Walk-up Mass Confirmation of Glucagon-like Peptides (GLP-1) Using the Xevo™ G3 QTof Combined with RemoteAnalyzer® Open Access Software
Chris Henrya, Isabella Jonesa, Scott Campbellb
a Waters Corporation, Wilmslow, United Kingdom
b SpectralWorks Limited, Runcorn, United Kingdom
Published on May 15, 2026
Abstract
This application note demonstrates quick, simple sample submission for the acquisition and reporting of deconvolved accurate mass data for GLP-1 analogues tirzepatide, liraglutide and semaglutide using the Xevo G3 Quadrupole Time-of-Flight (QTof) Mass Spectrometer. The system is fully controlled by the walk-up RemoteAnalyzer (SpectralWorks Ltd., Runcorn, UK) Software and fully integrated with the waters_connect™ Intact Mass Application. Both targeted and untargeted accurate mass confirmation experiments are accessible to expert and non-expert mass spectrometrists accelerating productivity during the drug development process.
Benefits
- Utilizes ballistic UPLC™ gradients for rapid sample turnaround.
- Walk-up solution for automatically deconvolved accurate mass confirmation of GLP-1 analogues for a wide range of mass spectrometry expertise.
- RemoteAnalyzer is compatible with both MassLynx™ and waters_connect instrument control environments.
- Processed data generated in the Intact Mass App for review if desired.
- Results automatically emailed for convenient data review at any location.
- Raw data available for UNIFI™ Application processing when desired.
Introduction
GLP-1 receptor agonists (GLP-1 RAs) are highly effective treatments for type 2 diabetes, and more recently, have gained significant attention as a treatment for obesity.1 This increase in demand has required pharmaceutical companies to reassess their strategies to meet the increased market opportunity and regulatory expectations. In parallel, drug manufacturers need consistent and rapid analytical methods to deliver drug products that meet the quality and regulatory standards.
Early-stage discovery of GLP-1 RAs relies heavily on mass spectrometry (MS) to confirm the molecular weight, structure, and purity of synthesized peptides and small molecules.2,3 As these compounds, such as GLP-1 (7–37) analogs, are often large (30+ amino acids), high-resolution mass spectrometry (HRMS) is critical to verify that the experimental molecular weight of the target molecule matches the theoretical weight. This process ensures the identity of the target peptide, distinguishes it from synthesis byproducts, and confirms the efficiency of the purification.
The need for continuous productivity increases in drug development departments has also led to an increased reliance on open access software.4 This is both to facilitate a simplified sample submission process and the receipt of condensed, easily digestible results for rapid decision making. Walk-up analysis software provides a simple, intuitive sample submission platform for a wide range of mass spectrometry expertise. Combining walk-up analysis software with ballistic UPLC™ gradients enables simple, efficient sample turnaround and decision making in mass confirmation experiments.
RemoteAnalyzer is a vendor neutral, browser-based open-access software that integrates with waters_connect and the Intact Mass App via an acquisition client and forms a seamless walk-up platform for GLP-1 analogue analysis. Samples can be submitted individually or as a batch using multiple plate configurations such as a 48 vial or 96-well plate.
For this workflow, samples are submitted via RemoteAnalyzer with the acquisition, processing, and reporting being carried out by waters_connect combined with the Intact Mass Application. RemoteAnalyzer retrieves the automatically generated result report, and emails to pre-designated recipients.
Both targeted and untargeted workflows are available using this platform. In this experiment, an untargeted workflow is chosen with the Intact Mass method incorporating expected peptide modifications with BayesSpray™ deconvolution determining the zero-charge state monoisotopic masses of the samples from the raw data generated.
In this work, a mass confirmation method is developed, designed to facilitate efficient, high-throughput analysis of GLP-1 RAs. By using the Xevo G3 QTof Mass Spectrometer controlled by RemoteAnalyzer Software (Figure 1), integrated with the Intact Mass Application, an intuitive walk-up method is demonstrated for rapid decision making in the synthesis of GLP-1 analogues.

