This work describes a selective sample preparation strategy and LC-MS compatible sample storage plates with high performance surfaces to mitigate pramlintide loss due to non-specific binding. In addition, it shows that optimized UPLC separation and column chemistry, along with a highly sensitive tandem quadrupole mass spectrometry method, can produce significantly lower limits of quantification.
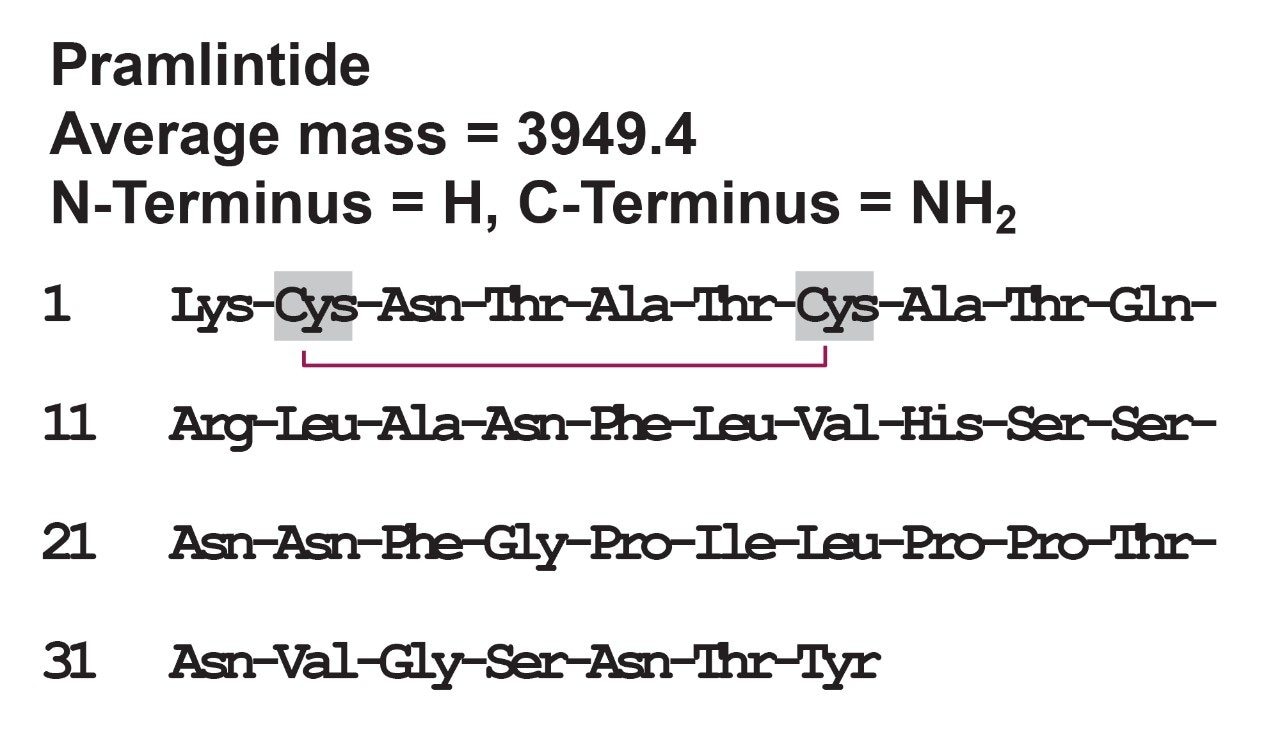
Pramlintide acetate (SYMLIN) is a 37 amino acid (MW 3949.4 Da) synthetic analogue of the human hormone amylin (Figure 1). Developed as an adjunctive therapy for patients with Type 1 and 2 diabetes, this peptide can improve treatment outcomes for those who have failed to achieve glycemic control despite optimal insulin therapy.1 With nearing patent expiry dates for pramlintide,2 and recent research indicating a role for amylin in Alzheimer’s Disease models,3 interest in amylin and amylin agonists is rising. Characterized by fast absorption (<30 minutes), elimination (~1 hour), and low circulating levels (pg/mL), amylin agonists such as pramlintide can be challenging to quantify. LC-MS/MS assays have become increasingly popular for peptide quantification due to the high sensitivity and specificity afforded by selective MRM fragments. However, method development and accurate quantification for hydrophobic peptides like pramlintide can still be challenging because peptides notoriously suffer from non-specific binding (adsorption). This can lead to poor recovery, loss of analyte, and poor limits of detection. The work described here uses a selective sample preparation strategy and LC-MS compatible sample storage plates with high performance surfaces to mitigate pramlintide loss due to non-specific binding. In addition to this, we will show that optimized UPLC separation and column chemistry, along with a highly sensitive tandem quadrupole mass spectrometry method can produce significantly lower limits of quantification. For this assay, an LLOQ of 25 pg/mL of pramlintide was achieved, extracted from 100 μL of rat and human serum.

