The automated analysis of a set of peptide mapping samples that have been spiked with varying levels of a peptide impurity, using the Biopharmaceutical Platform Solution with UNIFI is demonstrated in this application note.
The ability to acquire, process, and report LC-MS peptide mapping data has been integrated and automated using the Biopharmaceutical Platform Solution with the UNIFI. This enables laboratories to generate communicable information directly following sample analysis or data review. The UNIFI Scientific Information System organizes data and information and simplifies the review of large amounts of complex peptide mapping results, limiting opportunities for human error, and saving untold hours of formerly manual effort that go into producing the results and reports needed to advance molecules through the development process.
Characterization of protein biopharmaceuticals requires the analysis of primary structure and thorough investigation of sequence variants. Confirmation of protein sequence, quantification of minute changes in the protein covalent structure, and assessment of disulfide bond patterns are routinely performed using LC/UV-MS with the assistance of specialized informatics tools. Most of these informatics tools for the biopharmaceutical laboratory are designed for facilitating intense characterization of novel samples, not for higher throughput data processing and results management for large sets of routine samples.
Increases in peptide mapping samples from increasingly complex quality by design (QbD) methods, formulations, stability, and clone screening workflows have created the need for larger batch processing and reporting of routine analytical studies within a biopharmaceutical laboratory. The Biopharmaceutical Platform Solution with UNIFI merges robust UPLC/UV/Tof-MS characterization technologies with automated data acquisition, processing, and reporting workflows. The platform is designed to address the challenges of routine analysis with large sample sets in either regulatory compliant or non-compliant
laboratory organizations.
This application note details the automated analysis of a set of peptide mapping samples that have been spiked with varying levels of a peptide impurity, using the Biopharmaceutical Platform Solution with UNIFI. Such controlled studies are useful for verifying system performance, and for establishing expectations for subsequent studies conducted with unknown samples. In addition to describing the results from this spiking study, we focus on demonstrating the tools that enable efficient review and reporting of peptide mapping data within UNIFI.
|
System: |
ACQUITY UPLC H-Class |
|
Column: |
ACQUITY UPLC BEH130 C18, 2.1 x 100 mm, 1.7 μm |
|
Column temp.: |
65 °C |
|
Mobile phase A: |
Water |
|
Mobile phase B: |
Acetonitrile |
|
Mobile phase C: |
1% formic acid |
|
Detection: |
UV 214 nm |
|
Injection vol.: |
10 μL |
|
Time(min) |
Flow(mL/min) |
%A |
%B |
%C |
%D |
Curve |
|---|---|---|---|---|---|---|
|
Initial |
0.2 |
89 |
1 |
10 |
0 |
Initial |
|
90.00 |
0.2 |
35 |
55 |
10 |
0 |
6 |
|
91.00 |
0.2 |
10 |
80 |
10 |
0 |
6 |
|
94.00 |
0.2 |
10 |
80 |
10 |
0 |
6 |
|
95.00 |
0.2 |
89 |
80 |
10 |
0 |
6 |
|
115.00 |
0.2 |
89 |
1 |
10 |
0 |
6 |
|
Capillary: |
3.0 kV |
|
Sampling cone: |
25 V |
|
Extraction cone: |
2.0 V |
|
Source temp.: |
100 °C |
|
Desolvation temp.: |
250 °C |
|
Cone gas flow: |
0 L/Hr |
|
Desolvation gas flow: |
500 L/Hr |
|
Scan time: |
0.5 sec |
|
Low energy: |
4 V |
|
High energy ramp: |
25 V to 40 V |
|
Mass range acquisition: |
100 to 2000 m/z |
|
LockMass compound: |
GFP 500 fm/μL, 10 sec interval, 0.5 sec scan time |
UNIFI Scientific Information System
Denatured trastuzumab (2.1 mg/ml, 8 M guanidine chloride, 1 M Tris, pH 7.5) was reduced with 0.5 M DTT and alkylated with 0.5 M iodoacetamide. Buffer exchange (100 mM Tris, pH 7.5) over a NAP-5 column (GE Healthcare) enabled efficient tryptic digestion (Sample: Enzyme = 42:1, 4 hrs at 37 °C) for subsequent analysis by peptide mapping.
