The purpose of this application note is to demonstrate a cost-effective LC-UV-MS-based workflow using ProMass with MassLynx for mass confirmation and impurity monitoring of synthetic peptides using the ACQUITY QDa Detector as an in-line orthogonal detector.
The market for peptide-based drugs is growing due to the broad range of activity and low toxicity of peptides.1 The use of solid-phase synthesis to produce the majority of peptide drugs often introduces process impurities such as incomplete deprotection of peptides and side reaction products.2,3 In addition, product-related impurities, such as oxidation and deamidation, add to the complexity of the impurity profile of the drug product.2,3 To ensure product efficacy and safety, the International Council for Harmonisation (ICH) guidelines require impurities to be identified based on toxicity and dosage levels of a new product.4 Optical thresholding based on LC-UV detection is commonly used to determine the impurity levels. However, optical detection can only be applied to known or pre-characterized samples, which typically require a laborious process of fractionation and enrichment followed by analysis using MS-based techniques for identification. The continuous growth in peptide-based therapeutics requires efficient workflows that can bring products to market in a timely manner while preserving product quality. Techniques that incorporate orthogonal detection methods, such as mass spectrometry (MS), offer the ability to address the challenges of impurity profiling in synthetic peptide production.
The ACQUITY QDa Detector offers an efficient and cost-effective solution to incorporating MS detection to LC-UV-based workflows.5 As demonstrated by previous work, the software ProMass (Novatia, LLC) works with MassLynx to provide an automated processing method for MS data generated by the ACQUITY QDa Detector and therefore enables high throughput screening for synthetic peptides.6 The purpose of this application note is to demonstrate a cost-effective LC-UV-MS-based workflow using ProMass with MassLynx for mass confirmation and impurity monitoring of synthetic peptides using the ACQUITY QDa Detector as an in-line orthogonal detector. To demonstrate this workflow, a clinically relevant synthetic peptide, eledoisin, is used for analysis.
