For research use only. Not for use in diagnostic procedures.
This application note describes a hydrophilic interaction chromatography (HILIC) based approach for the separation of lipids by class prior to MS analysis and thereby reducing identification ambiguity. It describes the use of the LipidQuan platform that utilizes a HILIC-based approach to perform a targeted screen for Cer, HexCer, and SPH.
Sphingolipids (SLs) are a class of lipid comprising ceramides (Cer), sphingomyelins (SMs), and more complex glycosphingolipids.
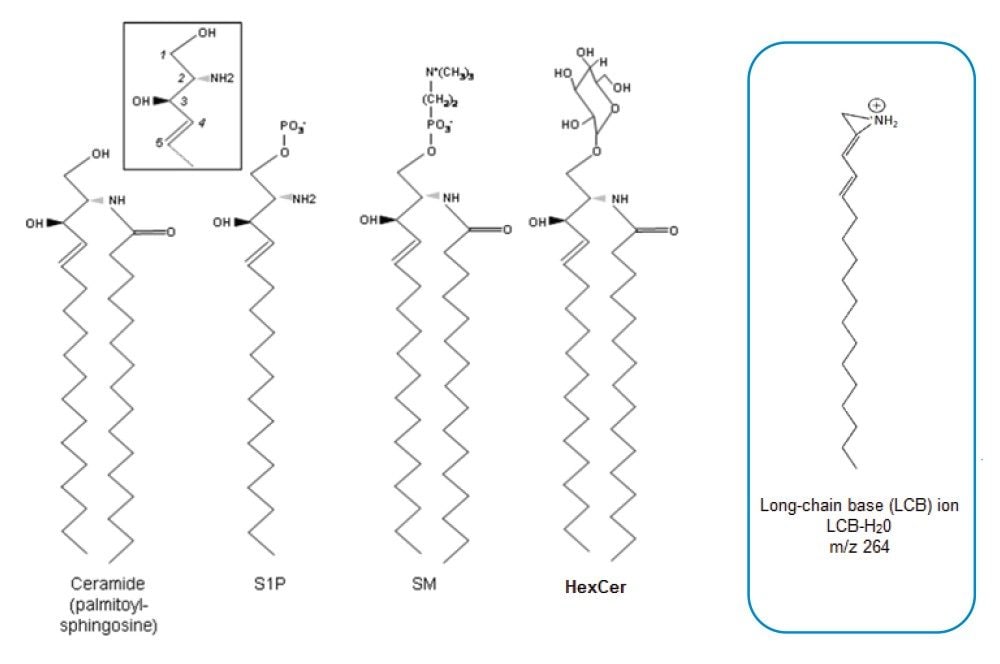
A common constituent of all SLs is a “long chain” or “sphingoid” base.1 These bases are organic aliphatic amino alcohols such as sphingosine (SPH) or a structurally similar compound (Figure 1). They are essential components of cellular membranes and are implicated in a variety of biological functions including cellular signal transduction and apoptosis.2,3 SLs play key roles in the induction of tumour cell proliferation and conference of chemotherapeutic resistance,4 while Cer and hexosylceramide (HexCer) have been found to be at elevated levels in Parkinson’s disease patients.5 The rapid metabolic interconversion of SLs as well as the high number of derivatives with similar functions creates a challenge for characterizing functional profiles of individual sphingolipids.
Although advances in mass spectrometry (MS) have allowed for more in depth lipidomic analysis, unambiguous identification and quantification have proven difficult. Lipids exhibit a high number of isomeric and isobaric lipid species. Furthermore, MS spectra often contain peaks and fragments from multiple compounds, making clear identification and relative quantification of specific molecular species difficult and time consuming. As a result the transfer of lipidomic data between laboratories is severely hindered, making it difficult to draw biological interpretation using multiple-site studies.
