In this application note, we propose a novel method of acquiring data called High Definition MSE (HDMSE). This method maintains the independent nature of PMF but extends its utility by providing molecular and MS/MS-like peptide sequence information in a simple, generic fashion
Peptide Mass Fingerprinting (PMF) is an unbiased protein identification technique that offers limited sequence information for identification purposes due to the reliance on the m/z measurement of proteolytic peptides.
An alternative technique commonly used conducts MS/MS sequencing experiments by fragmenting the tryptic peptides through sequential selection using the first analyzer of a mass spectrometer. This MS/MS approach has the following limitations:
In this application note, we propose a novel method of acquiring data called High Definition MSE (HDMSE). This method maintains the independent nature of PMF but extends its utility by providing molecular and MS/MS-like peptide sequence information in a simple, generic fashion. Unlike traditional MS/MS analyses, where selection of the precursor ions by the first mass analyser is used, ion mobility separation (IMS) is utilized as an orthogonal separation of peptides in the gas phase, followed by simultaneous fragmentation and high resolution mass analysis. Peptides are associated with their fragments for protein identification using ion mobility (drift time) alignment.
Protein extracted from normal kidney cortex was separated by 2D PAGE. Gels were stained with ProteoSilver Silver Stain, and 23 spots were excised and destained. Gel pieces underwent a classic tryptic digestion for 4 hours at 37 °C. Supernatants, with two further extractions, were pooled together and dried. These were reconstituted in 10 μL of 0.1% trifluoroacid aqueous solution. Samples were mixed with 5 mg/mL of α–cyano-4- hydroxycinnamic acid CHCA matrix and spotted onto a MALDI target.
|
MS system: |
MALDI SYNAPT G2 HDMS |
|
Ionization mode: |
Positive |
|
Mass range: |
100 to 3,000 Da |
|
Low energy transfer collision voltage: |
4 eV |
|
Elevated energy transfer collision voltage: |
30 to 199 eV |
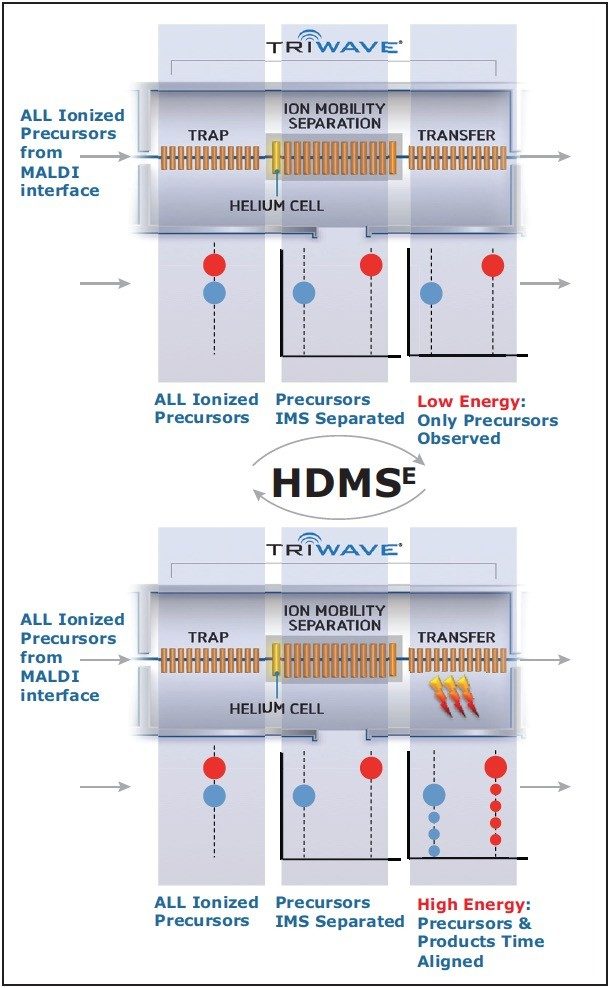
The first function was manually processed as a Peptide Mass Fingerprint (PMF) with only the m/z and intensities of the peptides submitted for a MASCOT database search.
