This is an Application Brief and does not contain a detailed Experimental section.
This application brief enables the systematic analysis of fluorescently labeled N-glycan pools released from recombinant biotherapeutics and separated by HILIC-UPLC chromatography. Data are interpreted with the aid of a robust online glycan database generated from standardized retention values.
NIBRT’s GlycoBase 3+ is a web-enabled proprietary resource that contains normalized retention data, expressed as Glucose Units, or GU values, for more than 600 2AB-labeled N-linked glycan structures.
Glycosylation is the most complex posttranslational protein modification and it is estimated that more than half of all eukaryotic proteins are glycosylated. In order to elucidate the functional outcomes of glycosylation and to characterize the glycoprotein more fully, it is essential to define the monosaccharide sequence, linkage, and anomericity of the covalently attached oligosaccharides.
Glycosylation of biotherapeutics is a critical product attribute and adverse changes in glycosylation can significantly alter product efficacy and safety. Furthermore glycosylation of biotherapeutics can be influenced by dissolved oxygen, pH, carbon source, temperature during manufacture, as well as by the choice of expression system. These variations can put product integrity at risk and therefore the ability to monitor glycosylation accurately and rapidly is essential at all stages of the process.
The U.S. FDA, the EMA, and other regulatory bodies have started to increase pressure on manufacturers to analyze the glycosylation of therapeutics more fully, and also to demonstrate how their process can affect glycan composition. However, there are no robust workflows for systematic detailed, quantitative, sensitive, glycan analysis.
The National Institute for Bioprocessing Research and Training (NIBRT) in Ireland is a world-class institute that provides training and research in the biopharmaceutical industry. NIBRT’s mission is to provide a unique experience for trainees in an environment that replicates a modern bioprocessing facility. In parallel, NIBRT also undertakes leading bioprocessing research in collaboration with industry partners and provides contract analytical services.
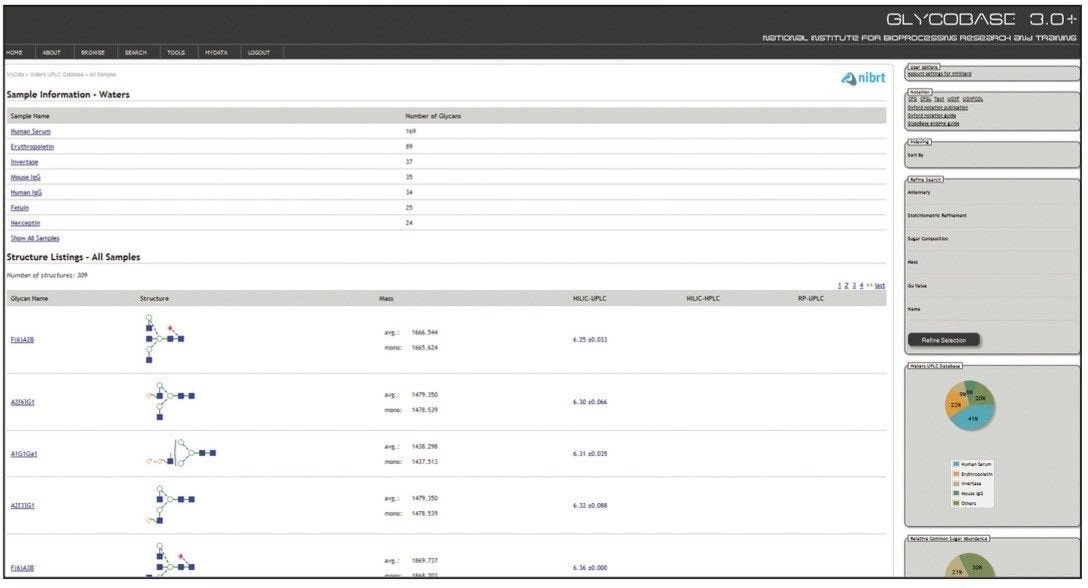
Waters Corporation and NIBRT, led by Prof. Pauline Rudd, have developed a comprehensive solution to glycan analysis that combines NIBRT’s novel bioinformatics database solution, GlycoBase 3.0+, with the unique capabilities of the Waters ACQUITY UPLC System and Glycan Separation Technology chemistries for HILIC-UPLC separations. GlycoBase 3.0+ is a web-enabled proprietary resource that contains normalized retention data (expressed as Glucose Units, or GU values) for more than 600 2-AB labeled N-linked glycan structures. These values were obtained by systematic analysis of released glycans from a diverse set of glycoproteins using Waters HPLC and UPLC technologies and the NIBRT glycan analytical platform.