The synthetic peptide eledoisin (pGlu-Pro-Ser-Lys-Asp-Ala-Phe-Ile-Gly-Leu-Met-NH2) was purchased from New England Peptide. HPLC grade water, acetonitrile, and formic acid were purchased from Fisher Scientific and used as received. A stock solution of the peptide was prepared at a concentration of 2 mg/mL in water and diluted to the desired concentration. Assay optimization was performed by varying the mass load of the peptide from 0.05 µg to 5.00 µg, while the injection volume was kept constant at 5 µL. A 2 µg sample load was found to be the optimal mass load and was used for all analyses unless otherwise noted.
|
LC system: |
ACQUITY UPLC H-Class Bio |
|
Detectors |
ACQUITY UPLC TUV Detector, 5 mm flow cell, λ = 215 nm ACQUITY QDa Detector (Performance model) |
|
LC column |
ACQUITY UPLC CSH C18, 1.7 μm, 130 Å, 2.1 mm x 100 mm |
|
Column temp. |
60 °C |
|
Sample vial |
12 mm x 32 mm glass vial, Total recovery |
|
Injection volume |
5 μL |
|
Mobile phase A |
H2O, 0.1% formic acid |
|
Mobile phase B |
Acetonitrile, 0.1% formic acid |
|
Time (min) |
Flow rate (min) |
%A |
%B |
|---|---|---|---|
|
Initial |
0.2 |
82 |
18 |
|
2 |
0.2 |
82 |
18 |
|
2.01 |
0.2 |
82 |
18 |
|
22.01 |
0.2 |
72 |
28 |
|
22.02 |
0.2 |
20 |
80 |
|
24.02 |
0.2 |
20 |
80 |
|
24.03 |
0.2 |
82 |
18 |
|
30 |
0.2 |
82 |
18 |
|
Mass range: |
350–1,250 m/z |
|
Mode: |
ESI Positive |
|
Collection mode: |
Continuum |
|
Sample rate: |
2 points/sec |
|
Cone voltage: |
10 V |
|
Probe temp.: |
500 °C |
|
Capillary voltage: |
1.5 kV |
|
Data management: |
MassLynx v 4.1 ProMass for MassLynx |
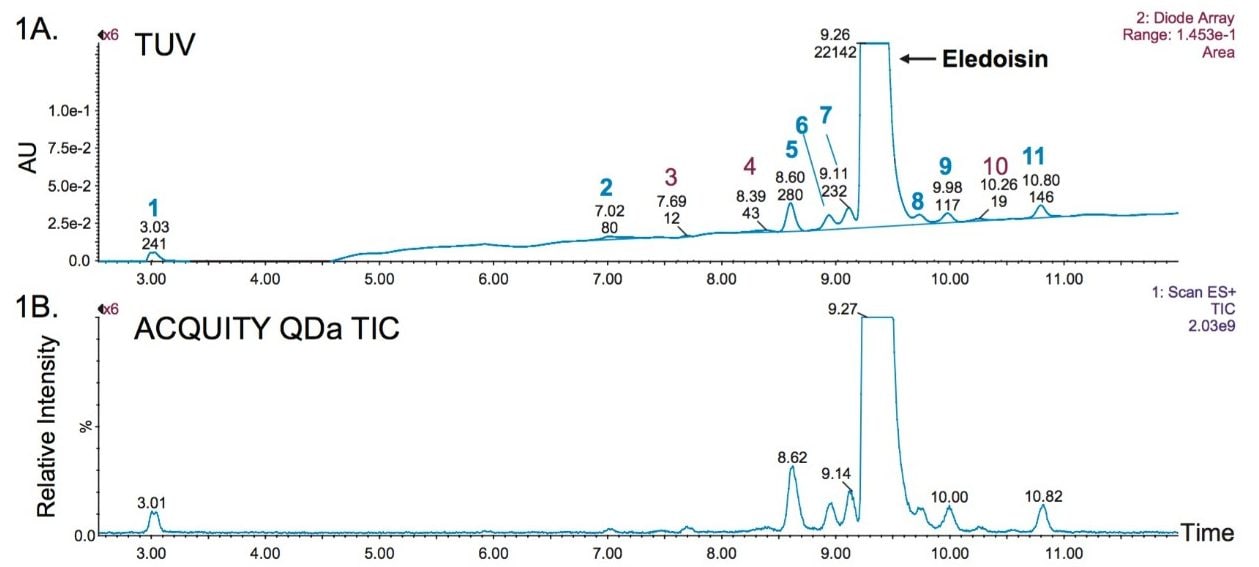
Synthetic peptide impurity profiling via LC-UV-based techniques requires optimization of peak resolution and detector response. For this study, a 20 minute gradient was found to resolve 11 impurity peaks from the main peak as shown in Figure 1. ICH guidelines recommend impurity identification to be based on total daily intake. For this study, a cut-off of 0.2% relative abundance was used along with S/N ≥ 10 to determine the broadest applicability of the method. Eight of the 11 peaks shown in Figure 1 were identified as peaks meeting this criteria based on automatic peak integration performed by MassLynx and are annotated in blue. Peaks above the optical cut-off of S/N ratio were all observed in the MS spectrum (Figure 1B).
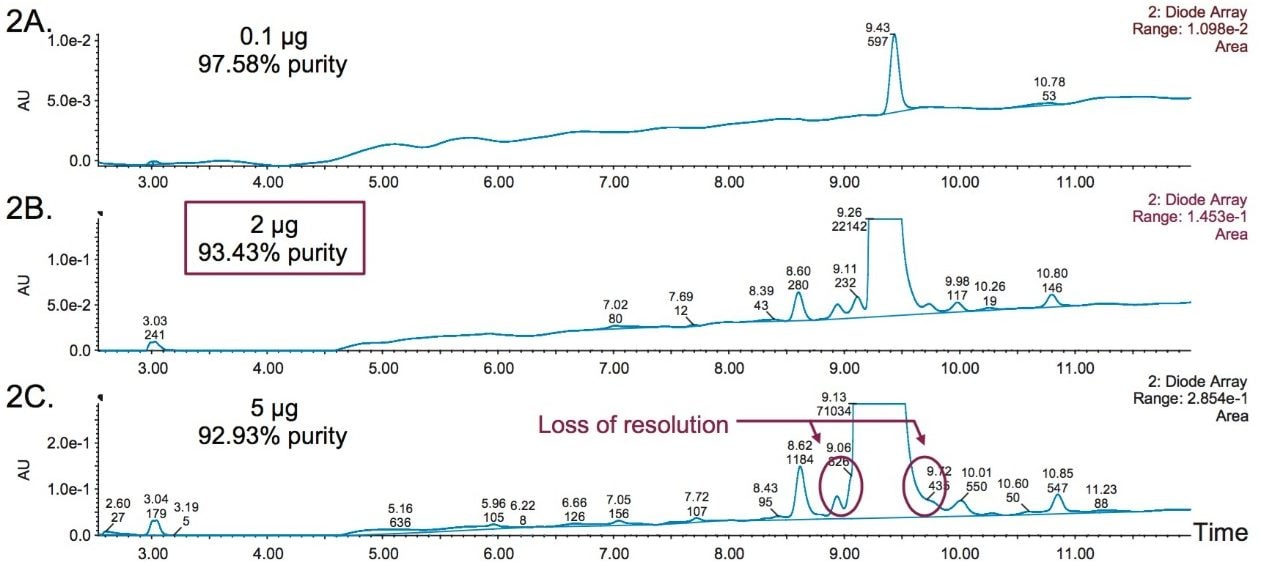
