For research use only. Not for use in diagnostic procedures.
This application note describes the use of the LipidQuan platform that utilizes a HILIC-based approach to perform a targeted screen for FFA without the need for complex sample preparation and long chromatographic separation.
Fatty acids (FA) are hydrocarbon chains comprising both carboxyl (-COOH) and methyl (-CH3)functional groups. Traditional nomenclature designates the carbon atom next to the carboxyl group as α and the subsequent one β carbon with the methyl group carbon designated ω (Figure 1). Fatty acid chains may contain one or more double bonds at specific positions (unsaturated and poly unsaturated with cis (Z) or trans (E) configuration) or they may be fully saturated. Branched chain fatty acids with one or more methyl branches on the carbon chain are more commonly found in prokaryotic organism but have been reported in cow’s milk fat and the gastrointestinal tract of newborns.1,2
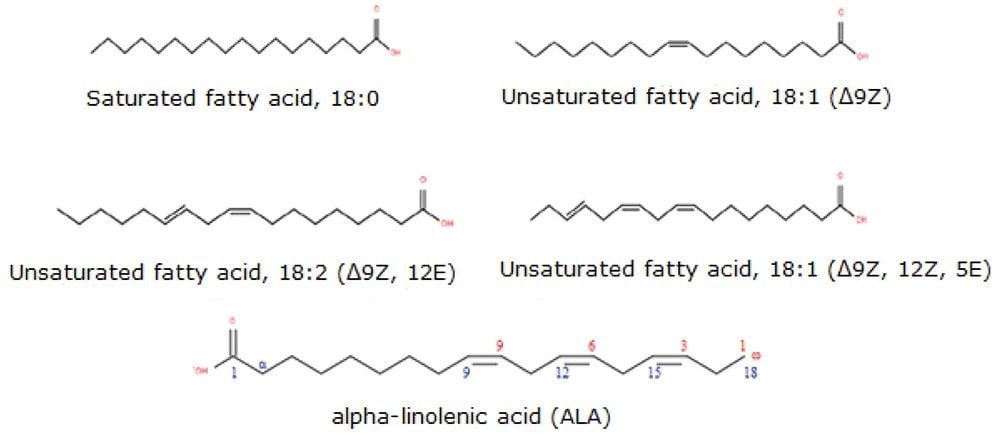
Although free (non-esterified) fatty acids (FFA or NEFA) represent only a small fraction of total fatty acids in plasma, they represent a highly metabolically active lipid class. The most abundant FFAs in plasma consist of oleic acid (18:1), palmitic acid (16:0), and stearic acid (18:0) and together these make up 78% of all FFAs. Some FA species within the unsaturated family are long chain polyunsaturated fatty acids (LC-PUFAs) and are termed “essential” since they cannot be synthesized de novo. Examples include the main PUFAs such as linoleic acid (18:2), arachidonic acid (20:4), ω -linolenic acid (18:3 ω -3), eicosapentaenoic acid (20:5, EPA), and docosahexanaenoic acid (22:6; DCA).3 Eicosanoids are locally acting bioactive signalling lipids derived from arachidonic acid and related polyunsaturated fatty acids (PUFAs). Eicosanoids regulate a diverse set of homeostatic and inflammatory processes linked to numerous diseases including metabolic syndrome disorders and cancers.5
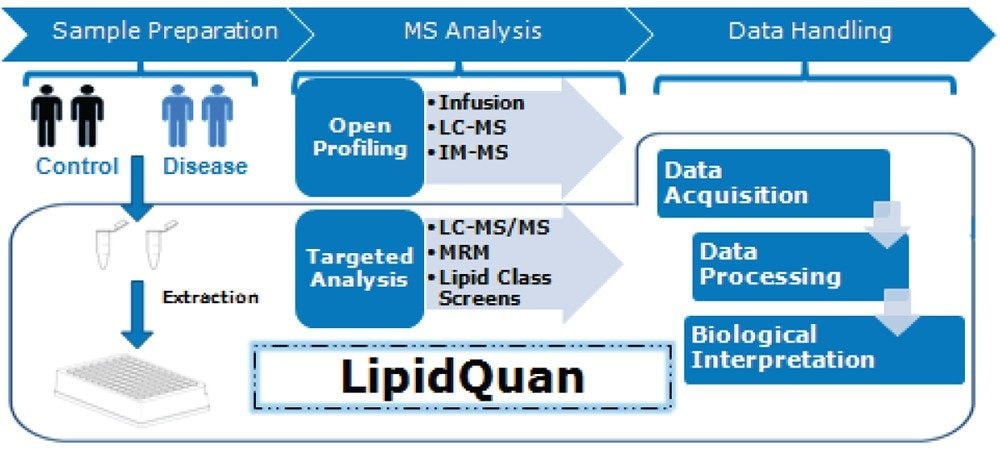
Here, we describe a hydrophilic interaction chromatography (HILIC) based approach for the separation of fatty acids by class prior to Mass Spectrometry (MS) analysis and thereby reducing identification ambiguity.6 An additional benefit of separating lipid species by class is that fewer stable isotope labelled (SIL) standards are required for quantification, conferring a cost savings. This application note describes the use of the LipidQuan platform (Figure 2) that utilizes a HILIC-based approach to perform a targeted screen for FFA without the need for complex sample preparation and long chromatographic separation.