Two orthogonal technologies, exoglycosidase sequencing and mass spectrometry, were used to confirm every structure, generating a high-quality glycan library that can be used as a general tool or interrogated in various ways, e.g., for each of the biotherapeutic glycoproteins analyzed. Data stored in a web-accessible database is accessed through a customized software application with a simple intuitive interface.
Each entry/glycan is comprehensively annotated and includes:
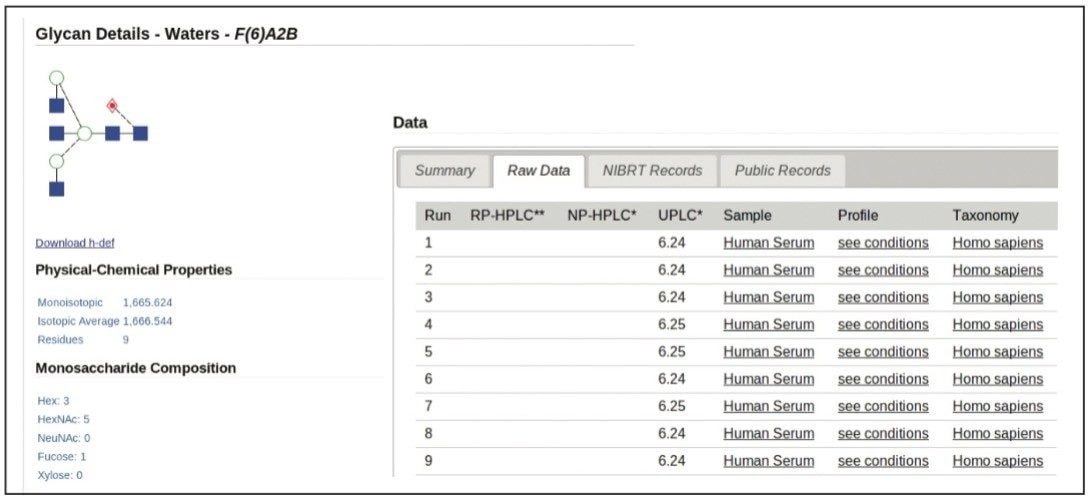

A glycan database, GlycoBase 3.0+, of protein N-linked oligosaccharides released from a range of biotherapeutic proteins has been developed for the biopharmaceutical industry using a combination of Waters and NIBRT technologies. Reproducible, sensitive, standardized and high-throughput glycan HILIC separations, in combination with an online glycan bioinformatics resource, GlycoBase 3.0+, allows the identification of glycans based on three orthogonal data attributes.
An additional feature of 2-AB fluorescently labeled N-linked glycans is that the data generated by chromatography can be used for relative quantitation, which when combined with Glycobase 3.0+ is a powerful tool for the biopharmaceutical industry. The dynamic range allows glycans present at <1% of the glycan pool to be quantified.
These novel capabilities will facilitate the more rigorous characterization of biopharmaceutical protein products required by today’s regulatory environment.
Waters would like to acknowledge Mark Hilliard, Weston Struwe, Giorgio Carta, John O’Rourke, Niaobh McLoughlin, and Pauline Rudd from NIBRT for this work.
720004203, December 2012