In this study, an advanced LC-MSE peptide mapping workflow is demonstrated for the detection and identification of disulfide linkages, including scrambled disulfide linkages, in a recombinant IgG1 mAb. We develop a viable, integrated approach for automatic mapping or monitoring of disulfide bonded peptides in a protein with multiple cysteines.
Disulfide bond formation is critical for establishing and maintaining proper three-dimensional folding and therefore functions of therapeutic proteins, e.g., monoclonal antibodies (mAbs). Localization and assignment of disulfide bonds are an important aspect of protein structural analysis.
However, the identification of disulfide pairing in a protein with multiple cysteine residues is generally more time-consuming and challenging than the determination of the protein sequence due to incomplete disulfide bond formation and disulfide bond scrambling.1,2
The combination of enzymatic digestion with liquid chromatography-mass spectrometry (LC-MS) is frequently applied as a routine method for the assignment of disulfide bonds.3,4 However, peptide mapping via an LC-MS approach is traditionally labor-intensive and time-consuming in data processing and interpretation.
Furthermore, the assignment of disulfide linkages sometime is inconclusive when a protein contains multiple cysteines due to the significantly increased number of possible disulfide-bonded peptide isomers.1 For example, there are totally 105 possible disulfide-bond pairing schemes for a protein containing eight cysteine residues (while an IgG1 mAb typically has 32 or more cysteines in its two light chains and two heavy chains).
This, plus the reality of potential disulfide bond scrambling, makes the disulfide bond assignment extremely difficult when the analysis is performed manually. An automated workflow is highly preferred for quick assessment of heterogeneities of therapeutic proteins caused by the disulfide linkages.
Recently, we have developed an integrated peptide mapping workflow for protein sequence confirmation and characterization using a UPLC-MSE method for data collection and a targeted bioinformatic tool, the BiopharmaLynx Application Manager for MassLynx Software, for automated data processing and annotation.5
The data independent MSE approach alternatively collects mass spectrometric data of precursors and fragments of eluting peptides from protein enzymatic (e.g., tryptic) digests in an unbiased manner.6 The integrated workflow overcomes most shortcomings of traditional peptide mapping methods7 and has been demonstrated to be fast, robust and suitable for routine analysis of recombinant proteins such as therapeutic mAbs5 and subunit vaccines.8 The approach has also been successfully applied to identify expected disulfide bonds in an IgG1 mAb.9
In this study, an advanced UPLC-MSE peptide mapping workflow is demonstrated for the detection and identification of disulfide linkages (including scrambled disulfide linkages) in a recombinant IgG1 mAb. The goal is to develop a viable approach for automatic mapping or monitoring of disulfide bonded peptides in a protein with multiple cysteines.
The significances of the integrated workflow include:
To reduce the complexity of protein digest mixture, the mAb was digested with endoproteinase Lys-C. The obtained digests were separated by reversed-phase LC with an ACQUITY UPLC System, followed by online MSE detection and BiopharmaLynx 1.3 analysis. Digests prepared with and without alkylation protection of free sulfhydryl groups2 in the sample were used to differentiate native scrambled disulfide linkages in the mAb from those potentially formed during sample preparation stage.