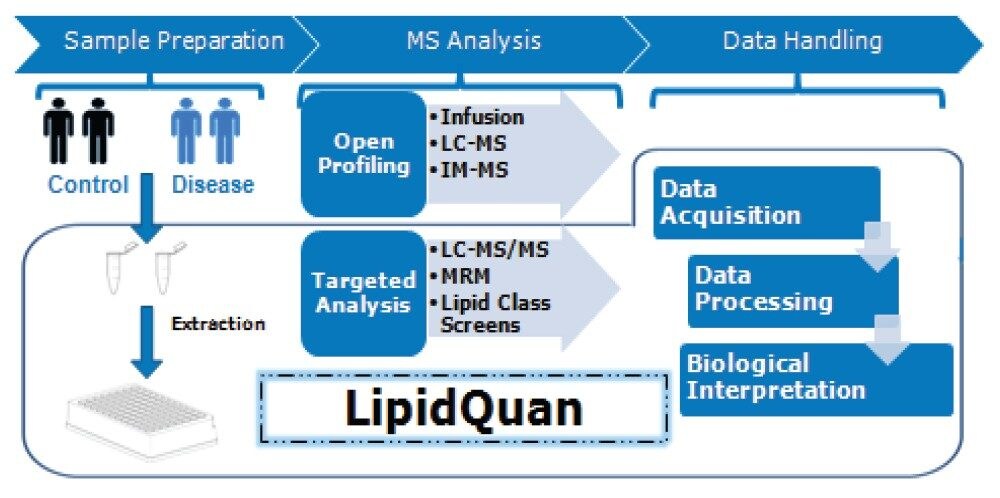
Here, we describe a hydrophilic interaction chromatography (HILIC) based approach for the separation of lipids by class prior to MS analysis and thereby reducing identification ambiguity.6 Separating lipid species by class also results in fewer stable isotope labelled (SIL) standards being required for quantification, thereby saving cost. This application note describes the use of the LipidQuan platform (Figure 2) that utilizes a HILIC-based approach to perform a targeted screen for Cer, HexCer, and SPH.
Pooled healthy human plasma was spiked with stable isotope labelled (SIL) SLs, (Deuterated CERAMIDE LIPIDOMIX, Avanti Lipids, Alabaster, AL) at nine concentration levels to generate calibration curves for quantification. The concentration ranges covered were; C16 Ceramide-d7 (d18:1-d7/16:0) = 2.2-1090 ng/mL, C18 Ceramide-d7 (d18:1-d7/18:0)= 1.2-575 ng/mL , C24 Ceramide-d7 (d18:1-d7/24:0) =2.6-1315 ng/mL and C24:1 Ceramide-d7 (d18:1-d7/24:1) = 1.3-665 ng/mL are used to illustrate this.
Six replicates of the NIST Standard Reference Material 1950 plasma (Sigma Aldrich, Poole, UK) were also spiked with 5% SIL, prior to extraction.
A simple sample preparation procedure was adopted using protein precipitation with pre-cooled isopropanol (IPA) (1:5, plasma:IPA). Samples were vortex mixed for 1 minute and placed at -20 °C for 10 minutes. Samples were vortex mixed again for one minute and placed at 4 °C for two hours to ensure complete protein precipitation. The extracted samples were centrifuged at a maximum of 10,300 g for 10 minutes at 4 °C before transferring the supernatant to glass vials for LC-MS/MS analysis.
|
LC system: |
ACQUITY UPLC I-Class System Fixed Loop (FL) or Flow Through Needle (FTN) |
|
Column(s): |
ACQUITY UPLC BEH Amide 2.1 × 100 mm, 1.7 μm |
|
Column temp.: |
45 °C |
|
Flow rate: |
0.6 mL/min |
|
Mobile phase A: |
95:5 Acetonitrile/water +10 mM Ammonium acetate |
|
Mobile phase B: |
50:50 Acetonitrile/water +10 mM Ammonium acetate |
|
Gradient: |
0.1% to 20.0% B for 2 minutes, then 20% to 80% B for 3 minutes followed by 3 minutes re-equilibration |
|
Run time: |
8 minutes |
|
Injection volume: |
1 μL |
|
MS systems: |
Xevo TQ-S micro, Xevo TQ-XS, and Xevo TQ-S |
|
Ionization mode: |
ESI (+) |
|
Capillary voltage: |
2.8 kV (+) |
|
Acquisition mode: |
MRM |
|
Source temp.: |
120 °C |
|
Desolvation temp.: |
500 °C |
|
Cone gas flow: |
150 L/hr |
|
Desolvation flow: |
1000 L/hr |
|
Nebulizer gas: |
7 bar |
|
Ion guide offset 1: |
3 V |
|
Ion guide offset 2: |
0.3 V |
A LipidQuan Quanpedia method file (version 1.4) was generated, that contains the LC conditions, MS method, and associated TargetLynx processing method (including retention times and MRM transitions). The resulting data were processed with either TargetLynx or Skyline (MacCoss Lab Software, University of Washington).