Calibration curve standards and quality control (QC) samples of pramlintide (ProSpec Bio, Rehovot, Israel, p/n: HOR-300) were prepared in commercially available human and rat serum at various concentration levels (25–50,000 pg/mL). All calibration curve standards, QC levels, and blank (non-spiked) samples were prepared in triplicate.
One hundred microliters of serum was diluted with 100 μL of water and vortexed. All wells of an Oasis WCX 96-well μElution Plate (p/n: 186002499) were conditioned with 200 μL of methanol and then equilibrated with 200 μL of water. The diluted serum samples (200 μL) were loaded onto the SPE plate, subsequently washed with 200 μL of water, and followed by 200 μL of 20% acetonitrile in water. Pramlintide was eluted from the sorbent using a 1 × 25 μL aliquot of the elution solvent containing 1% trifluoroacetic acid in 75:25 (v/v) acetonitrile:water. Eluates were collected in a QuanRecovery LC-MS compatible sample plate with MaxPeak High Performance Surfaces (p/n: 186009184), and then diluted with 25 μL of water for a final sample volume of 50 μL. Ten microliters of each sample were injected onto an ACQUITY UPLC I-Class PLUS System equipped with an ACQUITY UPLC Peptide CSH C18, 2.1 × 50 mm Column (p/n: 186006936) and a Xevo TQ-XS Mass Spectrometer.
|
LC system: |
ACQUITY UPLC I-Class PLUS (fixed loop) |
|
Detection: |
Xevo TQ-XS Mass Spectrometer, ESI+ |
|
Column: |
ACQUITY UPLC Peptide CSH C18, 130 Å, 1.7 μm, 2.1 × 50 mm (p/n: 186006936) |
|
Column temp.: |
60 °C |
|
Sample temp.: |
15 °C |
|
Injection volume: |
10 μL |
|
Mobile phase A: |
0.1% Formic acid in water |
|
Mobile phase B: |
0.1% Formic acid in acetonitrile |
|
MS system: |
Xevo TQ-XS |
|
Ionization mode: |
ESI+ |
|
Capillary: |
1.0 kV |
|
Cone: |
15 V |
|
Source offset: |
30 V |
|
Source temp.: |
150 °C |
|
Desolvation temp.: |
600 °C |
|
Cone gas flow: |
150 L/hr |
|
Desolvation gas flow: |
1000 L/hr |
|
Collision gas flow: |
0.15 mL/min |
|
Nebulizer gas flow: |
7 bar |
|
System calibration: |
Low resolution |
|
Data management: |
MassLynx (v4.2) |
|
Quantification software: |
TargetLynx XS |
|
Time(min) |
Flow rate (mL/min) |
%A |
%B |
Curve |
|---|---|---|---|---|
|
Initial |
0.400 |
80.0 |
20.0 |
6 |
|
0.50 |
0.400 |
80.0 |
20.0 |
6 |
|
1.00 |
0.400 |
78.0 |
22.0 |
6 |
|
4.00 |
0.400 |
73.0 |
27.0 |
6 |
|
4.75 |
0.400 |
5.0 |
95.0 |
6 |
|
5.50 |
0.400 |
5.0 |
95.0 |
6 |
|
6.00 |
0.400 |
80.0 |
20.0 |
6 |
|
7.00 |
0.400 |
80.0 |
20.0 |
6 |
Larger and more hydrophobic molecules, including peptides, often suffer from non-specific binding (NSB) or adsorption to any labware that samples come in contact with (e.g., plates, vials, pipette tips).4 A commonly used strategy to mitigate the effects of non-specific binding of the molecules of interest is to add carrier protein(s) to the sample. This can be done prior to sample cleanup and/or immediately prior to LC-MS/MS analysis of the analyte. Although generally highly effective, adding carrier proteins to a sample also adds more complexity to the sample, which sample preparation seeks to eliminate. In recent years, scientists have found more innovative ways to mitigate non-specific binding. Waters developed QuanRecovery LC-MS sample plates and vials with MaxPeak High Performance Surfaces which mitigate, and in some cases eliminate, non-specific binding of peptides to the surface of the container.