|
LC system: |
ACQUITY UPLC |
|
Column: |
ACQUITY UPLC BEH300 C4, 1.7 μm, 2.1 x 150 mm (p/n 186004497) or ACQUITY UPLC BEH300 C18, 1.7 μm, 2.1 x 150 mm (p/n 186003687) |
|
Column temp.: |
65 °C |
|
Flow rate: |
200 μL/min |
|
Injection vol.: |
5 μL (partial loop injection mode) |
|
Mobile phase A: |
0.1% FA in water |
|
Mobile phase B: |
0.1% FA in ACN |
|
Gradient: |
1–40% in 90 min |
|
MS system: |
SYNAPT HDMS G2 |
|
Ionization mode: |
ESI+ |
|
Acquisition mode: |
MSE |
|
Low collision energy: |
4 eV |
|
Elevated collision energy: |
Ramping from 22 to 48 eV |
|
Capillary voltage: |
3.0 kV |
|
Cone voltage: |
30 V |
|
Scan mode: |
Resolution mode (resolution ≥ 20000) |
|
Scan time: |
0.5 second |
|
Source temp.: |
100 °C |
|
Desolvation temp.: |
350 °C |
|
Mass range: |
100 to 2500 Da |
|
Mass accuracy: |
Lockmassed by GFP sprayed from lockmass channel every 30 seconds |
BiopharmaLynx 1.3 Application Manager for MassLynx Software
Endoproteinase Lys-C (Wako) and iodoacetamide (Sigma) were used along with RapiGest (Waters, p/n 186001861). The IgG1 mAb Trastuzumab digests were prepared by incubating the sample (protein/Lys-C of 50:1) in 50 mM tris buffer (pH ~7.5) at 37 °C and the presence of 0.1% RapiGest SF for 18 hours. Prior to digestion, the protein was denatured at 80 °C for 10 min. For testing artificial scrambling potentially introduced during digestion, a digest prepared without alkylation was directly compared with a digest prepared in parallel with IAM alkylation before the digestion.
Like all IgG1 mAbs, the Trastuzumab antibody should have nine unique disulfide bonds when formed properly. In-silico digestion (by Lys-Cp) shows that eight expected disulfide linked peptides (or unique linkages) are generated after digestion, including two in each of light chain, four in each of heavy chain, one between light chain and heavy chain, and one between the two heavy chains (which has two disulfide bonds connecting with).
To demonstrate that the workflow works properly to identify the formed disulfide linkages, the collected LC-MSE data were first searched against the mAb sequences with properly linked cysteines to form the 16 expected disulfide bonds in the BiopharmaLynx method setting.