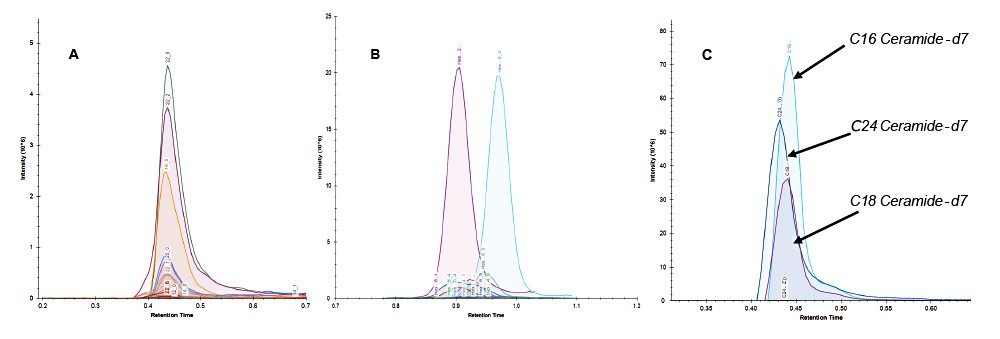
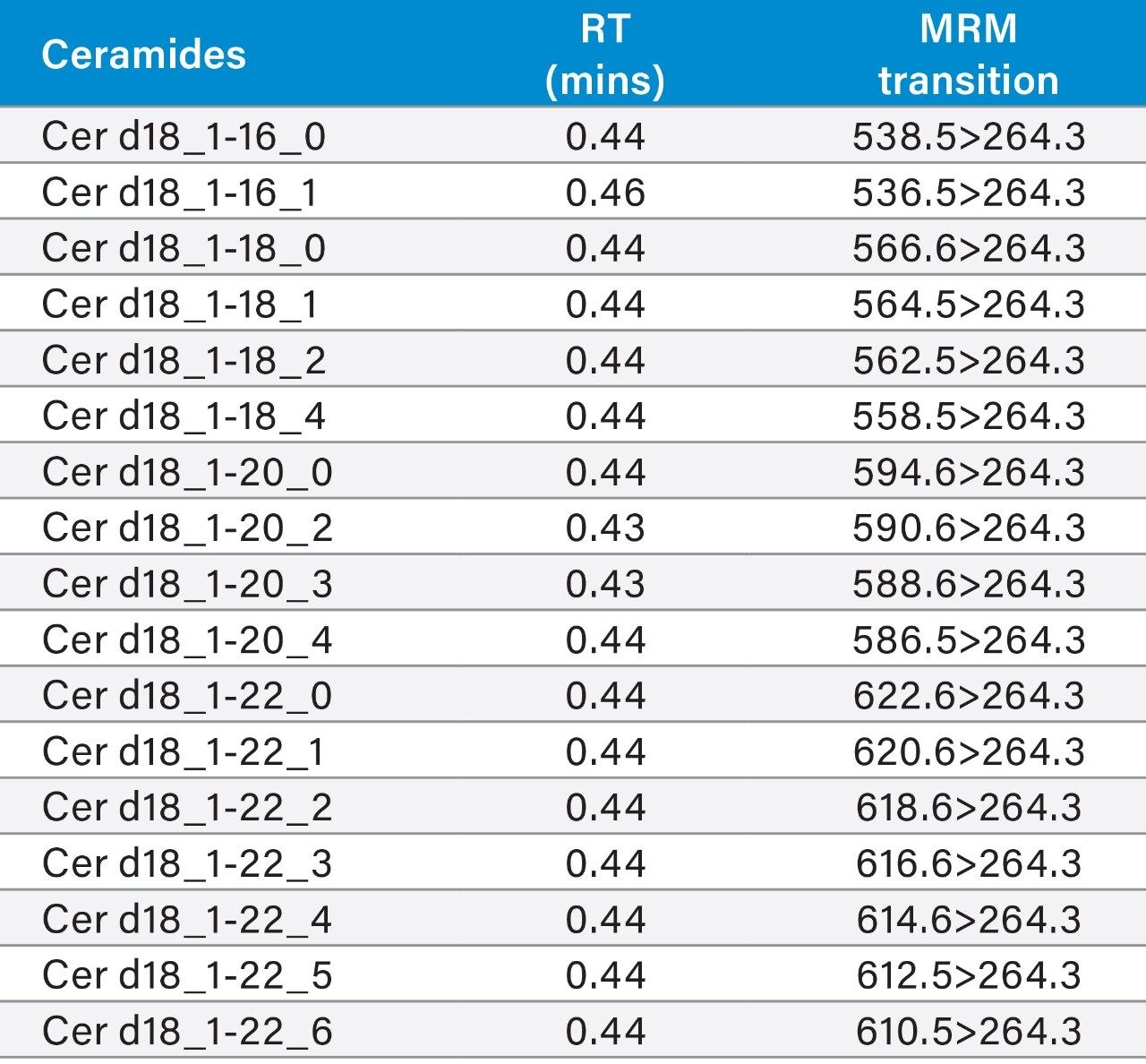
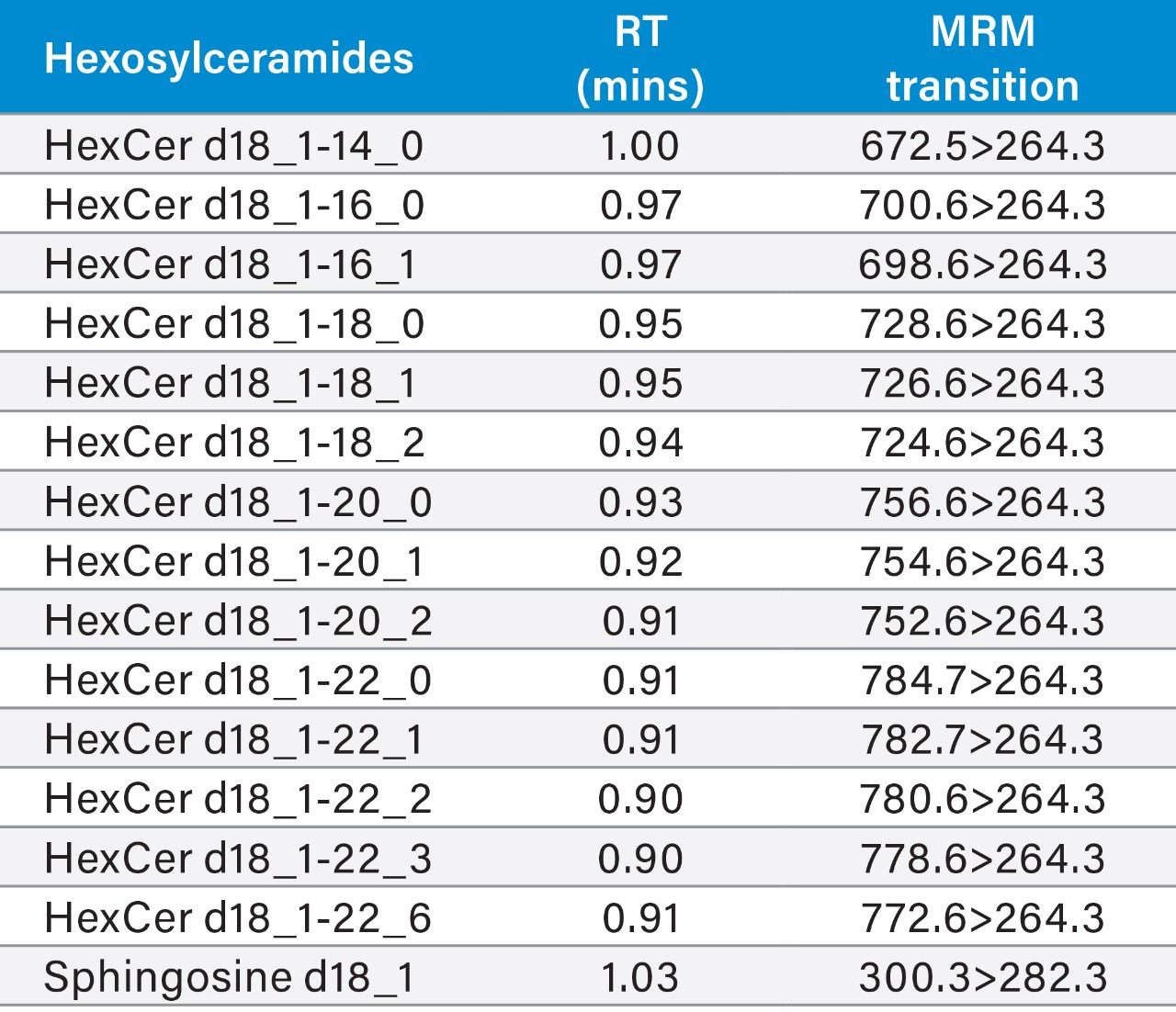
LipidQuan employs MRM in positive ion mode to identify and quantify Cer and HexCer species contained in human plasma whilst simultaneously analyzing several other lipid classes such as TG, PC, and SM. Cer elutes at the solvent front (~0.4 mins) while HexCer and SPH lipids elute as discrete bands (~0.8–1.1 mins) under HILIC-based conditions (Figure 3). A total of 24 Cer, 23 HexCer, and SPH were analyzed. The method sensitivity easily facilitated the detection of these lipids in human plasma at normal circulating levels from 50 μL of plasma while demonstrating a linear dynamic range over four orders of magnitude. The Cer