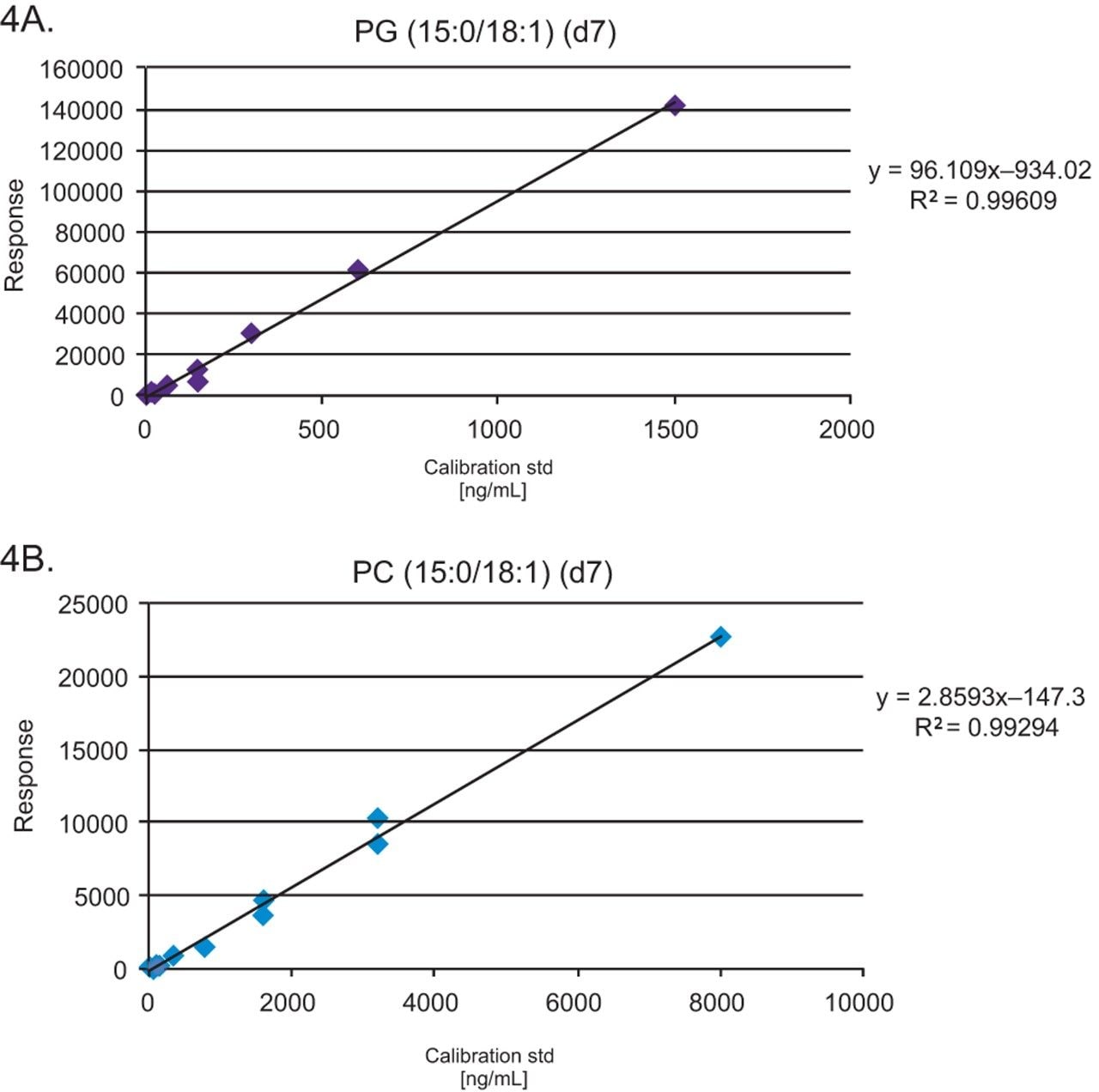
Pooled healthy human plasma was spiked with stable isotope labeled (SIL) standards, (SPLASH LIPIDOMIX, Avanti Lipids, Alabaster, AL) at nine concentration levels to generate calibration curves for quantification. SPLASH LIPIDOMIX does not contain a suitable surrogate standard for the quantification of FFA but the linearity data can be used to assess the quality of the data generated. Typical R2 values of 0.95 and deviations from the line of best fit (CVs <30%) are routinely achieved. PG (15:0–18:1) (d7) = 0.5–1500 ng/mL and PC (15:0–18:1) (d7) = 16–8000 ng/mL are used to illustrate the linearity. Six replicates of the NIST Standard Reference Material 1950 plasma (Sigma Aldrich, Poole, UK) were also spiked with 5% SIL standards, prior to extraction.
