This application note reports a routine system check to assess LC-MS system performance prior to intact protein quantification (intact or subunit analysis). Also this procedure assures optimal and robust performance. The method can also be used for troubleshooting a system that is presenting abnormal behaviors.
In recent years, there has been a significant increase in interest in the (direct) bioanalysis of intact proteins, such as monoclonal antibodies (mAbs).1 As a result, bioanalytical scientists are increasingly interested in performing intact protein analysis. To be successful and generate reliable data, it is imperative that the user ensures: proper conditioning of the LC-MS system, careful sample preparation, and general care when setting up for analysis. Without proper system setup, it is common to observe: little or no signal of the analyte; poor chromatographic performance; and suboptimal sensitivity.
This document is designed to help the user performing intact and/or subunit protein analysis evaluate the performance or suitability of the instrument by undertaking a simple system check (SC) and running of QC standards. The status of the system is evaluated by running the NIST mAb subunit standard and assessing the following parameters: retention time reproducibility, chromatographic peak resolution, sensitivity, accuracy, and precision. Herein we show how to ensure an LC-QTof platform is ready for performing intact-level protein and subunit quantification using full scan Tof-MS.
This test should be routinely performed prior to the bioanalysis of intact proteins (e.g. mAbs: intact and subunit), particularly when the lab is routinely switching molecule classes and changing LC conditions. This simple guide will walk users through the steps required to ensure optimum performance of the Xevo G2-XS QTof Mass Spectrometer prior to running large molecule bioanalysis, thus ensuring confident sample analysis.
This should be performed as needed and is useful when switching from other application work, such as peptide and/or small molecule analysis (Figure 1).
