This is an Application Brief and does not contain a detailed Experimental section.
With the UNIFI Scientific Information System, we have taken these proven tools and integrated them seamlessly into user-defined workflows.
Building on the chemical intelligence pioneered in MetaboLynx Application Manager for MassLynx Software, newly developed software tools more intelligently engage the metabolite identification process to significantly reduce misidentifications and provide confidence in the automated results returned into your workflows.
We have previously discussed the benefits of the use of intelligent in-silico tools to support the metabolite identification process.1 These innovative software applications, which have been built through strong scientific partnerships, have facilitated the development and commercial deployment of intelligent workflow software throughout the industry. These partnerships have also allowed scientists to realize gains of an order of magnitude in throughput.2-4 With this next-generation solution in hand, Waters continues to drive the drug discovery process forward by building upon our core technology.
The next question for everyone is where do we go now?
With the UNIFI Scientific Information System, we have taken these proven tools and integrated them seamlessly into user-defined workflows. No longer is the output of an experiment predefined by strict rules; it is now built around tools that allow you to apply them as you see fit. You’ll not only produce results more rapidly, but also have more confidence and control in the results of your metabolite identification studies.
If you need to work on more than one drug at any given time, the task of determining the specific analysis conditions, processing settings, and reporting strategy holds back progress. UNIFI Software allows you to submit all of your compounds in a single, simple analysis list. Using a generic acquisition and automated structure-driven processing methodology, all of your samples are submitted, acquired, processed, and reported with no intervention from the user. All of this happens behind the scenes so your drug metabolism studies are not hindered by analytical tools.
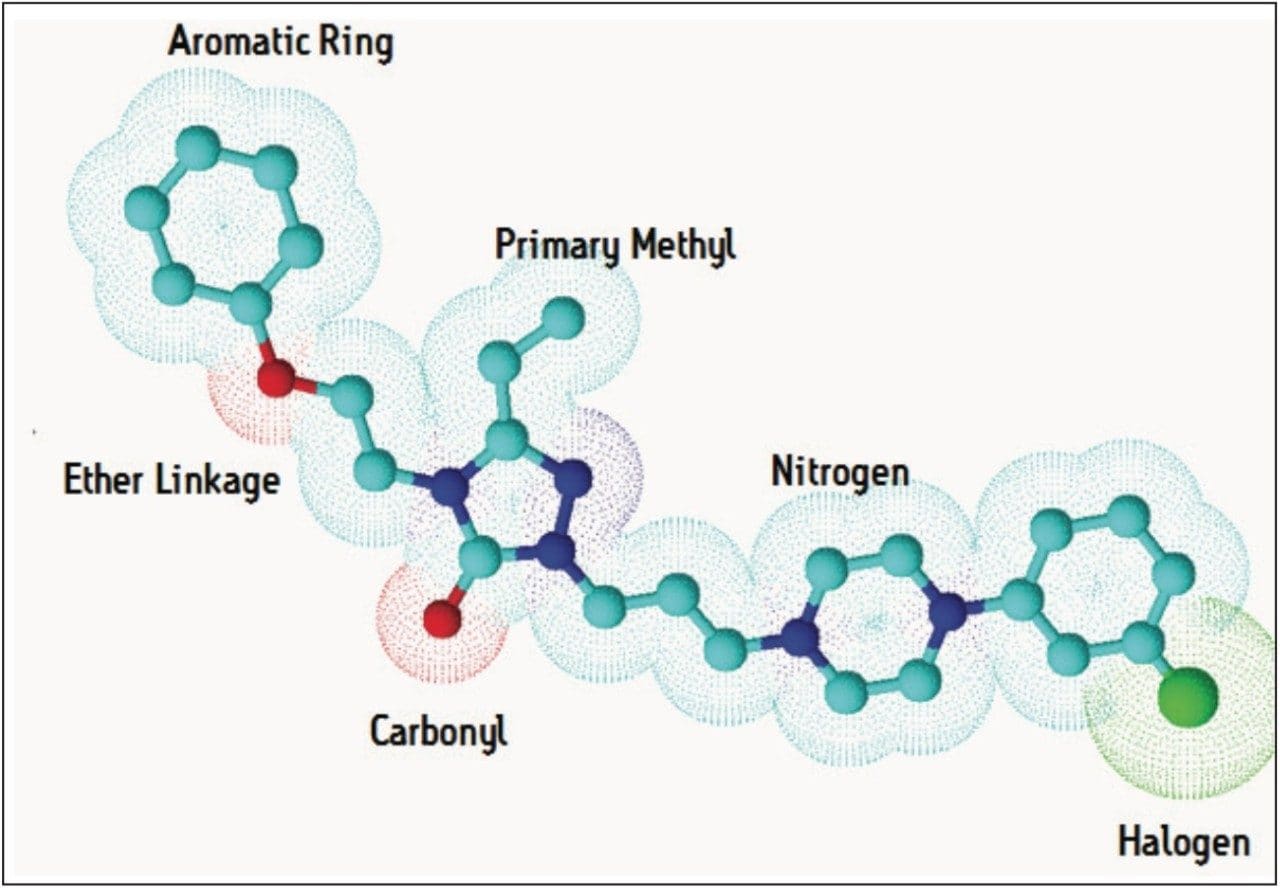
In order to fully understand a molecule you must understand its structure. Software algorithms interrogate each molecule submitted and fingerprint its metabolic potential through Functional Group Recognition (FGR). Now you don’t have to worry about manually selecting the correct standard biotransformations or mapping out potential secondary metabolites – the software will calculate this for you and only allow identifications to be made using the scaffold of what is actually in your drug ; an example is shown in Figure 1. FGR ensures that the answers returned are chemically valid and you don’t have to spend time reassigning incorrectly reported metabolic transitions.