[Glu1]-Fibrinopeptide B (GFP) (Sigma) peptide was prepared at 6.4 pmol/μL and 160 fmol/μL in 0.1% formic acid. A dilution series was made using the concentrated trastuzumab digest (19.6 pmol/μL) and GFP solution, then adjusted to final concentrations of 3% acetonitrile and 0.1% TFA, prior to analysis.
GFP peptide was spiked at 0.0% (digest control at 10 pmol/μL), 0.1% GFP (10 fmol/μL), 0.25%, 0.5%, 2.5%, 5.0%, 10% (1 pmol/μL).
In this study, a trastuzumab tryptic digest spiked with GFP enabled us to evaluate the performance of automated UPLC-MSE peptide mapping workflows within the Biopharmaceutical Platform Solution with UNIFI. To generate the data for this assessment, the trastuzumab digest was supplemented with spiked GFP peptide spanning two orders of magnitude (0.1% to 10% GFP mol/mol) dynamic range, with each level of spiked sample analyzed in triplicate.
UNIFI Software includes a workflow that is specific for accurate mass peptide mapping, constructed of a holistic method that combines UPLC-MSE acquisition settings, data processing parameters, bioinformatic assignment criteria, and linkage to templates for automated report generation. In this workflow, peptides are assigned by accurate mass, confirmed by accurate mass fragmentation, and assigned to peptides or modified peptides generated by in silico digests of one or more proteins defined within the method. Applying this methodology, the GFP peptide was identified and quantified using the automated processing and bioinformatics parameters down to the lowest levels studied (0.1% relative to trastuzumab peptide levels).