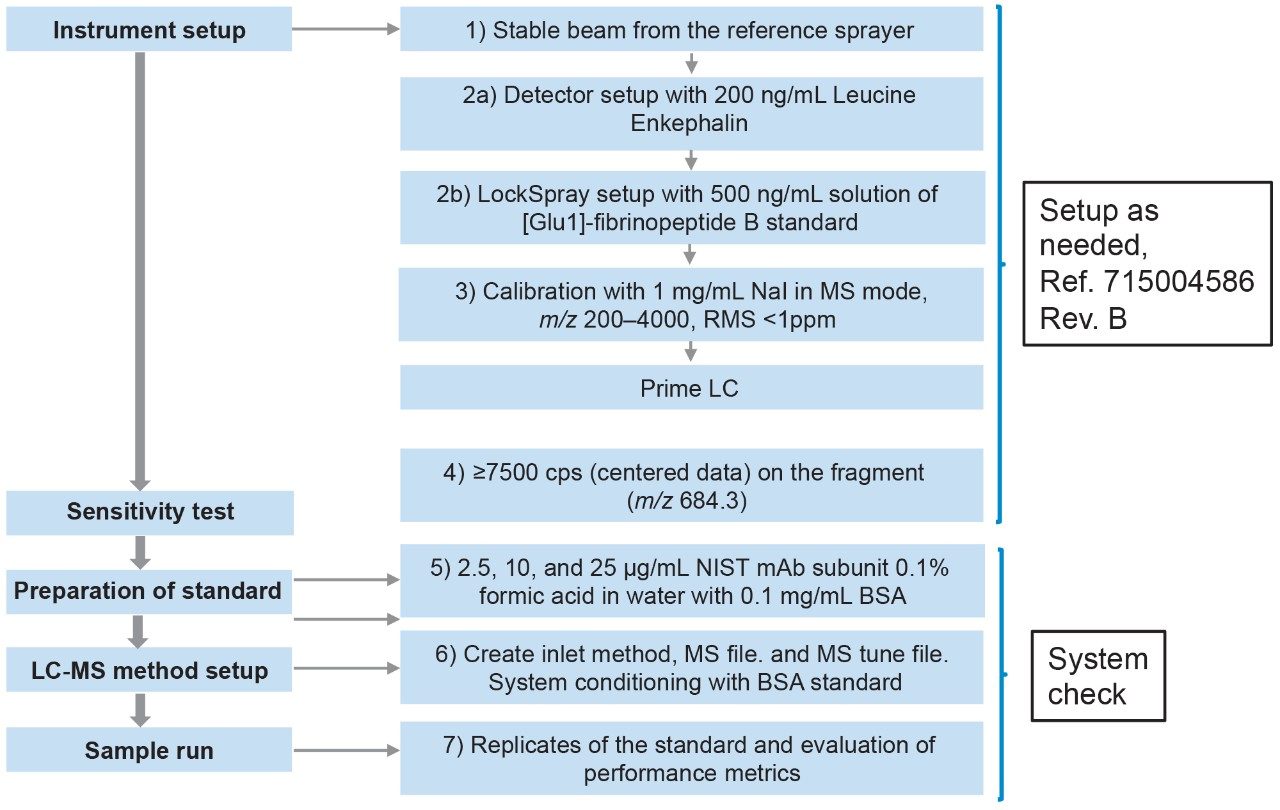
1. Check that the tune page shows a stable beam from the reference sprayer.
2. IntelliStart LockSpray setup should be performed using a 500 ng/mL solution of [Glu1]-Fibrinopeptide B standard (GFP).
3. Calibration of the instrument over the mass range of m/z 200–4000 Da should be performed with a solution of 1 mg/mL NaI in positive MS mode. Mass calibration should give an RMS of <1 ppm for a well calibrated system.
Note 1: If sensitivity or calibration is poor, detector setup should be performed using IntelliStart with a 200 ng/mL leucine enkephalin solution. It is not necessary to perform the detector setup on a regular basis, unless the instrument has been vented, or after critical maintenance has been undertaken on the instrument. Once completed, repeat sensitivity test and calibration procedure.
Note 2: In general, large molecule instrument setup parameters (also called proteomics parameters) described in the installation manual should be followed.
This document is intended as a guide for optimizing a system that is setup according to the official installation manual (715004586 Rev. B). Key steps are summarized here. For detailed test instructions, please refer to the protocol described in the Xevo G2-XS QTof installation manual.
4. A 100 fmol/μL solution of GFP is directly infused through the analyte sprayer at 5 μL/min. The signal height must be ≥7500 counts per second (centered data) on the fragment ion (m/z 684.3) from the MS/MS. The mass resolution must be ≥15,000 (FWHM), measured on the fragment ion (m/z 175.1).
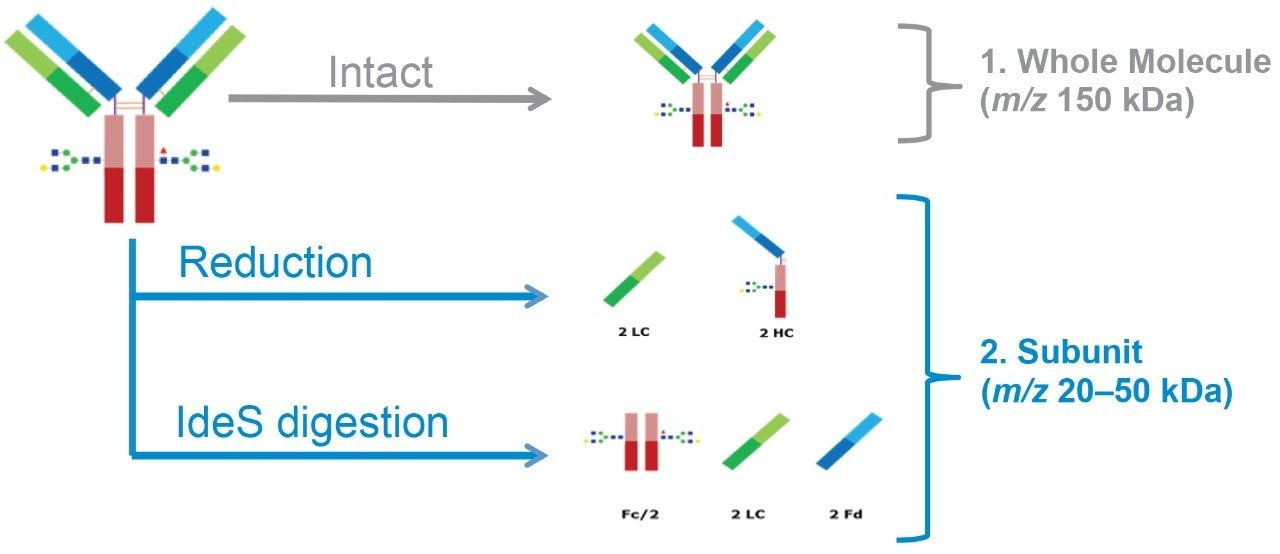
The NIST mAb subunit results from the reduction and IdeS digestion of the NIST Reference Material 8671 (Figure 2). Following successful setup of the LC and MS, the samples are prepared as follows:
5a. Reconstitute NIST mAb subunit standard vial in 100 μL of 0.1 mg/mL BSA in 0.1% formic acid in water. Aliquots can be made and stored at -20 °C long term.