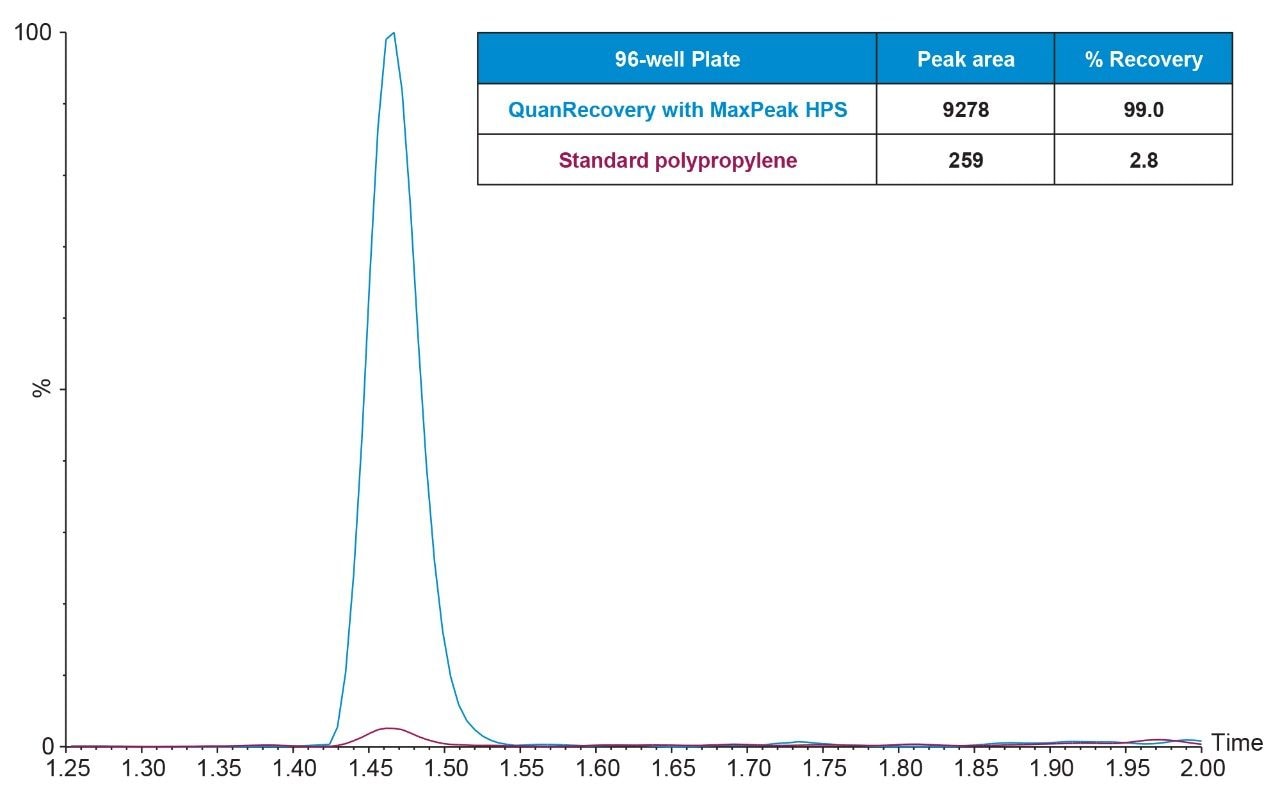
Pramlintide is a large peptide (MW 3949.4 Da) which is hydrophobic (HPLC Index: 88.7) and can therefore be expected to suffer from some degree of non-specific binding. During method development, pramlintide was stored in either a QuanRecovery sample plate or a standard polypropylene plate prior to LC-MS/MS analysis in order to assess the plates’ effectiveness in mitigating NSB. Test samples were prepared and stored in neat solution, while additional samples prepared in a solution containing carrier proteins were used as a benchmark for 100% recovery. Results from this assessment showed that pramlintide suffered from a high degree of NSB to a standard polypropylene plate, while the QuanRecovery Plate was able to significantly mitigate the effects of NSB as seen in Figure 2. Recovery of 10 ng/mL pramlintide from the QuanRecovery Plate was almost 100%, leading to a peak area ~36 times higher than samples stored in a standard polypropylene plate where analyte recovery was only ~3%. Although this strategy combats pre-analytical NSB of our peptide of interest, it is important to keep in mind that peptides can adsorb to any surface they come in contact with prior to entering the mass spectrometer (including any plate transfer steps or system loss). For all analytes, it is important to determine if pre-analytical mitigation of non-specific binding is enough to reach the desired limits of detection of your assay. In some cases, it may still be necessary to use both carrier protein addition and high-performance surface sample storage containers to ensure that your analyte is not lost due to adsorption. Pramlintide benefits from the use of QuanRecovery Plates and Vials post SPE extraction to reach the desired limits of detection of this assay.
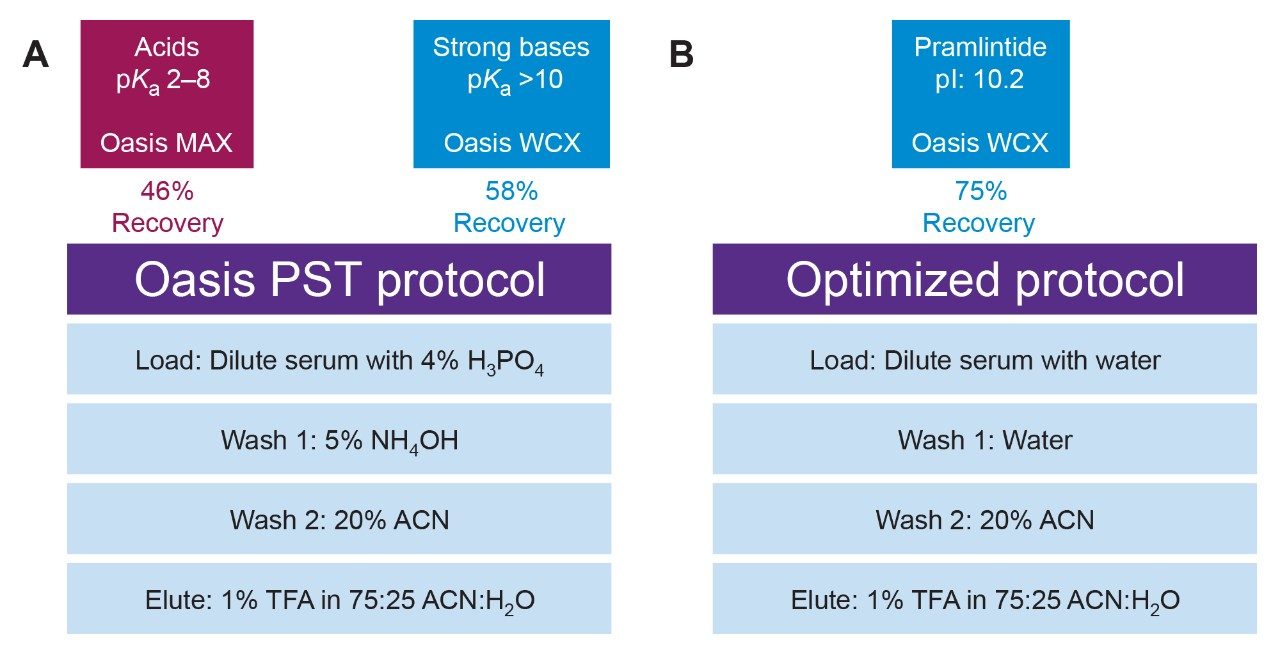