A simple sample preparation procedure was adopted using protein precipitation with pre-cooled isopropanol (IPA) (1:5, plasma:IPA). Samples were vortex mixed for one minute and placed at -20 °C for 10 minutes. Samples were vortex mixed again for one minute and placed at 4 °C for two hours to ensure complete protein precipitation. The extracted samples were centrifuged at a maximum of 10,300 g for 10 minutes at 4 °C before transferring the supernatant to glass vials for LC-MS/MS analysis.
|
LC system: |
ACQUITY UPLC I-Class Fixed Loop (FL) or Flow Through Needle (FTN) |
|
Column(s): |
ACQUITY UPLC BEH Amide 2.1 × 100 mm, 1.7 μm |
|
Column temp.: |
45 °C |
|
Flow rate: |
0.6 mL/min |
|
Mobile phase A: |
95:5 Acetonitrile/water + 10 mM Ammonium acetate |
|
Mobile phase B: |
50:50 Acetonitrile/water + 10 mM Ammonium acetate |
|
Gradient: |
0.1% to 20.0% B for 2 minutes, then 20% to 80% B for 3 minutes followed by 3 minutes re-equilibration |
|
Run time: |
8 minutes |
|
Injection volume: |
1 μL |
|
MS systems: |
TQ-S micro, TQ-XS, or TQ-S |
|
Ionization mode: |
ESI (-) |
|
Capillary voltage: |
1.9 kV (-) |
|
Acquisition mode: |
MRM |
|
Source temp.: |
120 °C |
|
Desolvation temp.: |
500 °C |
|
Cone gas flow: |
150 L/hr |
|
Desolvation flow: |
1000 L/hr |
|
Nebulizer gas: |
7 bar |
|
Ion guide offset 1: |
3 V |
|
Ion guide offset 2: |
0.3 V |
A LipidQuan Quanpedia method file (version 1.4) that contains the LC conditions, MS method and associated TargetLynx processing method (including retention times) was generated. The resulting data were processed with either TargetLynx or Skyline (MacCoss Lab Software, University of Washington).
Measurement of NEFA or free FA is typically performed using GC-MS.7 Prior to analysis, samples are typically fractionated using solid phases or liquidliquid extraction techniques before hydrolysis with derivatization agents to form fatty acid methyl esters (FAME). This protocol is time consuming, risks derivatising intact complex lipids, and does not always perform well for the longer chain, less volatile FAs (>C24).
Alternatively, reversed-phase (RP) LC-MS can also be applied for the analysis of FFA. This method, however, also requires time-consuming sample preparation and the use of toxic organic solvents that can be expensive to purchase and dispose of. Reversed-phase chromatography separates lipids according to the chain-length and the degree of unsaturation. The dual nature of reversed-phase separation (the fatty acyl chain length increases retention time and a double bond in the fatty acyl chain reduces retention time) can hamper the analysis of real samples. Further, with the large number of components that co-elute, identification becomes difficult.