5b. Prepare dilution of the standard at 2.5, 10, and 25 μg/mL in the same buffers mentioned above. These solutions should be stored at 4–8 °C and must be used over a period of five days. It is recommended to make the dilutions fresh prior to analysis in order to obtain reproducible results as it was noticed during stability and reconstitution testing that some losses of protein could occur even in the presence of BSA. It should be noted that the system may require further conditioning with a proteinaceous sample (BSA standard or precipitated plasma supernatant) prior to running the subunit standard. This also ensures that any available binding sites on the instrument, or any other material used for sample preparation, are blocked.
6. The BSA standard or plasma supernatant should be injected until signal intensity of the analyte is stable.
7. Run replicates/standards as outlined in the section below.
|
LC system: |
ACQUITY UPLC H-Class PLUS (FTN) |
|
Detection: |
Xevo G2-XS QTof, ESI+, sensitivity mode |
|
Column: |
ACQUITY UPLC Protein BEH C4, 300 A, 1.7 μm, 2.1 × 50 mm (p/n: 186004495) |
|
Column temp.: |
80 °C |
|
Sample temp.: |
8 °C |
|
Injection volume: |
5 μL |
|
Mobile phase A: |
0.1% formic acid in water |
|
Mobile phase B: |
0.1% formic acid in acetonitrile |
|
Time (min) |
Flow rate (mL/min) |
%A |
%B |
Curve |
|---|---|---|---|---|
|
Initial |
0.400 |
85.0 |
15.0 |
6 |
|
0.50 |
0.400 |
85.0 |
15.0 |
6 |
|
7.00 |
0.400 |
35.0 |
65.0 |
6 |
|
7.50 |
0.400 |
10.0 |
90.0 |
6 |
|
8.00 |
0.400 |
85.0 |
15.0 |
6 |
|
8.50 |
0.400 |
10.0 |
90.0 |
6 |
|
9.00 |
0.400 |