Percent purity determined by mass load was used in lieu of a spiking study to determine the working range of the assay. Ten samples were prepared with increasing concentration (from 0.01 mg/mL to 1.0 mg/mL) and evaluated with the optimized gradient. Purity assessment based on total peaks detected in the optical trace was determined to be stable above mass loads of 0.5 µg at 92.74 % ± 0.45%. To evaluate assay precision, three replicate runs were acquired to measure the %RSD of purity. Results showed the %RSD was below 5% at all mass loads. Using the combined data it was determined that a 2 µg mass load offered optimal chromatographic performance while maintaining an S/N ≥10 for eledoisin and its impurities as shown in Figure 2.
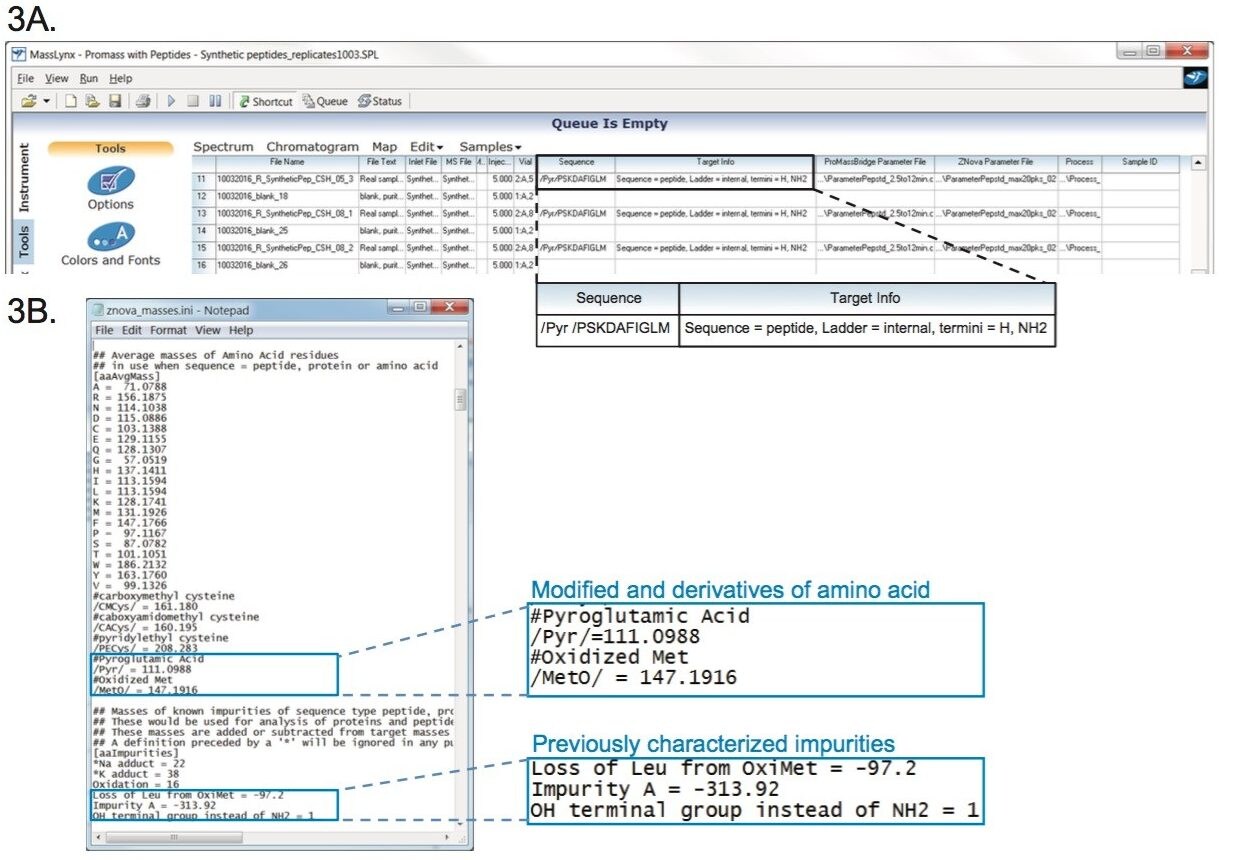
With the optimized method determined, the benefit of mass detection can now be fully utilized. The ACQUITY QDa Detector enables the software to determine mass difference between impurities and the target peptide for impurity identification. Automated data acquisition and processing of synthetic peptides can be performed by MassLynx when used in conjunction with ProMass. Figure 3A is an example of the modified Sample List in MassLynx, which contains the instrument methods for data acquisition, as well as information needed for target mass calculation, and parameter files for data deconvolution of synthetic peptides using ProMass. For peptide impurity profiling, the information of the target peptide needs to be entered in the Sequence column as well as the desired Target Info as shown in the zoom-in image of Figure 3A. In this example, ‘Ladder = internal’ was used to detect all possible fragments with either termini. The target synthetic peptide, eledoisin, has a primary amine group (NH2) at the C-terminus rather than an OH, thus requiring the termini to be specified as such for correct mass determination. Parameters for MS spectra analysis are defined by the files entered into the ProMassBridge Parameter File column and the Znova Parameter file column. The ProMassBridge Parameter File enables communication between ProMass and MassLynx, and contains user definable parameters such as the retention time window to process, smooth functions and baseline subtraction. As part of the ProMass software, the Znova™ Parameter file provides parameters for spectrum deconvolution, and thresholding for streamlining reporting.