Using the MSE Data Viewer software, all peaks in each function were detected using the Apex3D algorithm and aligned to correlate fragments to their precursor based on their drift time.
A peak list (.pkl) file was automatically created for each sample by combining all the precursors and their respective fragments.
The .pkl file was then submitted to a Mascot MS/MS ion search. Results were displayed using Protein Summary, where MS and MS/MS information contribute to the overall identification score.
Figure 2 shows an example plot of m/z versus drift time using DriftScope Software that displays both low and elevated functions in the case of the 2D-gel spot matching to ACTB_HUMAN.
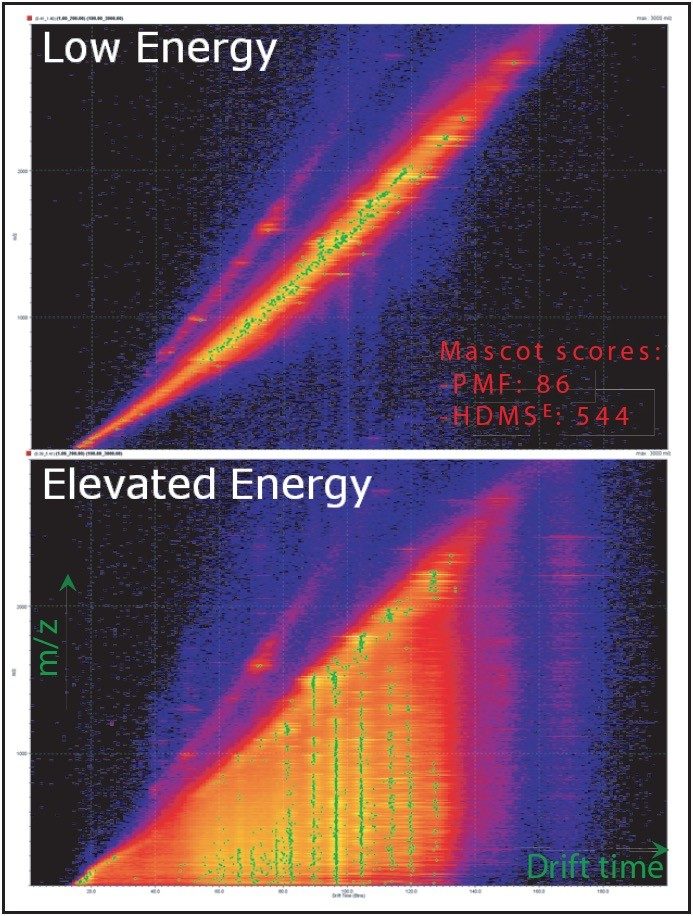
In the elevated energy plot, it is clearly seen that the fragments have similar drift time to their associated low-energy precursor ions.
Figure 3 shows the MASCOT scores for both PMF and MALDI-HDMSE analysis where identification was made in common.