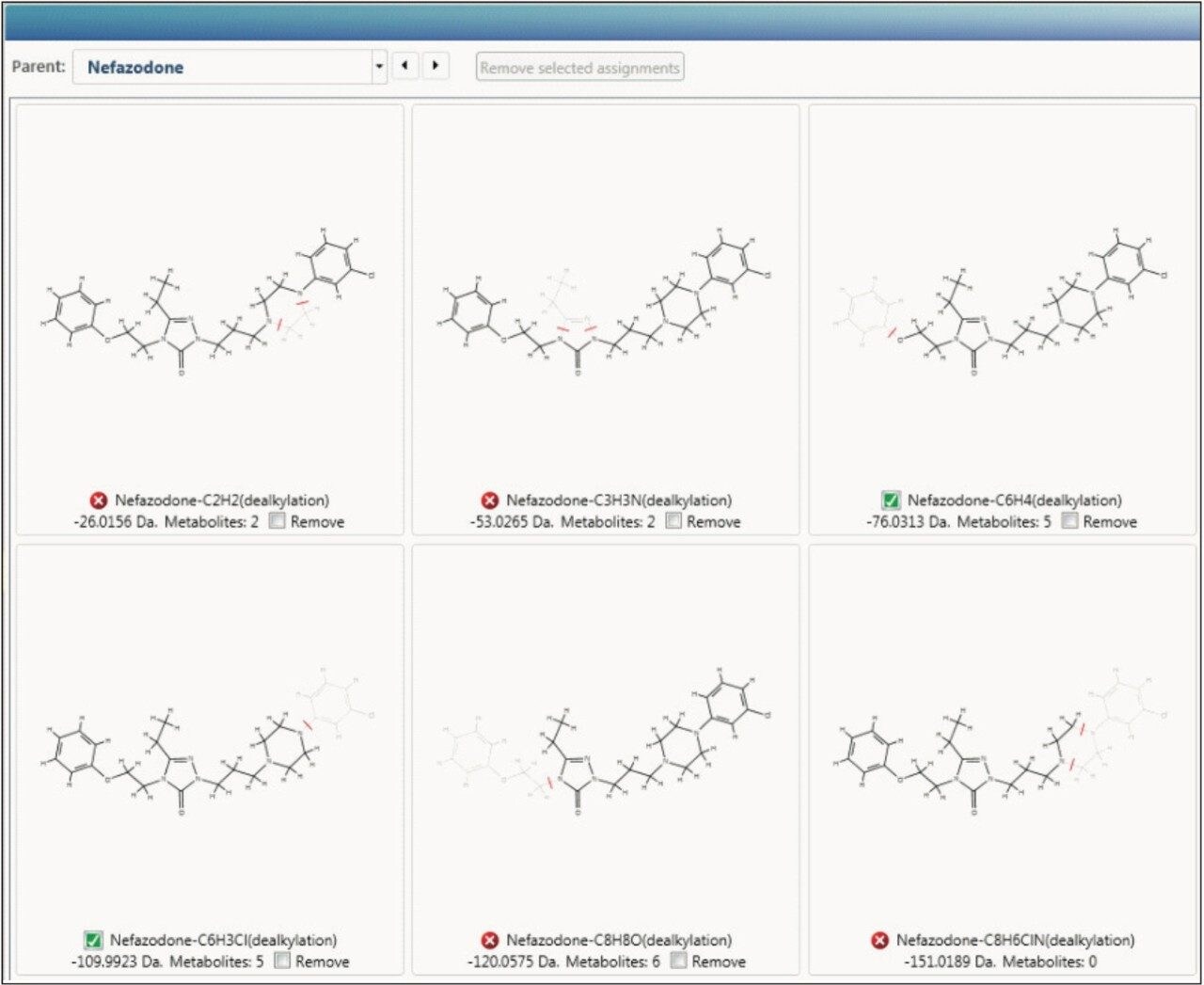
Understanding the structure of a molecule is only the first step in determining and characterizing the metabolic fate of a molecule. When you need go beyond this and think about the bond cleavage of your molecule either by enzymatic cleavage or collisionally induced dissociation, PrISE algorithms from Dotmatics, built based upon the principles published by Hill et al,5-6 are called to work.