and HexCer lipid transitions discussed here use the lipid precursor ion and characteristic long chain base (LCB) product ion (m/z 264) (Figure 1). These transitions are tabulated in Tables 1 and 2.
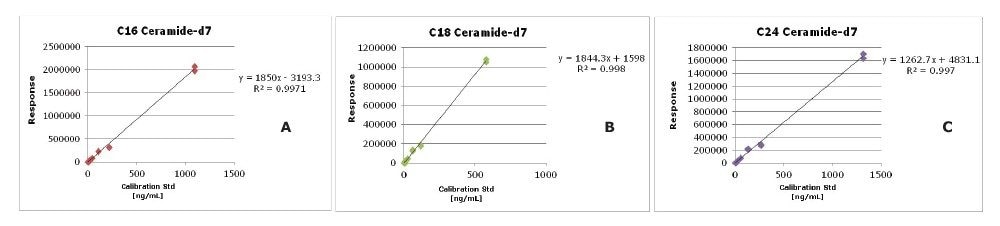
Quantification was achieved using calibration curves of plasma spiked with known concentrations of SIL standards prior to extraction. By using surrogate standards prepared and analyzed under identical conditions to those of endogenous lipids, the quantification of endogenous lipids within the same class was achieved. The use of a commercially available premixed SIL solution rather than a SIL for each measured lipid significantly reduces the overall cost of large studies. Example calibration curves for SIL standards C16 Ceramide-d7 (d18:1-d7/16:0), C18 Ceramide-d7 (d18:1-d7/18:0) and C24 Ceramide-d7 (d18:1-d7/24:0) are shown in Figure 4 and can be used for quantification of endogenous Cer and HexCer. Typical R2 values of 0.95 and deviations from the line of best fit (CVs <30%) are deemed acceptable using the available SIL standards from the deuterated CERAMIDE LIPIDOMIX. The calibration curve for C24:1 Ceramide-d7 (d18:1-d7/24:1) did not meet this criteria.
The development of a LipidQuan Quanpedia method file, allows for the simple download of MRM transitions and chromatographic conditions representing Cer and HexCer lipid species thereby eliminating manual input of LC-MS methods and reducing possible transcription errors.
720006451, February 2019