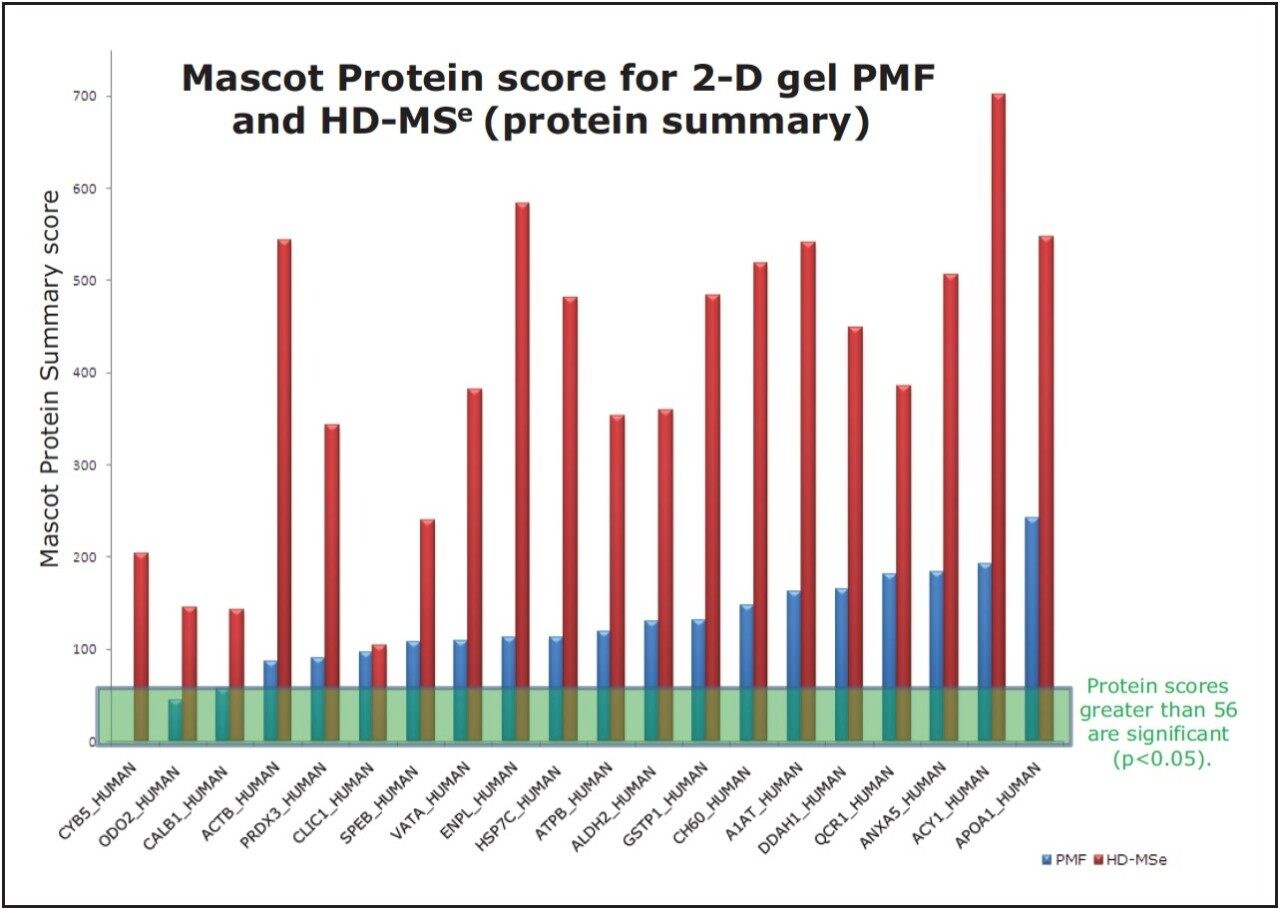
Out of the 23 2D-gel samples analyzed, 21 samples were identified by both PMF and HDMSE method, with just two that could not be identified by either method. For the remaining 21 samples, three samples were not identified above the 95% confidence level (in MASCOT) by PMF, whereas MALDI-HDMSE successfully identified these components above the confidence threshold. MALDI-HDMSE affords significant improvements to MASCOT identification scores for 20 out of the 21 identified samples.
Figure 4 displays the MSE Data Viewer data from the tryptic digest sample containing ACTB_HUMAN, with window B showing the low energy spectrum for the selected precursor, [M+H]+1790.8, within the drift time window.
