes.

Figures 3 through 5 detail how a scientist can use UNIFI Software to navigate through processed and raw chromatographic results. A typical dextran calibration curve shown in Figure 3 is applied to the GlycoWorks Reductive Amination Single Use Sample Preparation Kit (p/n 176003119), a mixture of N-glycans from mAb (Figure 4). Reproducibility of the results is depicted in Figure 5. The ability of UNIFI to automatically summarize data from multiple injections of standards and unknown samples is highlighted in these displays.
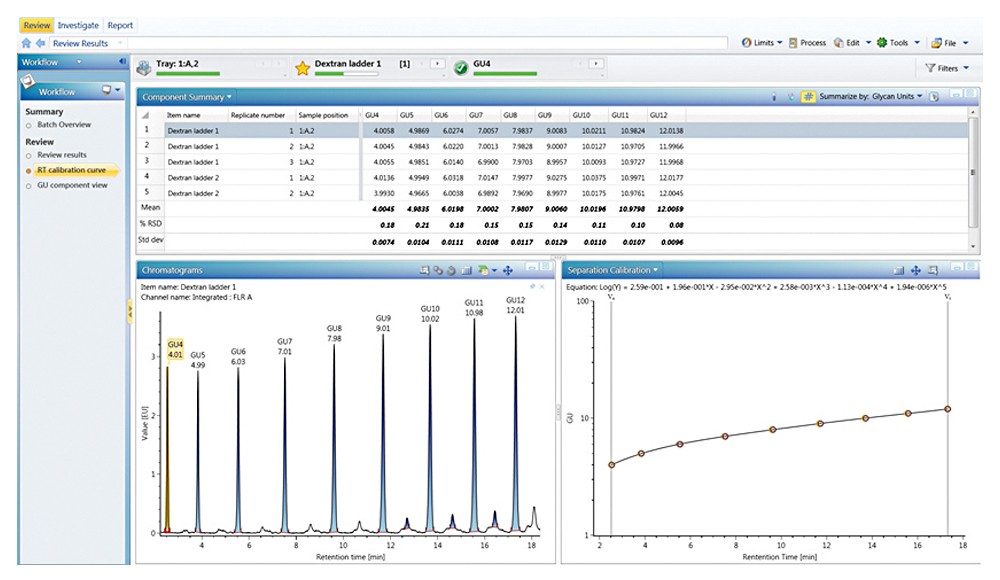
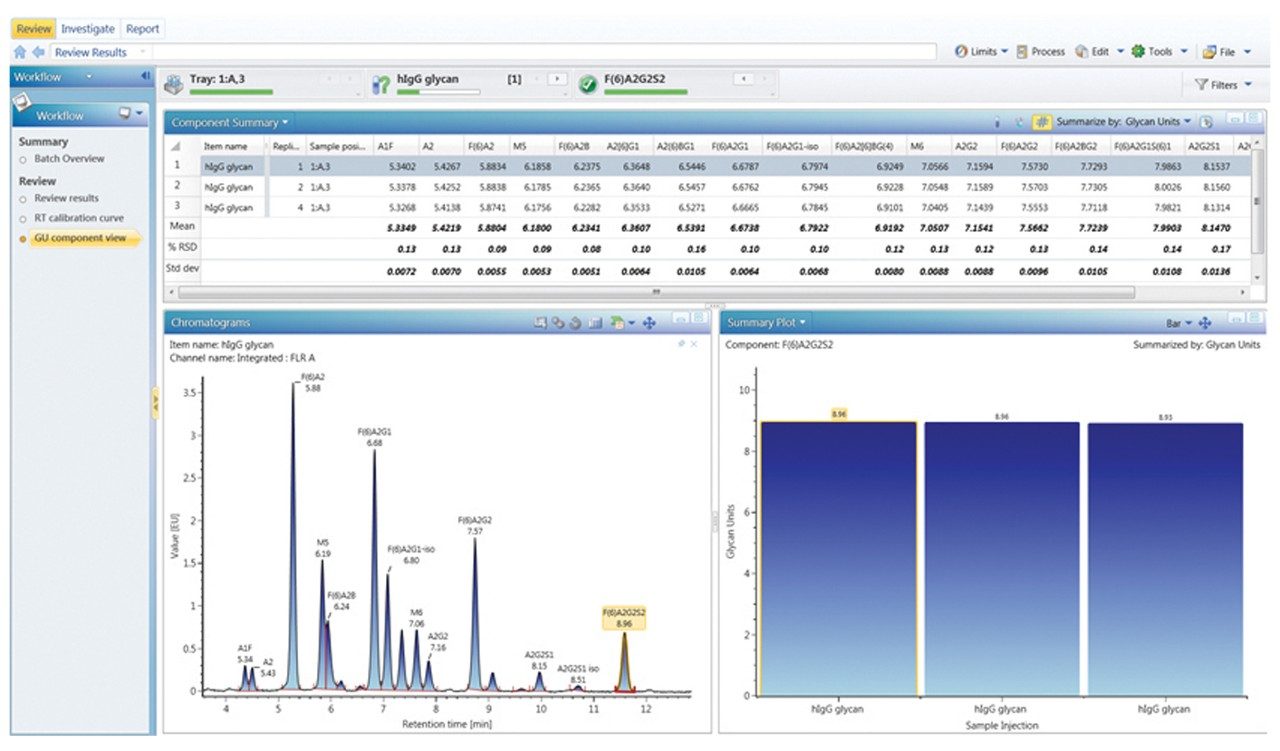
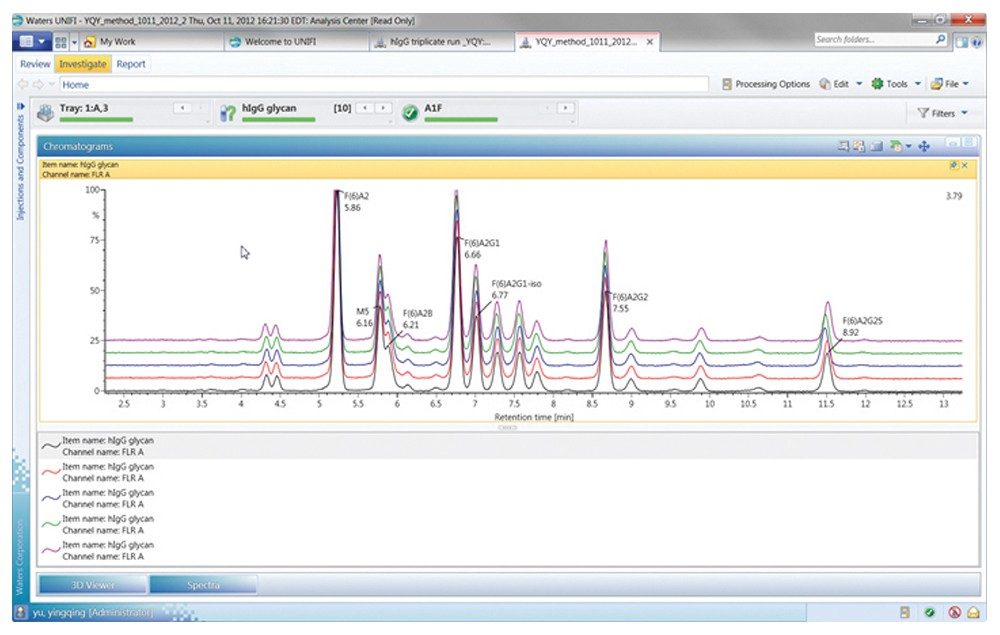
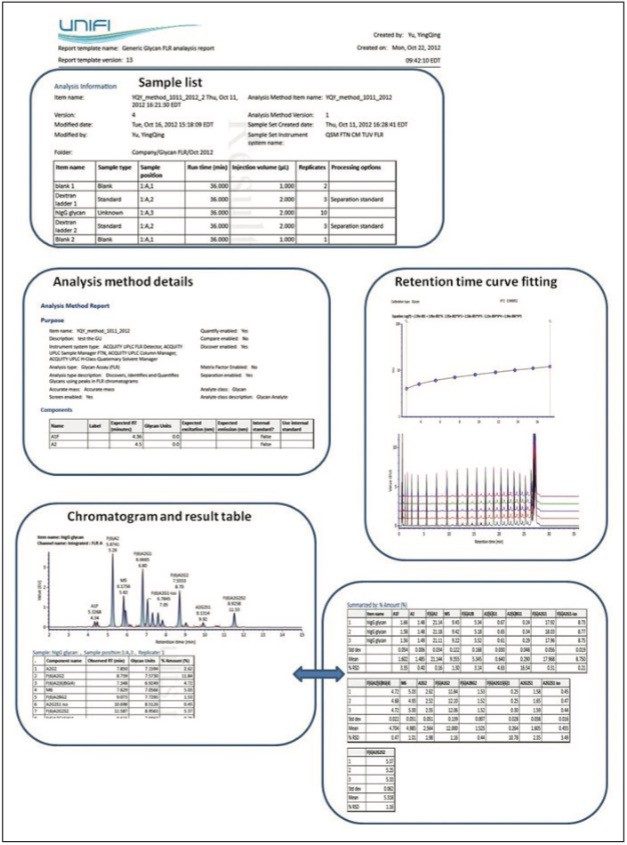
Figure 6 provides a snapshot of a typical glycan analysis report; information such as the sample list, retention time calibration curve fitting for 2-AB Dextran Calibration Ladder, and calculated GU values for unknown glycan samples are organized within single report template. Valuable information such as the relative abundance and the GU value for each glycan component were also summarized with appropriate statistical analysis.
UNIFI is the first comprehensive software that seamlessly integrates UPLC chromatography, optical detection, high resolution mass spectrometry, and integrated informatics within one platform. Released glycan analysis is one of the latest application workflows to be offered in UNIFI Software, v. 1.6.
This new UNIFI Application Solution enables a scientist in regulated or unregulated laboratory environments to acquire, process, and report qualitative and quantitative glycan information along with high confidence and minimal user intervention.
720004619, January 2016