The Znova processing file contains a customizable database for target identification and impurity profiling as shown in Figure 3B. Modified and derivatized amino acids can be entered along with naturally occurring amino acids as part of the “building-blocks” for calculating target masses. In the case of eledoisin, Pyroglutamic Acid (/Pyr/=111.1) was entered to account for the amino acid derivative at the terminus of the synthetic peptide. Similarly, previously characterized impurities can be manually entered as their calculated mass differences from the target peak. As shown in the blue rectangle of Figure 3B, deletion of leucine from the oxidized target peptide (loss of Leu from OxiMet = -97.2), possible peptide fragment (Impurity A = -313.9), and C-terminal substitution (OH terminal group instead of NH2 = 1) were previously identified by high resolution MS prior to this study. Matching mass differences will be shown in deconvolution reports along with their identities.
Using the method described above, an analysis of eledoisin was performed for target mass confirmation and impurity screening. Purity of the sample was reported to be 88.87%, which is estimated based on percent abundance of the target mass (including adducts) in the MS spectrum and normalized by chromatographic peak area percent. Percent purity can be found in the Target Mass Summary table, along with other information of the target peptide such as mass error and percent abundance (Figure 4). The sequence Ladder Summary table shows the sequence of identified peptide fragments. The Chromatogram Summary table lists all detected peaks above threshold (0.2%) across the chromatogram. The Spectral Quality, which is based on abundance and the number of charge states detected, was reported “ok” for all impurities with the exception of peaks 4 and 5 which were below the cutoff value (Score ≥4). In this example, the Low Score warning at 8.40 min (peak 4) results from a high degree of spectral noise, and the one at 8.62 min (peak 5) is from the misidentification of the base peak charge due to only one charge state being present in the spectrum. The Spectrum Quality threshold, which can be set by the user, enhances the confidence of impurity identification results by indicating the quality of both input and output spectra for deconvolution.