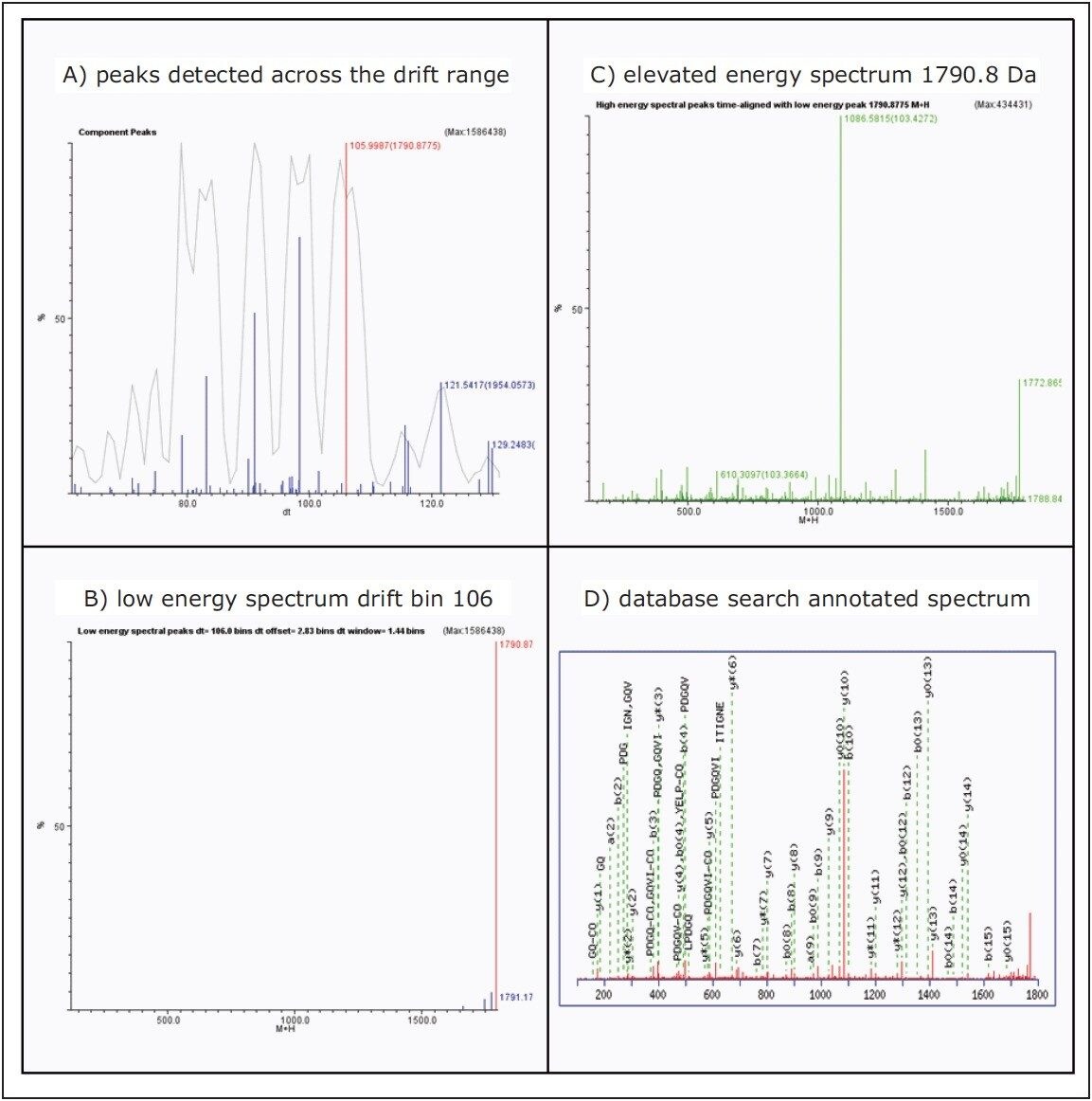
Window C is the elevated energy spectrum for the selected precursor that shows the associated fragment ions, more intuitive facilitation, and interactive results reviewing.
Window D shows the high number of fragments that have been used for scoring by MASCOT during protein database identification. The quality of the sudo MS/MS is clearly visible.
720004165, January 2012