Figure 1 shows the determination of the largest disulfide linkages 1:K1–1:K4 (an intra disulfide linkage of light chain, monoisotopic mass 10657.12 Da). The disulfide-bonded peptide was detected and annotated based on the precursor mass MH+ (Figure 1C) that was obtained from multiply-charged, isotope resolved raw MS data (Figure 1B). The assignment was also confirmed by elevated-energy fragmentation MSE spectrum (also charge-reduced, isotope-deconvoluted to singly-charged ions, as shown in Figure 1D) upon the retention time-alignment (as highlighted in Figure 1A). The MSE spectrum not only contains b/y ions from the two individual peptides (1:K1 and 1:K4), but also has ions corresponding to disulfide-bonded fragments from both peptides such as 7019.27 Da (1/y39–2/y24) and 10431.19 Da (1/y39–2/y56), providing unequivocal evidences for the disulfide linkage.
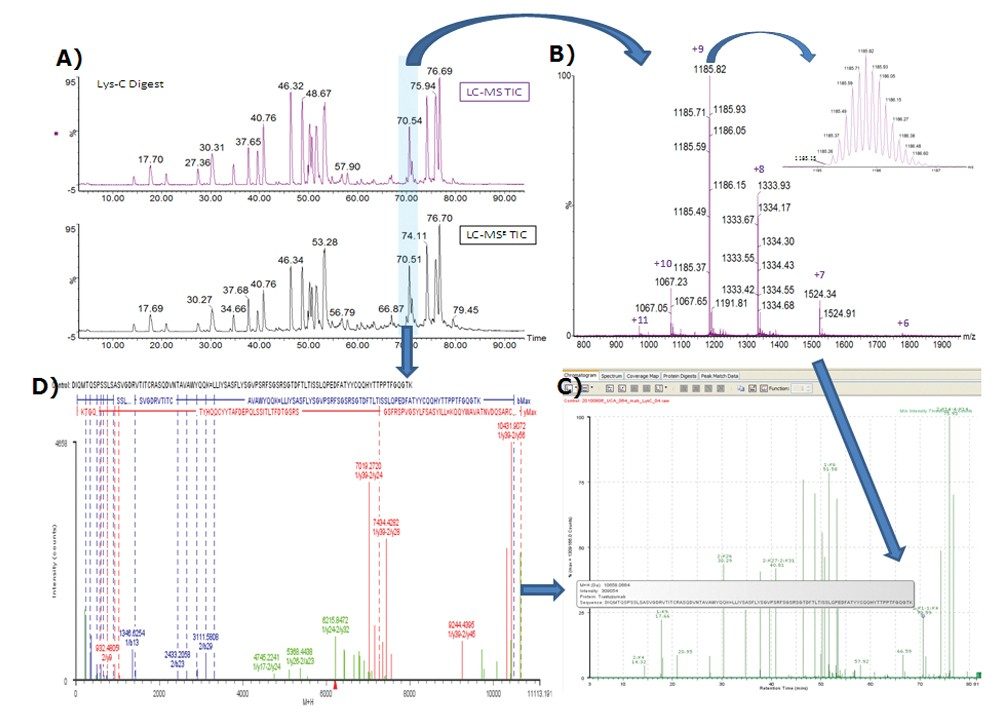
It should be stressed that the improved MS instrument resolution (see the inset in Figure 1B) is important to obtain the correct monoisotopic masses for the precursors as well as fragments of the large peptides. A well-resolved isotope pattern for such large peptides greatly facilitates the informatics tool for the identification of large disulfide linkages such as 1:K1–1:K4.
Similarly, BiopharmaLynx identifies other expected disulfide linkages. The mass accuracy of all identified disulfide bonded peptides was within 5.0 ppm. In addition, BiopharmaLynx is also capable of identifying the disulfide-bonded peptides with one of the linked peptides containing miscleavages.
For example, both disulfide linkages 2:K7–2:K8 (without miscleavage) and 2:K7–2:K8-9 (with one miscleavage) were identified (see corresponding peaks B and C in Figure 2A). Their MSE spectra are plotted in Figures 2B and 2C, respectively, showing confident identification. This capability is especially important because proteases such as Lys-C or trypsin tend to miss the cleavage site when a lysine (K) is followed by a proline (P) residue.
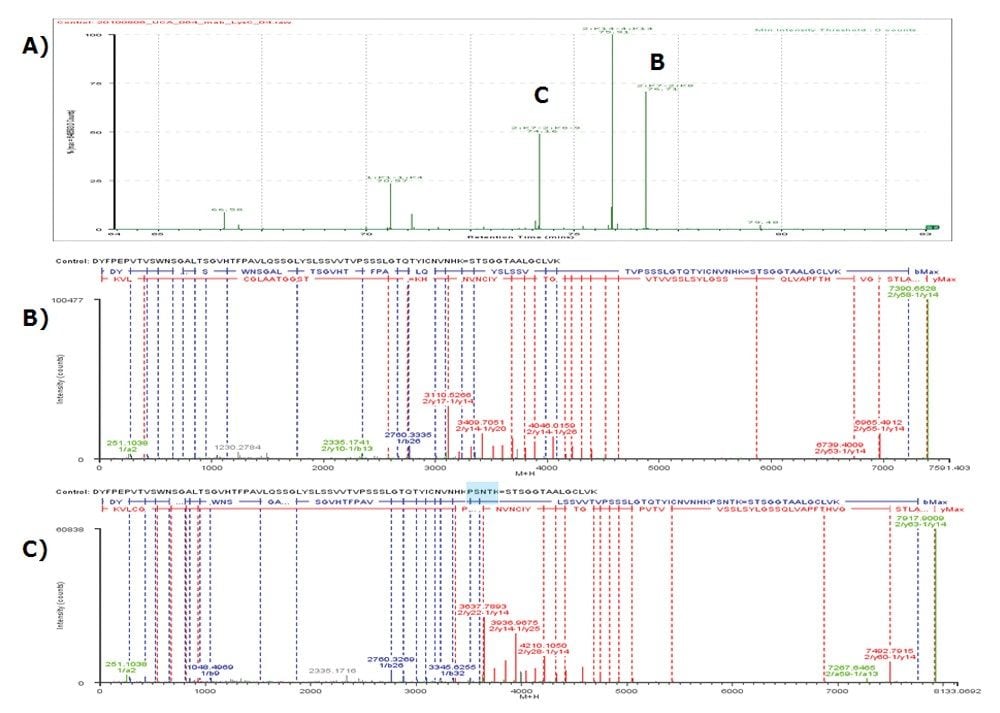
It is observed that high percentage of organic solvent (10% or more acetonitrile) is needed in the sample to help maintain the solubility of large disulfide-bonded peptides during LC-MS analysis. One disadvantage of the use of high organic in sample buffer is that this could result in the elution of the smallest disulfide-bonded peptides 1:K14–2:K13 (SFNRGEC=SCDK) in void volume.