Experimental
A 0.1 mg/mL solution of liraglutide, semaglutide and tirzepatide were prepared separately in 80:20 water:methanol. 0.1% v/v formic acid was added to each. The samples were placed in the autosampler and analyzed.
LC Conditions
|
LC system: |
Waters ACQUITY Premier UPLC I-Class System |
|
Vials: |
Waters QuanRecovery™ MaxPeak™ 12 x 32 mm Polypropylene 300 uL Screw Cap Vials (p/n: 186009186) |
|
Column: |
ACQUITY Premier BEH™ C18 1.7 µm 2.1 x 50 mm Column (p/n: 186009452) |
|
Column temperature: |
60 °C |
|
Sample temperature: |
6 °C |
|
Injection volume: |
0.5 µL |
|
Mobile phase A1: |
H2O 0.1% formic acid |
|
Mobile phase B1: |
Acetonitrile (ACN) 0.1% formic acid |
Gradient Table

MS Conditions
|
MS system: |
Waters Xevo G3 QTof Mass Spectrometer |
|
Acquisition mode: |
Sensitivity |
|
Mass range: |
50–2000 m/z |
|
Polarity: |
Positive |
|
Scan rate: |
10 Hz |
|
Cone voltage: |
30 V |
|
Source temperature: |
120 °C |
|
Desolvation temperature: |
350 °C |
|
Capillary voltage: |
2.8 kV |
|
MSE low energy (fixed): |
6 V |
|
MSE elevated energy ramp: |
30–55 V |
Results and Discussion
All samples were analyzed successfully in under five minutes with a high-throughput Intact Mass App report received by email (Figure 2), containing a UV aligned total ion current (TIC) and deconvolved spectra for each compound received by email (Figure 3). Highlighted in blue is the tirzepatide sample and the corresponding email delivered.

Example UV/TIC Aligned traces

BayesSpray Deconvolution
Using the BayesSpray algorithm within the Intact Mass processing application enables the deconvolution of a multiply charged species to a singly charged form. BayesSpray differs from maximum entropy (MaxEnt) deconvolution by using prior information provided in the form of sequence and modifications within the processing method to create the expected data and then comparing to the observed data. This approach is better suited to targeted workflows and produces cleaner spectra. In this case, the ‘targeted’ aspect refers to the potential modifications listed within the processing method. A resulting monoisotopic mass measurement can then be calculated as part of compound identification and characterization.
Deconvolved Sample Data
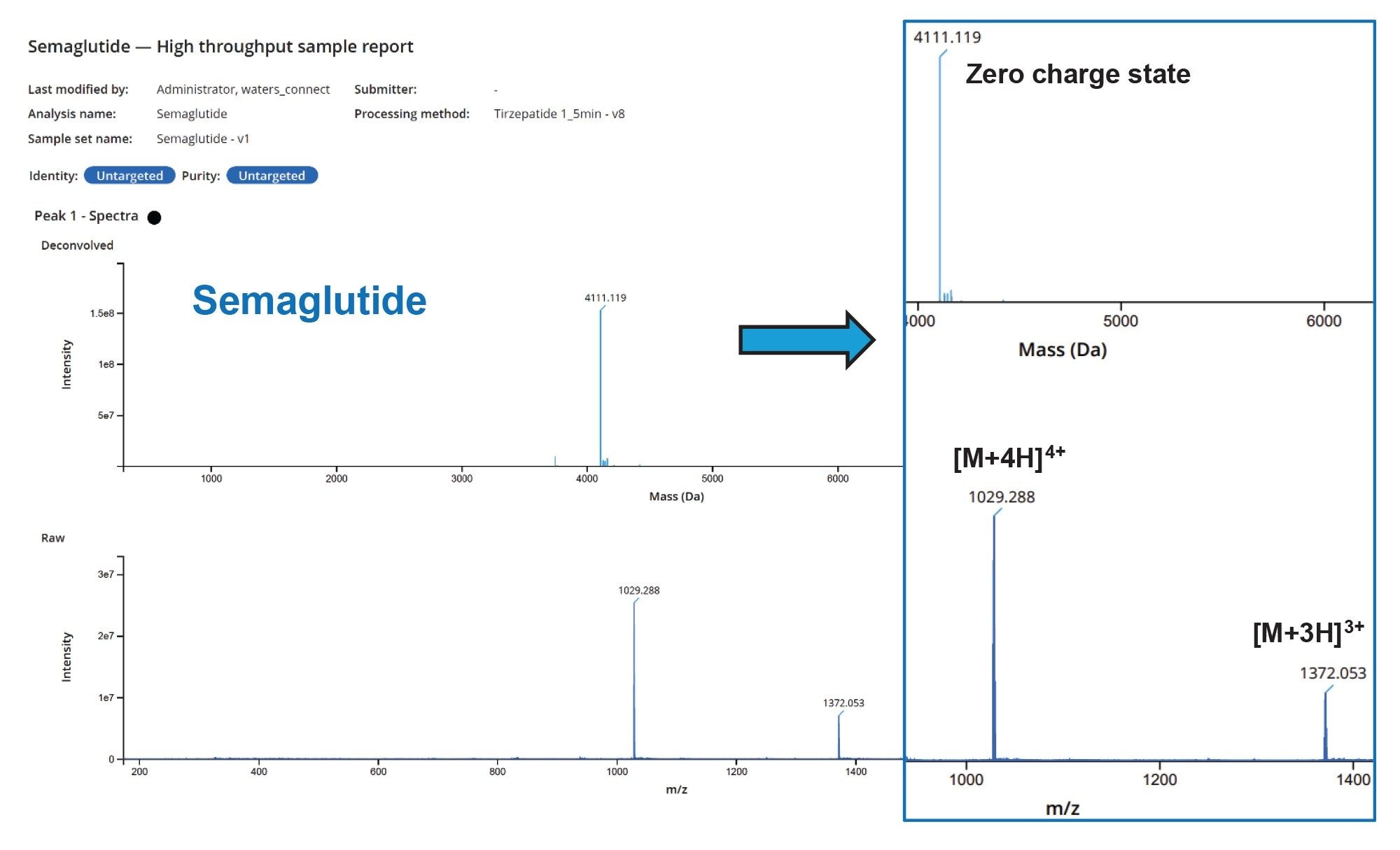
The deconvolution of the raw data returned for all three compounds showed excellent mass accuracy compared to the theoretical zero-charge state monoisotopic masses with parts per million (ppm) mass measurement accuracies of 0.9, 0.4, and 1.2 ppm for semaglutide, tirzepatide and liraglutide respectively.
Raw and deconvolved data are shown in Figures 4, 5, and 6.