85.0 |
15.0 |
6 |
|
11.00 |
0.400 |
85.0 |
15.0 |
6 |
|
Ionization mode: |
ESI+ |
|
Capillary: |
2.75 kV |
|
Cone: |
70 V |
|
Source offset: |
80 V |
|
Source temp.: |
125 °C |
|
Desolvation temp.: |
600 °C |
|
Desolvation gas flow: |
600 L/hr |
|
Cone gas: |
10 |
|
RF settings were default values as displayed in the installation guide document (715004586 Rev. B). |
|
Acquisition mode: |
MS continuum |
|
Acquisition time: |
3.5 min |
|
Range: |
500–4000 |
|
MS scan time: |
0.5 sec |
|
Target ions: |
941.654 (Fc/2), 1006.499 (LC) |
|
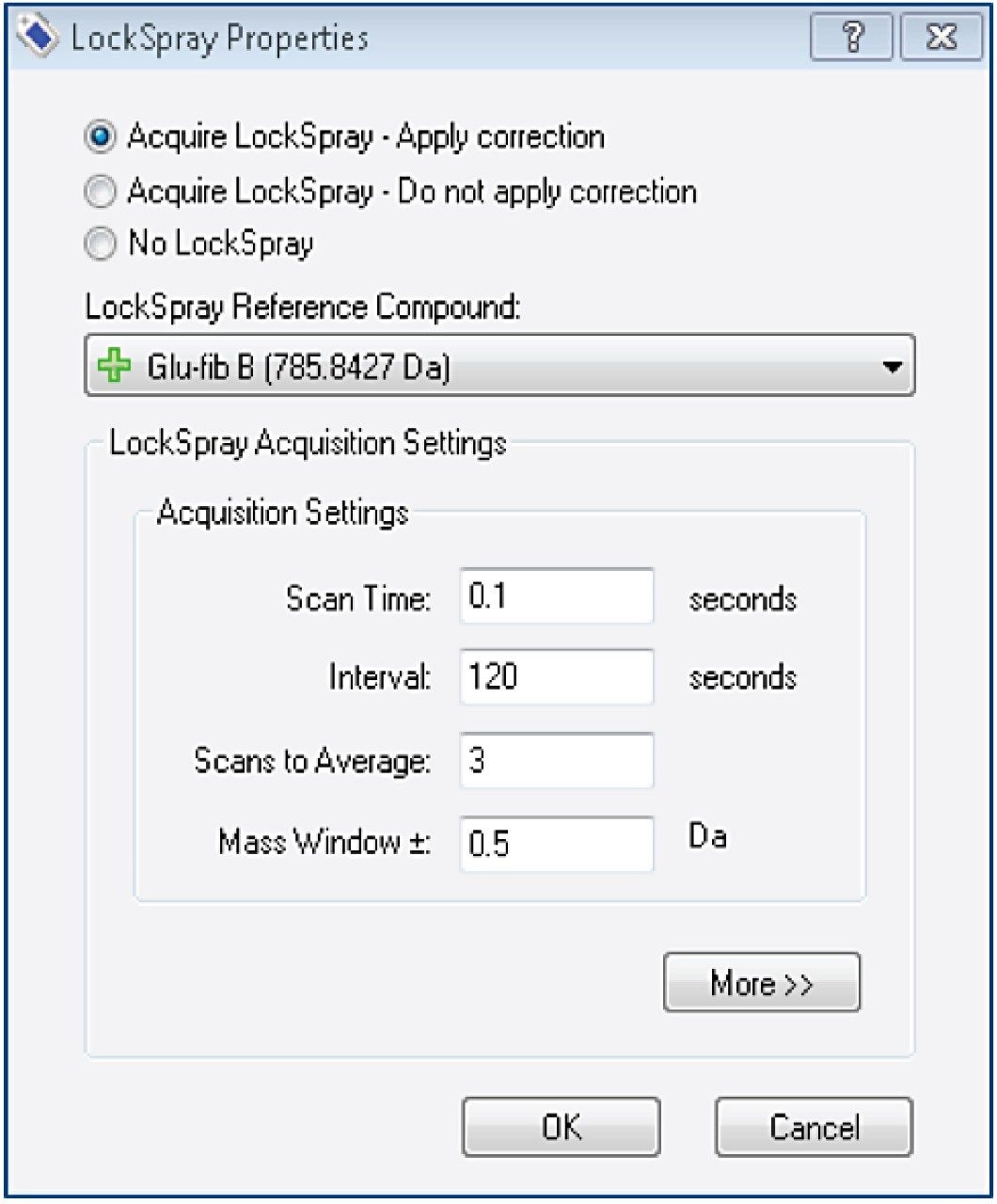
(acquire LockSpray – apply correction): |
|
|
LockSpray reference compound: |
GFP (785.8426) |
|
Reference scan time: |
0.1 sec |
|
Lock mass frequency (interval, sec): |
120 sec |
|
Scans to average: |
3 |
|
Mass window: |
±0.5 Da |
|
(Precursor-precursor): |
|
|
Acquisition mode: |
MS continuum |
|
Acquisition time: |
3.5 min |
|
Range: |
500–2000 |
|
MS scan time: |
0.2 sec |
|
MRM window: |
None |
|
Precursor mass: |
941.6>941.654 (Fc/2), 1006.5>1006.499 (LC) |
For the development of the system check, the mAb subunit standard derived from the NIST by reduction and IdeS digestion was used. The three main components of the standard are Fc/2, LC and Fd’. These mAb subunits have theoretical masses above 23,000 Da. There was nonspecific binding observed when the standard was diluted in aqueous or buffer solutions. To mitigate, 0.1 mg/mL bovine serum albumin (BSA) is recommended where possible. In this example, Fd’ coelutes with BSA, therefore, only the Fc/2 and LC subunits will be monitored for suitability.