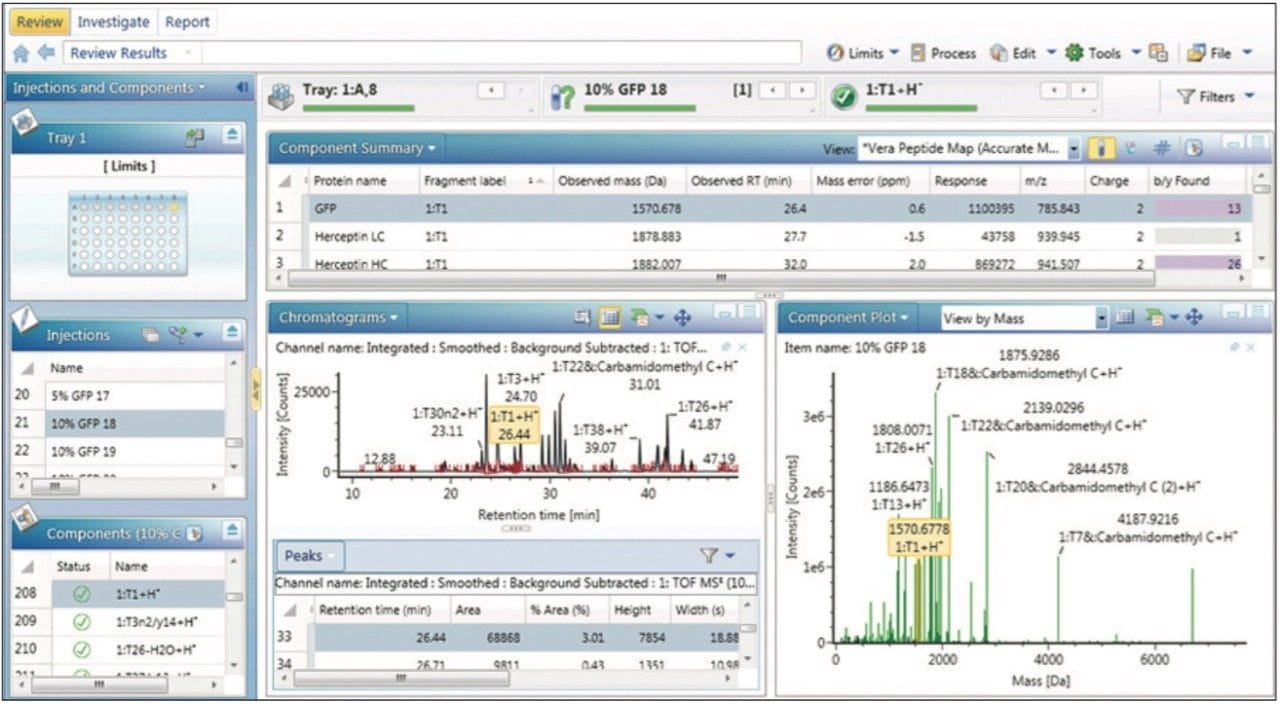
The Analysis Center in UNIFI Software provides a central hub to review processed data (Review panel), interrogate the underlying raw data (Investigate panel), examine reports (Report panel), and monitor analytical systems (Acquire panel). The Review panel for the spiked digestion experiment (Figure 1) enables scientists to format processed results to focus on questions about individual samples, to compare binary samples, or to summarize results for specific components across larger sets of data.
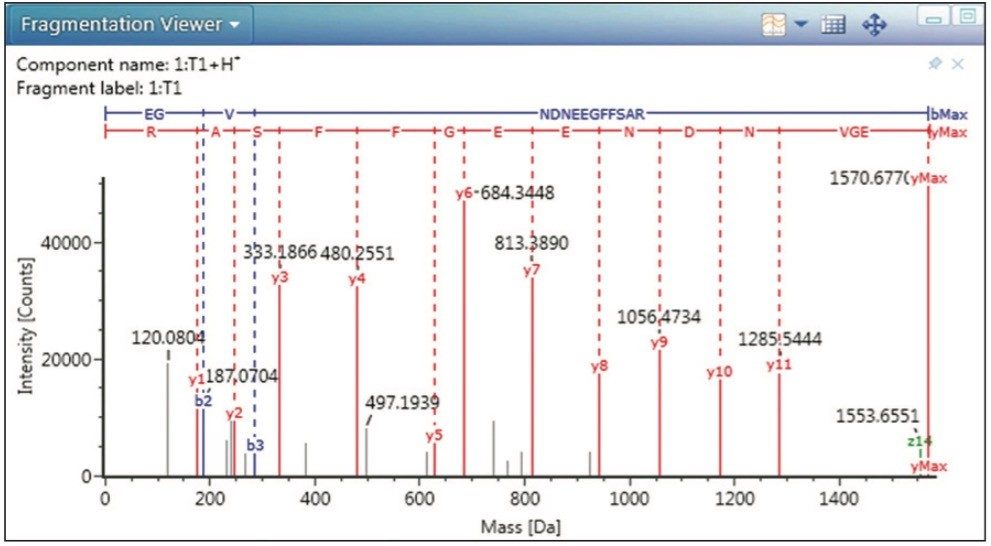
The spiked GFP peptide is displayed in Figure 3 as the undigested tryptic peptide, T1. Selecting this peptide in the main data table or the associated chromatogram or component plots highlights that component on all the visible panes and enables a researcher to easily scroll through multiple data sets for data relevant to that component. These plots are highly configurable and alternative displays can be used to answer other scientific questions. For example, selecting the Fragmentation Viewer (Figure 2) displays the annotated MSE spectrum of GFP, enabling the researcher to verify an accurate mass peptide assignment with corresponding characteristic fragment ion information.