The way the molecule is transformed biologically is predicted in-silico, both directly aiding identification of these metabolites and driving new cluster mass defect and intelligent elemental composition tools, as shown in Figures 2 and 3. The software is designed so that you can have greater confidence of the identification quality and coverage of the molecule’s metabolic space.
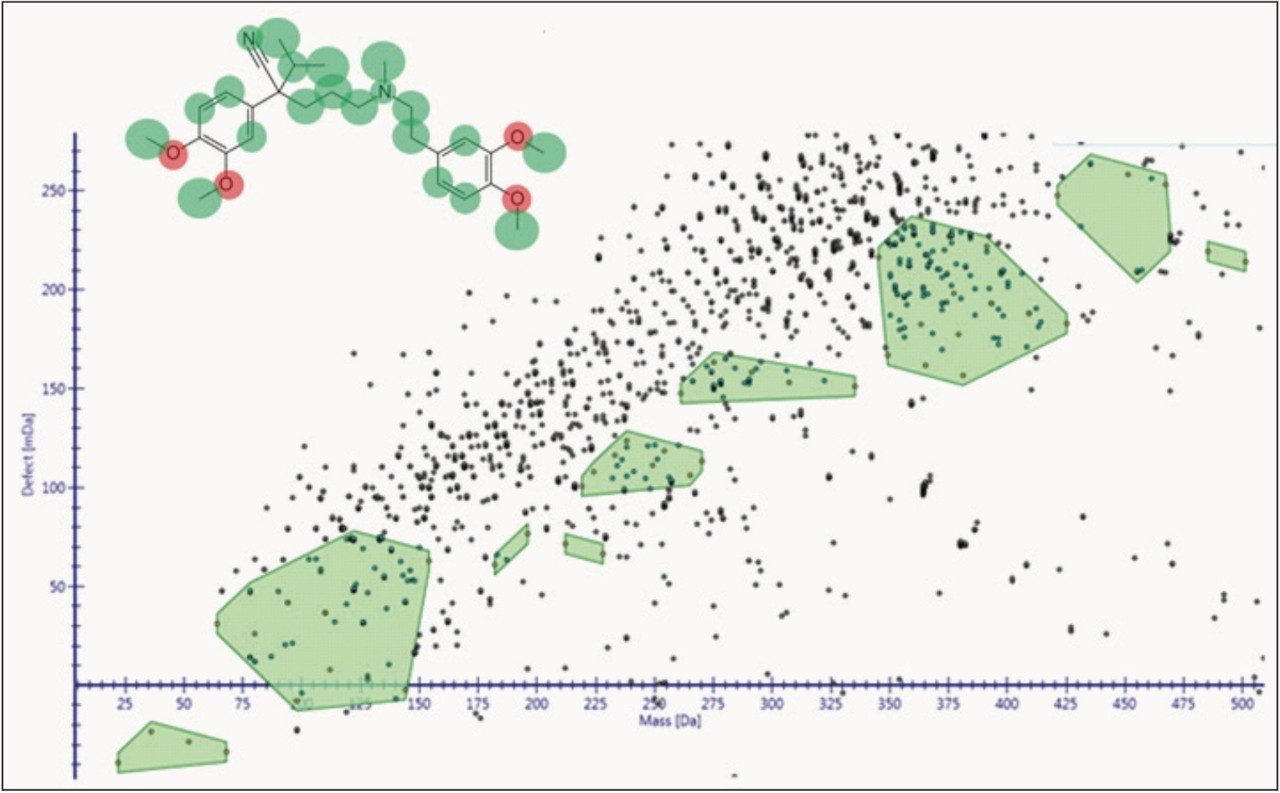
To aid in characterization, fragment spectra are automatically annotated with proposed structure, for both the parent drug and metabolites, giving you assurance that the identifications you are making are correct.
Taking this to the next level, UNIFI Software will also use this information to determine where the biotransformation is occurring, giving you instant access to decision-making information, as shown in Figure 4.