Solid-phase extraction (SPE) is a simple, robust, and fast sample preparation method which can be used to extract analytes from complex matrices. The Oasis Peptide Separation Technology (PST) μElution SPE screening strategy (Figure 3, Panel A), specifically designed for peptides, provides the user with simple methods to quickly screen multiple SPE sorbents (Oasis MAX and WCX) to assess recovery and matrix effects of the peptide of interest. Using this strategy, pramlintide was screened and it resulted in starting recoveries of ~46 and 58%, respectively. The calculated isoelectric point (pI) of pramlintide is 10.2 indicating that this peptide will be positively charged under most SPE conditions. Due to this, and the higher recovery achieved in the initial screening experiment, the Weak Cation-eXchange sorbent, Oasis WCX, was best suited for the extraction of pramlintide.
Through additional optimization experiments, it was determined that pramlintide was not adequately retained on the WCX sorbent during sample loading and Wash 1 with 5% NH4OH (pH ~12). Pramlintide is a basic peptide, as indicated by its pI, and is largely negatively charged at this pH, leading to poor retention on the negatively charged SPE sorbent. Changing the pretreatment solution and Wash 1 solvent to water (pH <7) ensured that the sorbent remained negatively charged, and pramlintide remained positively charged during both loading and Wash 1. Wash 2 and the elution solution used in the PST protocol remain the same in the optimized protocol, as it was determined during method optimization that trifluoroacetic acid was effective at fully eluting pramlintide from the SPE sorbent. Weaker acids such as formic acid and acetic acid were assessed and resulted in very low recoveries of the analyte (<10%). With the combination of this optimized protocol and chromatography gradient (discussed below), recovery of pramlintide was improved to ~75% (Figure 3, Panel B).
Optimization of chromatography was essential to the success of this assay. During method development of pramlintide, several reversed-phase columns were assessed for overall chromatographic performance (not shown). It was determined that the ACQUITY UPLC Peptide CSH C18 Column, with its positively charged surface, provided the best selectivity and sensitivity, with a 5-fold improvement in signal to noise compared to the ACQUITY UPLC Peptide BEH C18 Column.