Conclusion
This application note demonstrates a simple walk-up platform to deliver rapid accurate mass confirmation for the GLP-1 analogues semaglutide, tirzepatide and liraglutide. RemoteAnalyzer combined with the ACQUITY UPLC I-Class System coupled to the Xevo G3 QTof Mass Spectrometer delivers easy access to sub 2 ppm mass accuracy measurements using a 1.5 minute UPLC runtime in an open-access environment.
In addition, the ability to submit multiple samples as a batch and a short ballistic gradient provides a rapid throughput environment for efficient multi-experiment screening.
With the raw data remaining intact on UNIFI, further interrogation of the data is possible.
This approach provides drug development chemists with a flexible platform to make quick decisions on the success synthesis of a GLP-1 analogue.
References
- Alfaris, N.; Waldrop, S.; Johnson, V.; Boaventura, B.; Kendrick, K.; Stanford, F.C. “GLP-1 single, dual, and triple receptor agonists for treating type 2 diabetes and obesity: a narrative review”,eClinicalMedicine, Volume 75, 2024, 102782, ISSN 2589–5370, https://doi.org/10.1016/j.eclinm.2024.102782.
- Badgujar, D.; Bawake, S.; Sharma, N. A comprehensive study on the identification and characterization of major degradation products of synthetic liraglutide using liquid chromatography-high resolution mass spectrometry. J Pept Sci. 2025 Jan;31(1):e3652. doi: 10.1002/psc.3652. Epub 2024 Aug 20. PMID: 39162000.
- Jones, K. Tools and Techniques for GLP-1 Analysis. June 9, 2025. https://www.chromatographyonline.com/view/tools-and-techniques-for-glp-1-analysis
- Fontana, A.; et al. Automated open-access liquid chromatography high resolution mass spectrometry to support drug discovery projects. J Pharmaceut Biomed Anal. 2020;178:112908
Featured Products
720009354, May 2026