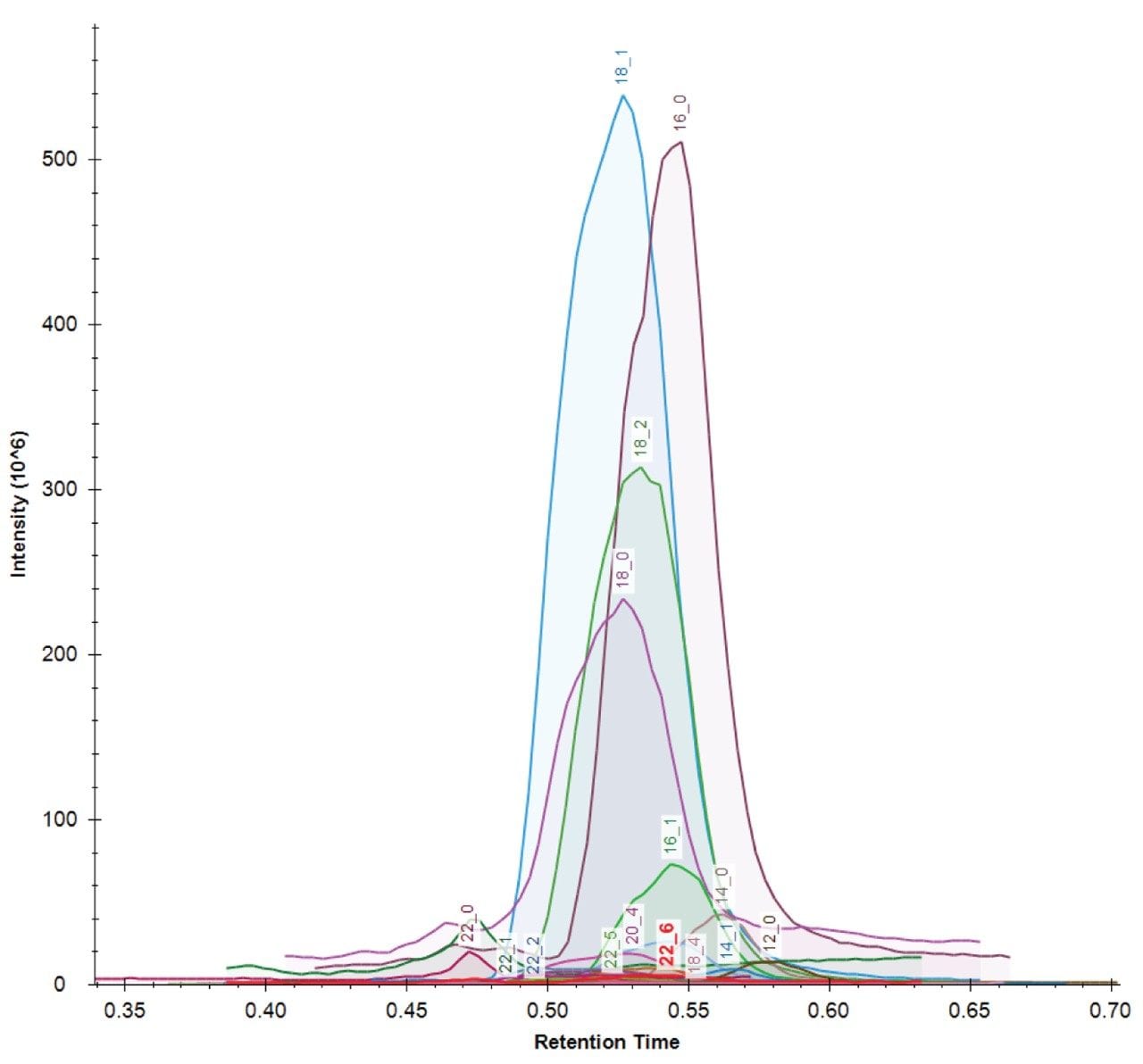
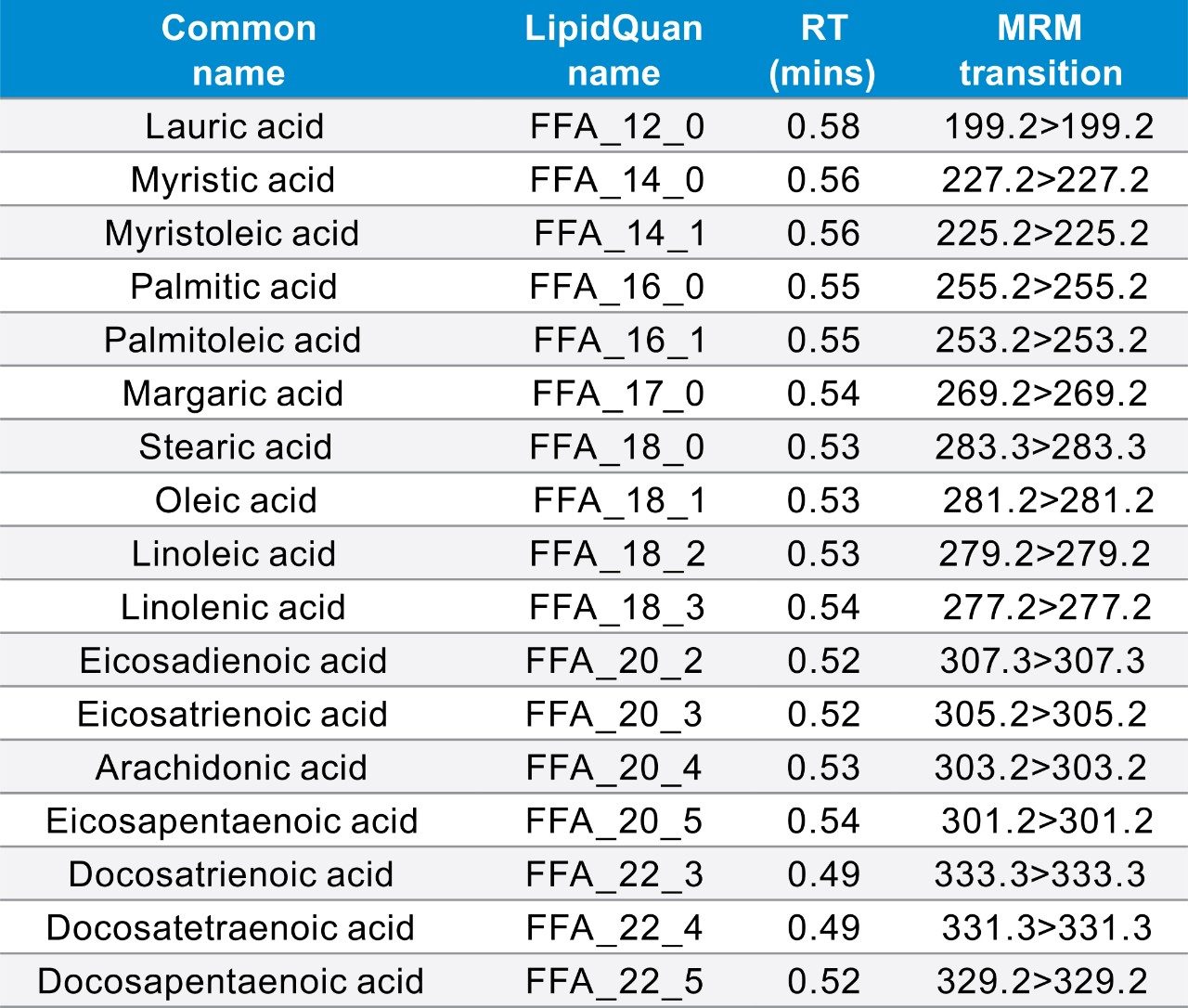
LipidQuan employs pseudo MRM in negative ion mode to identify and quantify 24 FFA species contained in human plasma whilst simultaneously analysing several other phospholipid classes from the same injection. FFAs elute as a discrete band (~0.5 mins) under HILIC-based conditions (Figure 3). The method sensitivity facilitated the detection of these lipids in human plasma at normal circulating levels from 50 μL of plasma. The dynamic range was linear over four orders of magnitude.
The data generated was acquired using the curated LipidQuan plasma screens (LipidQuan Quanpedia file v1.4) to provide relative quantification. Table 1 shows the lipid species with CVs <30%. Typical R2 values of 0.95 and deviations from the line of best fit (CVs <30%) are routinely achieved using the available SIL from SPLASH LIPIDOMIX. The calibration curve for PG (15:0–18:1) (d7) = 0.5–1500 ng/mL and PC (15:0–18:1) (d7) = 16–8000 ng/mL from the same analysis are shown (Figure 4).
A hydrophilic interaction chromatography (HILIC) based approach avoids the need for derivatization and complex sample preparation associated with FAME GC-MS. Additionally, since lipids are separated by class, the risk of co-elution associated with RP LC-MS is also minimised.
The development of a LipidQuan Quanpedia method file enables a simple download of MRM transitions and chromatographic conditions representing FFAs, eliminating manual input of LC-MS methods and reducing possible transcription errors.
720006464, January 2019