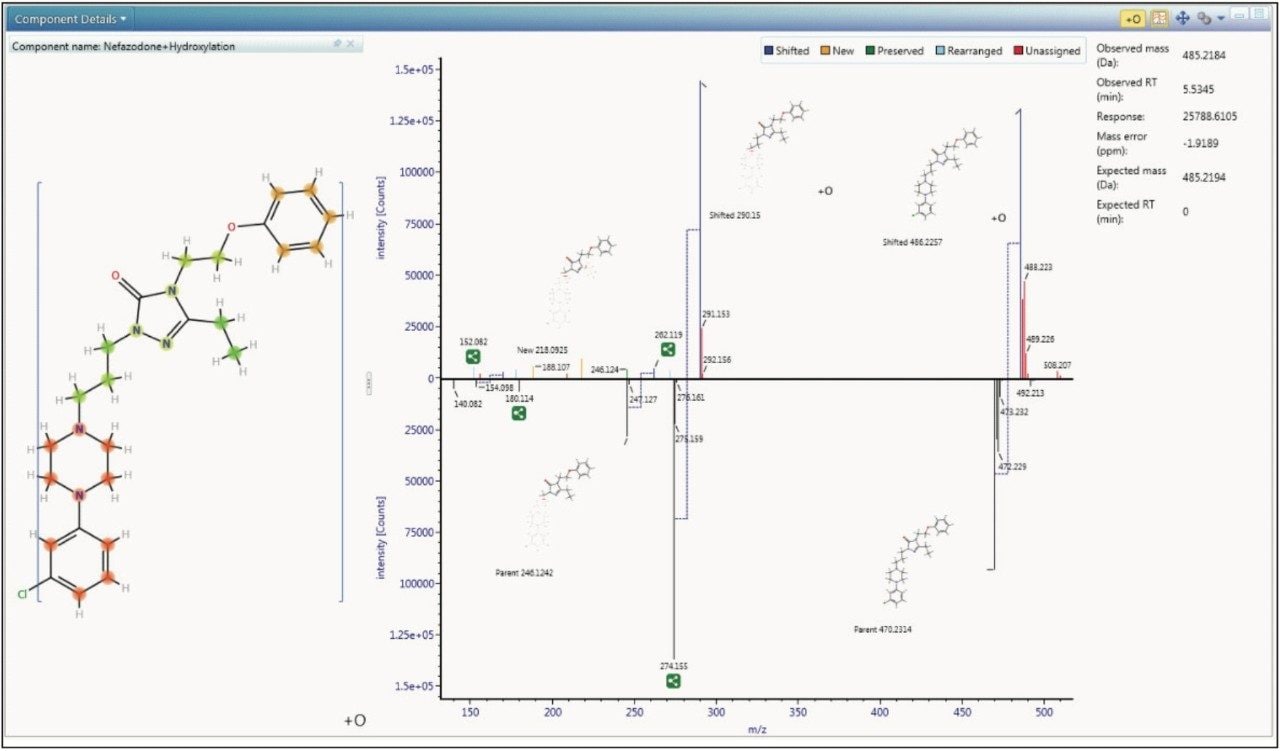
The described processing intelligence isn’t meaningful if data is not accessible and easily visualized. The Drug Related Properties view in UNIFI Software makes it easy to use all of the information from your data in a single window, showing you the criteria that provide evidence for the determination that a component is a metabolite. This view is not just for rapid identification of major metabolism, it can also be used in order to understand the complex relationships of the metabolites by identifying common fragments, neutral loss, and thus gaining a more definitive view of metabolic pathways.
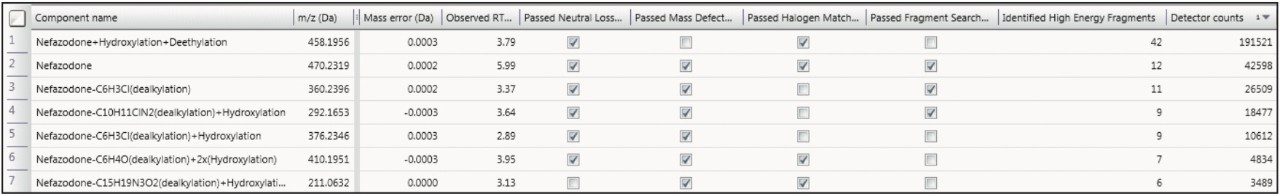
The data that you collect today is full of information. The key to success is in the development of tools that engage seamlessly with your LC-MS analysis to fully exploit this richness of content. Waters’ strength lies in our understanding of LC-MS data at the deepest level. This gives us the ability to pull even the smallest of relevant details from data and, more importantly, enables us to present this to you with confidence in the integrity of the answer. This allows you to focus on your strengths on using the data results to drive decisionmaking in the drug discovery and development process.
720004321, April 2012