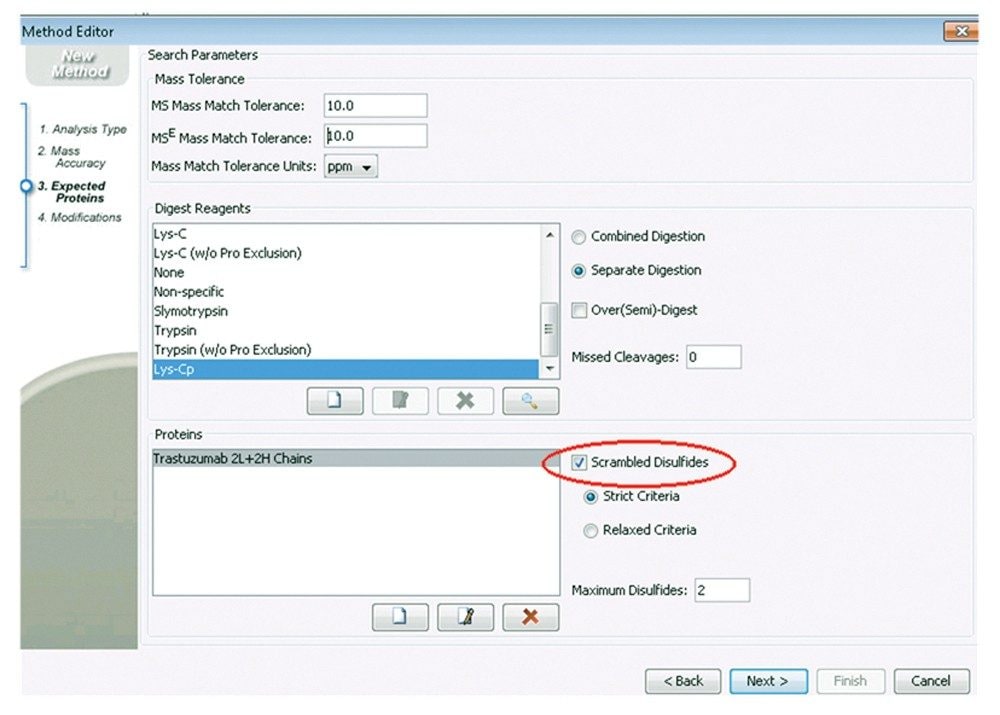
One challenge in disulfide bond mapping is to identify those peptides formed unexpectedly, e.g., due to scrambling. Therefore, the processing method was next created for Scrambled Disulfides (see the highlight part in Figure 3), which allows for detection and identification of all possible disulfide pairing formed between any two cysteines in the mAb. Lys-C digests of Trastuzumab with or without alkylation prior to digestion were analyzed for this investigation.
Using this new method, not only the expected disulfide linkages were identified, some minor scrambled disulfide linkages were also detected. Figure 4A illustrates three identified scrambled disulfide linkages (1:K13–2:K16, 1:K7–2:K21, 1:K14–2:K16) in the digest without alkylation. Again, elevated-energy fragmentation (MSE) data were available for confirmation of the assignments (the MSE spectrum of 1:K13–2:K16 is plotted in Figure 4B as an example).