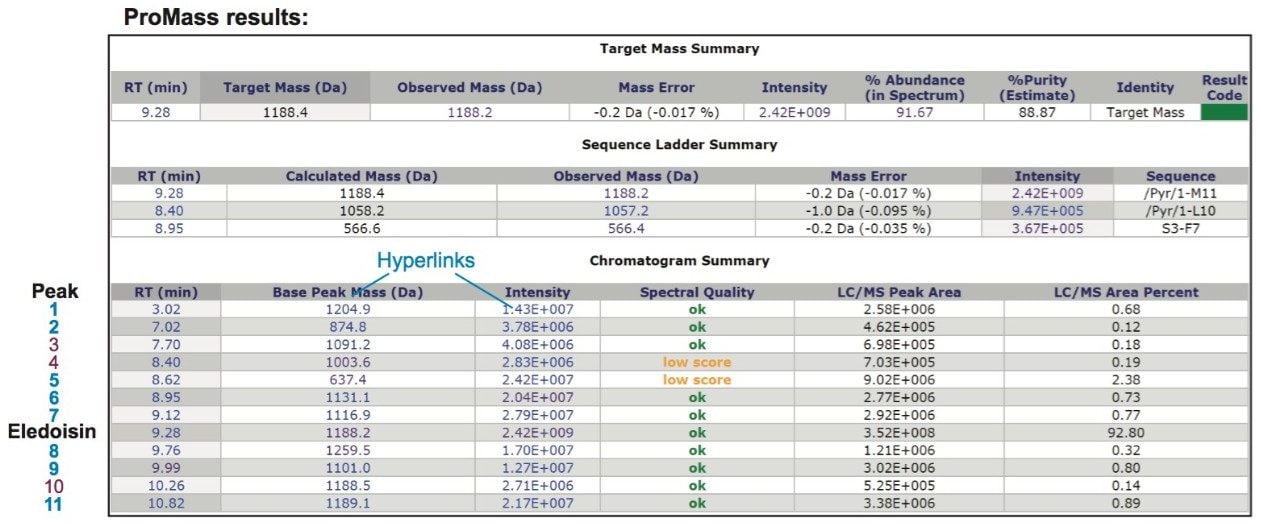
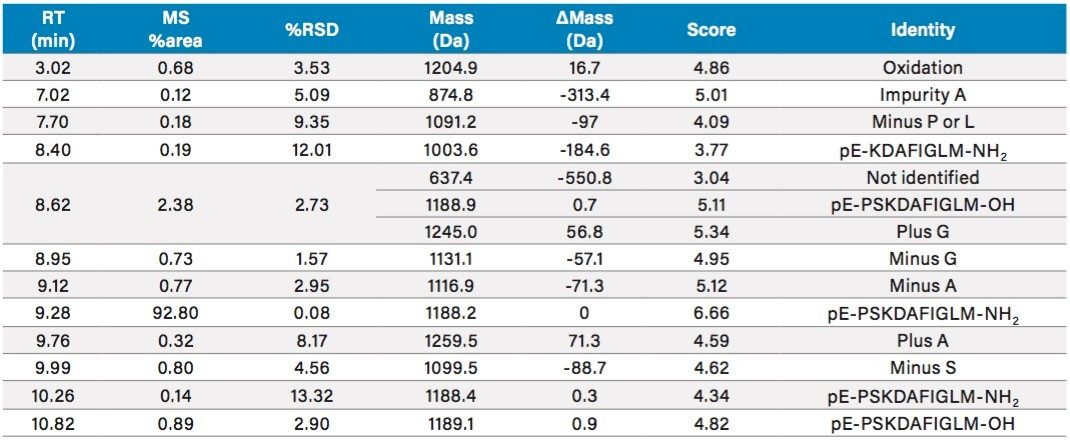
As part of the ProMass software, additional details can be obtained for each analysis by clicking the hyperlinks in Chromatogram Summary table. An example for the peaks at 9.99 min is shown in Figure 5. The top panel shows a list of peaks with their scores and presumed identities. The bottom panel shows the deconvoluted mass spectrum for all detected peaks at 9.99 min. In this list, it is suggested that the mass 1101.0 Da could be a deletion of Serine (S), from the target peptide. Data processing results obtained from ProMass are summarized in Table 1, including retention time, percent area and its %RSD, mass, mass difference between impurity and the target peptide, and presumed identity of detected impurities. The identification of impurities were highly reproducible for all monitored impurities in three replicate runs demonstrating the reliability of LC-MS based workflows in impurity profiling of synthetic peptides.

An automated workflow of synthetic peptide mass confirmation and impurities profiling was developed using the ACQUITY QDa Detector with MassLynx and ProMass. Impurities above 0.2% optical threshold were successfully detected and identified for eledoisin at 2 µg mass load. Retention time, mass, and presumed identity of both the target peptide and impurities were summarized in ProMass deconvolution summary tables and reports. Confidence of deconvolution was represented as a Score for each peak. The addition of the ACQUITY QDa Detector to an optical workflow significantly reduces the time of impurity analysis. With ProMass for MassLynx, the automated workflow allows high throughput screening and therefore improves the productivity of synthetic peptide impurity profiling in development and quality control environments.
720005921, February 2017