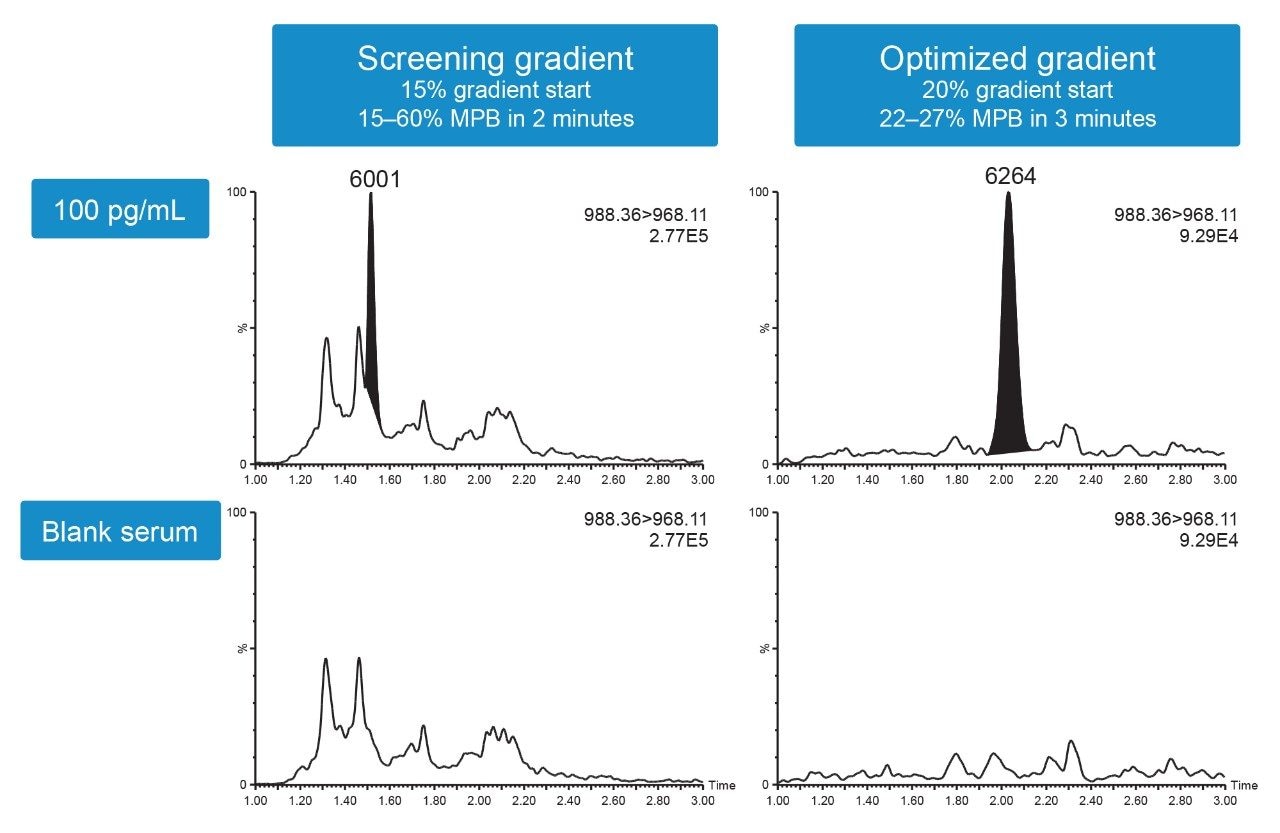
Despite the improved selectivity and sensitivity afforded by the ACQUITY UPLC Peptide CSH C18 Column, matrix interferences from extracted serum samples were still high, leading to signal suppression. During method development, a screening gradient from 15–60% mobile phase B (MPB) over two minutes was used. Initial results determined there was significant signal suppression (~20%) as compared to a non-extracted pramlintide sample. Modifying the gradient and increasing the starting percentage of organic solvent MPB from 15–20% significantly decreased the matrix background entering the mass spectrometer. In order to properly separate pramlintide from remaining matrix interferences at very low concentrations, the gradient was shallowed to 22–27% MPB over three minutes.
The combination of selective column chemistry and a highly optimized gradient resulted in reduction of observed matrix suppression (<10%) and improved pramlintide peak area, greatly improving both selectivity and sensitivity (Figure 4).