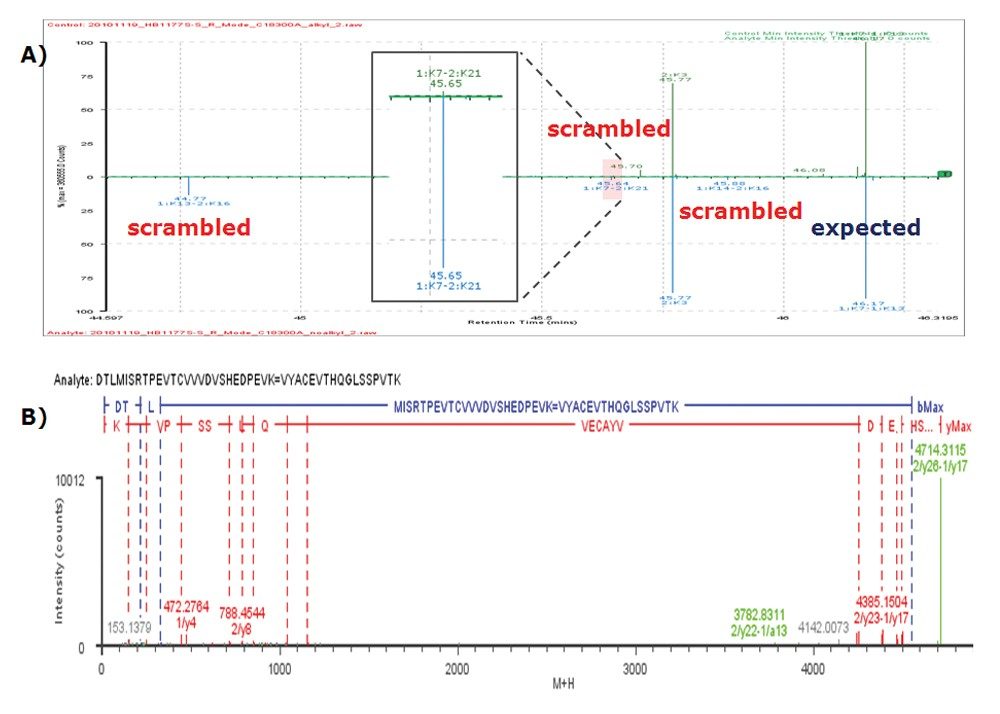
The scrambled 1:K13–2:K16 and 1:K14–2:K16 were only identified from the digest without alkylation and not detected in the digest with alkylation, suggesting these were artifacts introduced by sample preparation. The scrambled disulfide linkage 1:K7–2:K21 was identified in both digests (with or without alkylation), however, its abundance is much lower in the alkylated sample compared to the non-alkylated one (see the inset in Figure 4A). As expected, all the identified scrambled disulfide linkages are minor compared to the expected disulfide linkages (as illustrated in Figure 4A).
The successful determination of both expected and scrambled disulfide linkages demonstrated that the integrated workflow is capable of automatically mapping disulfide linkages in a protein. The enhanced function of Scrambled Disulfides requires no manual efforts, and achieves completely automatic data processing for the challenging task of disulfide bond mapping.
An integrated workflow, combining high mass resolution UPLC/MSE with BiopharmaLynx 1.3, was developed for rapid mapping of disulfide linkages in a recombinant IgG1 mAb. The workflow was demonstrated to be capable of simultaneous identification of both expected and scrambled disulfide linked peptides. Assignment of disulfide bond linked peptides is automated, based on accurate MS measurement and confirmed by elevated collision energy MSE fragmentation data.
The improved mass resolution – 20,000 for resolution mode and 40,000 for high-resolution mode – offered by the SYNAPT G2 HDMS System greatly enhances the MS data quality for large disulfide-linked peptides in enzymatic (e.g., Lys-C) digests. The added Scrambled Disulfides function setting of BiopharmaLynx 1.3 makes it possible for identification of any potential disulfide linkages of a protein in an automated mode.
This integrated approach should be applicable for routine mapping and monitoring of disulfide linkages in mAbs and other therapeutic proteins with multiple disulfide linkages.
720003939, June 2011