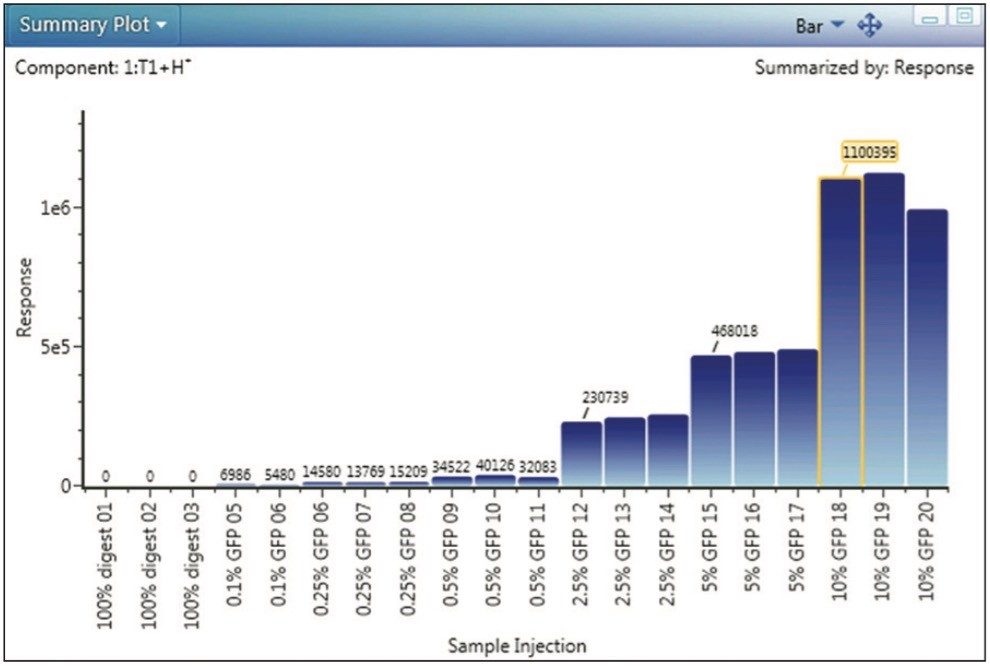
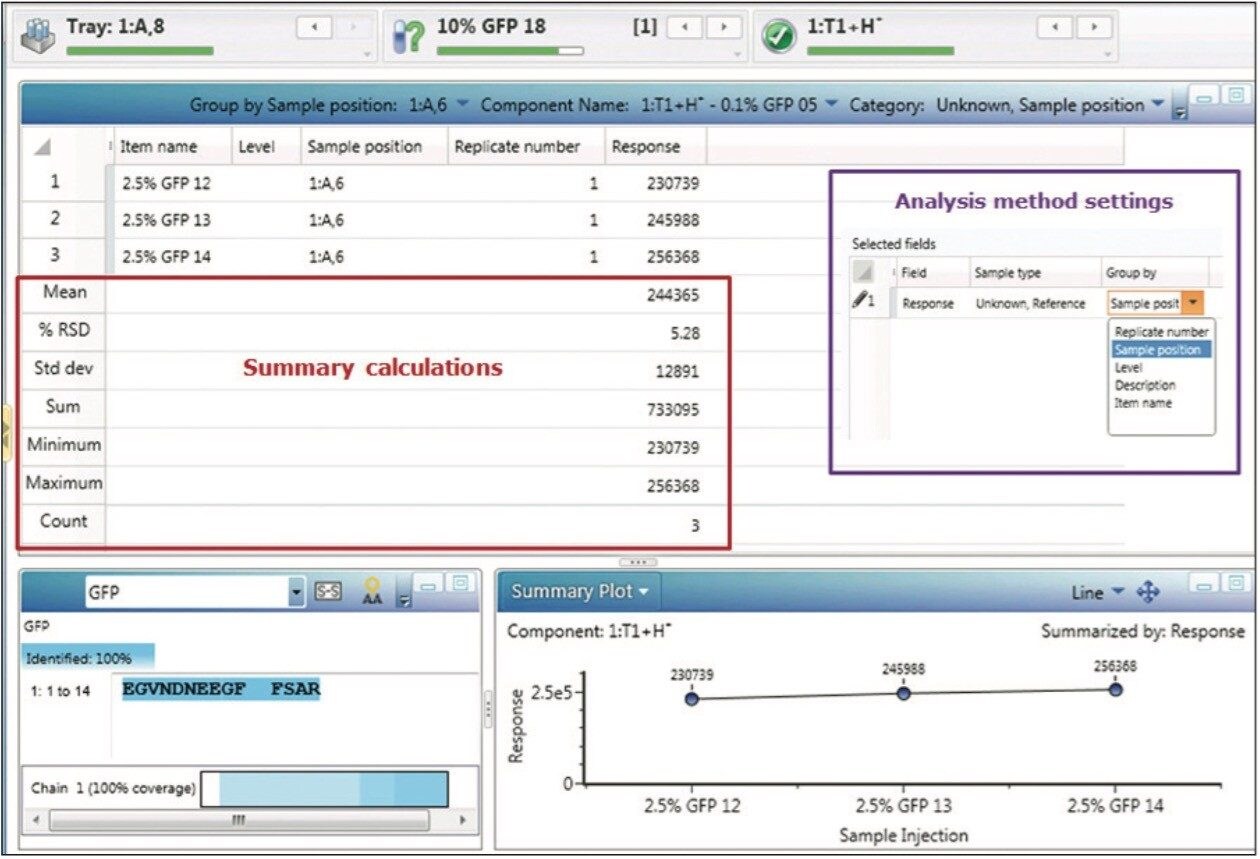
The GFP peptide summary plot (Figure 3) is one of the most informative displays addressing the central questions of the spiked impurity study. The Summary Plot shown in Figure 3 depicts total MS response for the GFP peptide across all experimental samples, on a per-injection basis, and enables us to see that response is proportional to spiked peptide level, and that detection and quantitation was robust across the entire analysis.
This ability to focus and summarize results from large sample sets goes beyond the summary plot, as injection replicates, sample replicates, or other data groupings can be utilized to generate tabular result summaries, along with appropriate statistical treatments. These summaries can be generated routinely during automated sample processing (Figure 4, purple box), and displayed within the Review pane.
Adjusting the main data table from a “Component Summary” to “Results Summary” focus enables the summarized display of grouped results for one or more component fields. Figure 4 displays summary results for the GFP peptide for the three injections of the 2.5% spiked sample, enabling automated calculation of average and StdDev and %RSD for this particular triplicate analysis. Executing the tasks of data organization and summarization as part of automated data processing and reporting enables organizations to avoid the transcription and other human errors common with the use of external software packages, and reduces the organizational costs of validating the external tools needed to accomplish this task.
Reporting templates are available for each specific Application Solution (peptide, protein, and glycan analysis) in UNIFI Software. These templates are composed of the individual report objects, which can be rearranged, filtered, and modified to automatically generate the tables, chromatograms, plots and spectra currently used in a researcher’s organization. The object properties as well as the whole report template are configurable by the user, and multiple report templates can be executed for an analysis to answer multiple scientific questions in an efficient format.