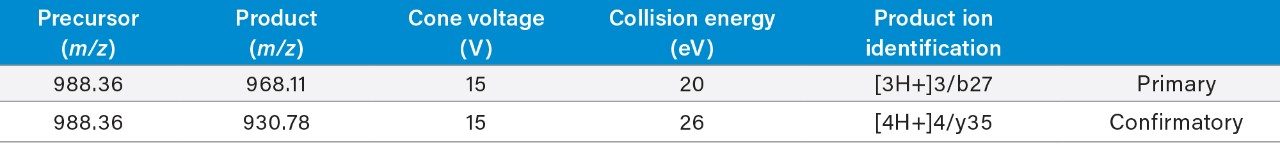
LC-MS/MS quantification of pramlintide was performed using a Xevo TQ-XS Mass Spectrometer and low resolution calibration (1.0 Da at FWHM) to achieve better assay sensitivity. When working with a lower resolution system calibration, it is critically important to use selective MRM fragments with high m/z values for quantification in order to avoid high background and matrix interferences. During method development, three predominant precursors were observed at 1317.48 (3+), 988.36 (4+), and 790.89 (5+) m/z, with the 4+ precursor resulting in the most intense fragments. Via manual tuning, an intense fragment of the 4+ precursor was identified at 968.11 m/z, corresponding to the b-ion cleavage between Leu-27 and Pro-28. This highly selective and intense fragment of pramlintide was used as the primary quantification transition, while the y-ion transition at 930.78 m/z was used as a qualifier. Optimized MS conditions and MRM transitions used for quantification of pramlintide are listed in Table 1.
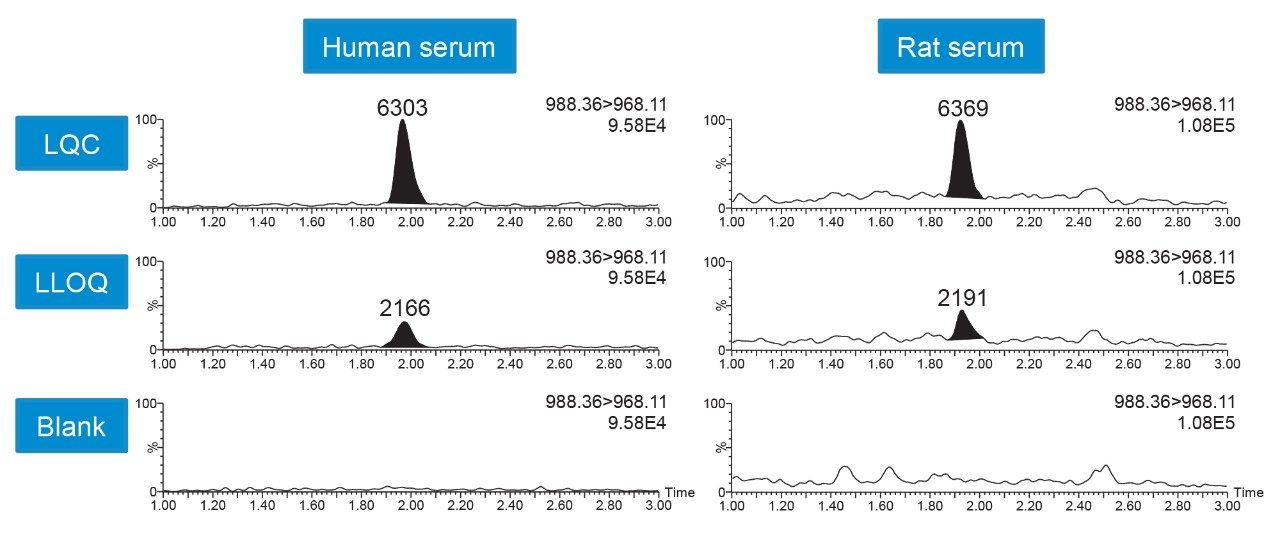
Linear, precise, and accurate quantification of pramlintide was achieved using only 100 μL of human or rat serum. Limits of detection (LOD) of 10 pg/mL and lower limits of quantification (LLOQs) of 25 pg/mL were achieved for both human and rat serum. Calibration curves were linear (r2>0.99) from 25–50,000 pg/mL using a 1/X2 linear fit (Table 2). All calibration curves and QC levels were accurate within ±15% and CVs were <5%. QC performance is highlighted in Table 3 where all statistics met recommended bioanalytical method validation guidelines, and chromatographic performance is illustrated in Figure 5.
The work described here employs a simple sample preparation strategy using Oasis WCX μElution SPE and QuanRecovery Sample Plates with MaxPeak High Performance Surfaces. Combining this approach with UPLC separation and a tandem quadrupole MS resulted in high sensitivity quantification of pramlintide from human and rat serum.
720006527, March 2019