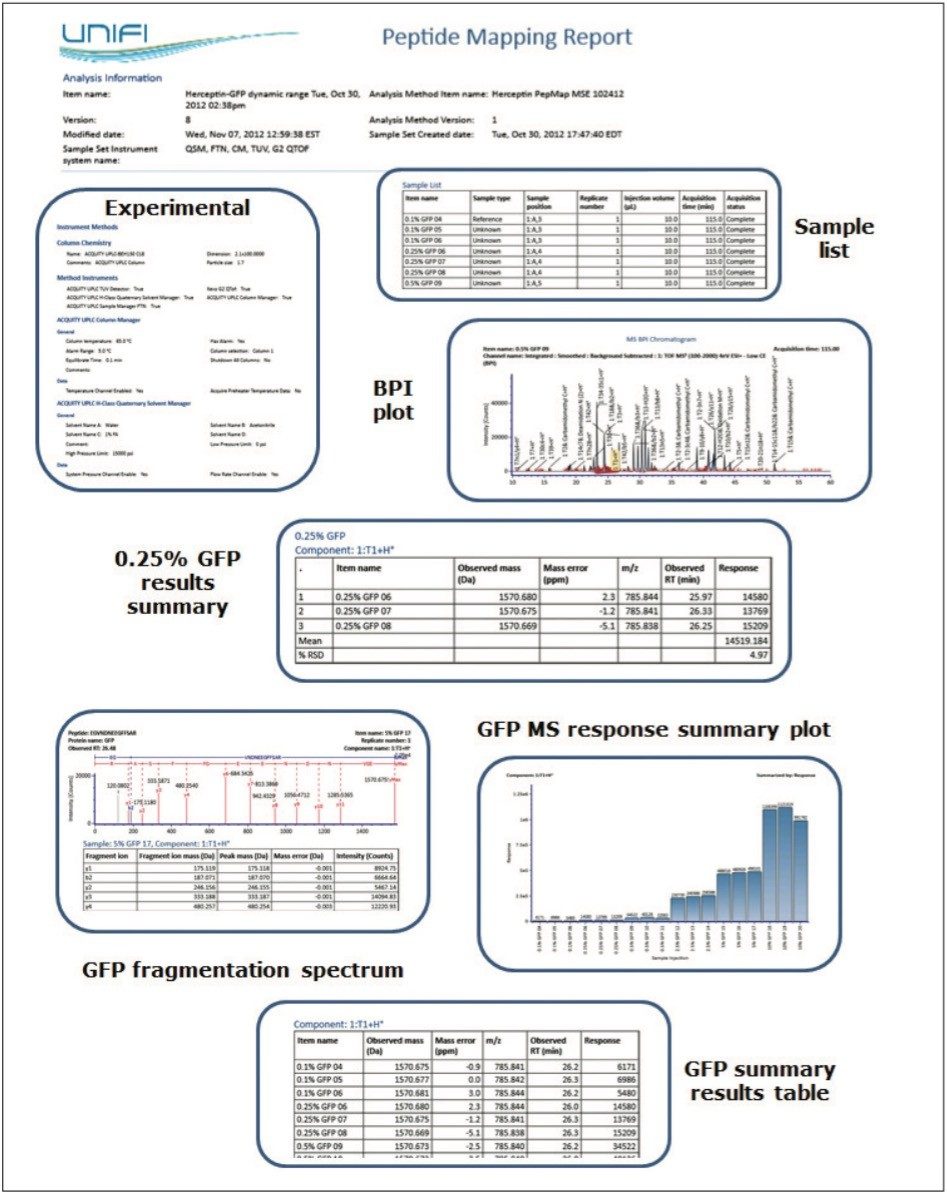
A collection of report elements appropriate for the spiked peptide map experiment are displayed within Figure 5. These objects enable a researcher to communicate experimental settings, chromatographic and mass spectral results for individual samples, and the component-focused result summaries available as part of the Review panel functionality described above.
Going from “big data” to communicable information often represents the largest bottleneck in the operations of many biopharmaceutical analytical laboratories. The ability of the Biopharmaceutical Platform Solution with UNIFI to standardize and automate the generation of such information, in both regulated and non-regulated laboratories, enables these organizations to address routine analysis with greater efficiency and effectiveness.
The automated UNIFI peptide mapping workflow was able to identify a spiked peptide impurity down to the lowest concentration tested (0.1% with 100 fmol of digest loaded on-column). Automated trending and summary statistical calculations executed within UNIFI Software facilitated rapid data review and efficient reporting of the key results from this study.
These results confirmed the utility and reproducibility of UPLC/UV/Tof-MS peptide mapping studies, and their capability to yield quantitative results with high precision and linearity of the MS response.
The ability to integrate and automate such peptide mapping workflows should greatly improve the efficiency of laboratories within regulated and non-regulated biopharmaceutical organizations that are challenged with interpreting complex peptide mapping studies.
720004620, February 2013