Criteria that was monitored included chromatographic peak width, baseline peak resolution of Fc/2 and LC, and sensitivity. The goal of this application was to develop a reproducible method that demonstrates a robust performance and suitability of the Xevo G2-XS QTof prior to running samples for protein quantification.
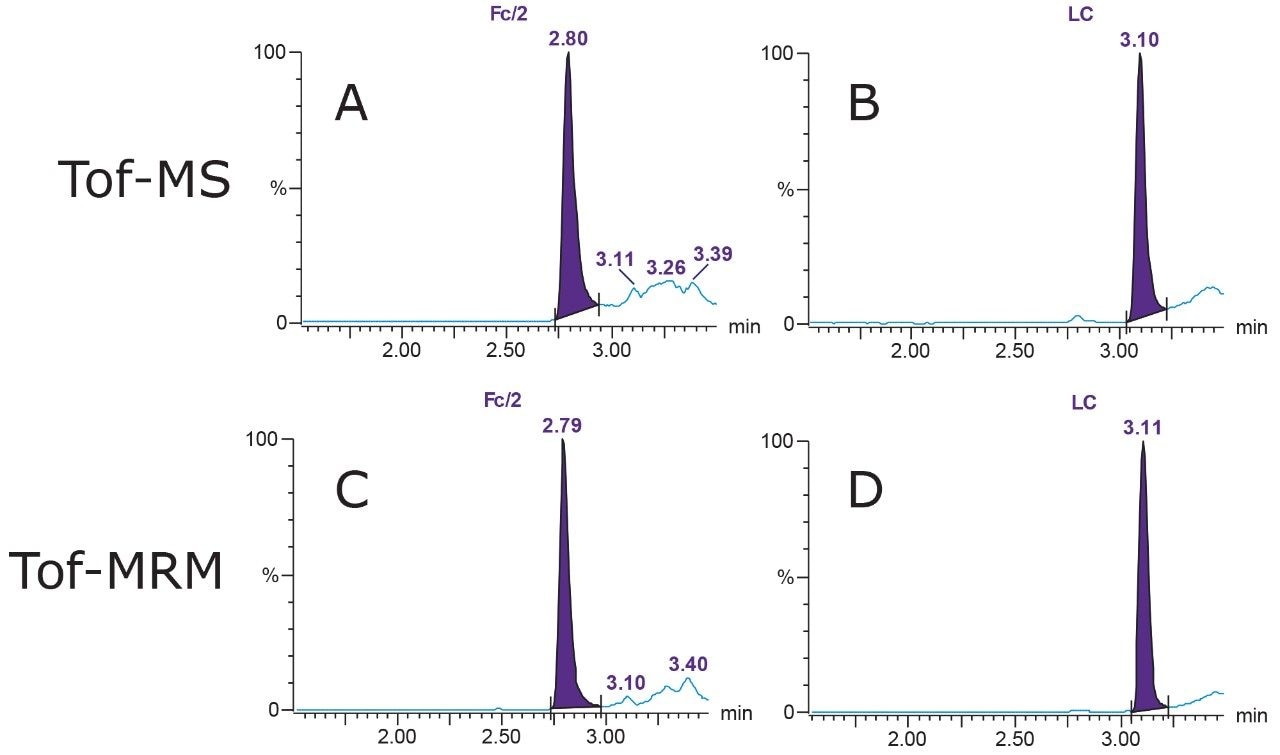
Subunit standards were analyzed on the Xevo G2-XS QTof. To quantify Fc/2, m/z of 941.654, corresponding to the +27-charge state of the G1F isoform, was selected. To quantify LC, m/z of 1006.499, corresponding to the +23-charge state of the molecule, was monitored. Extracted ion chromatograms (XICs) of Fc/2 and LC are shown in Figures 3A and 3B, respectively.
(Optional: Testing these modes ensures targeted modes are available.)
The same m/z masses used in full scan Tof-MS mode were considered, and target enhancement was engaged on the precursors (Figures 3C and 3D).
Linearity
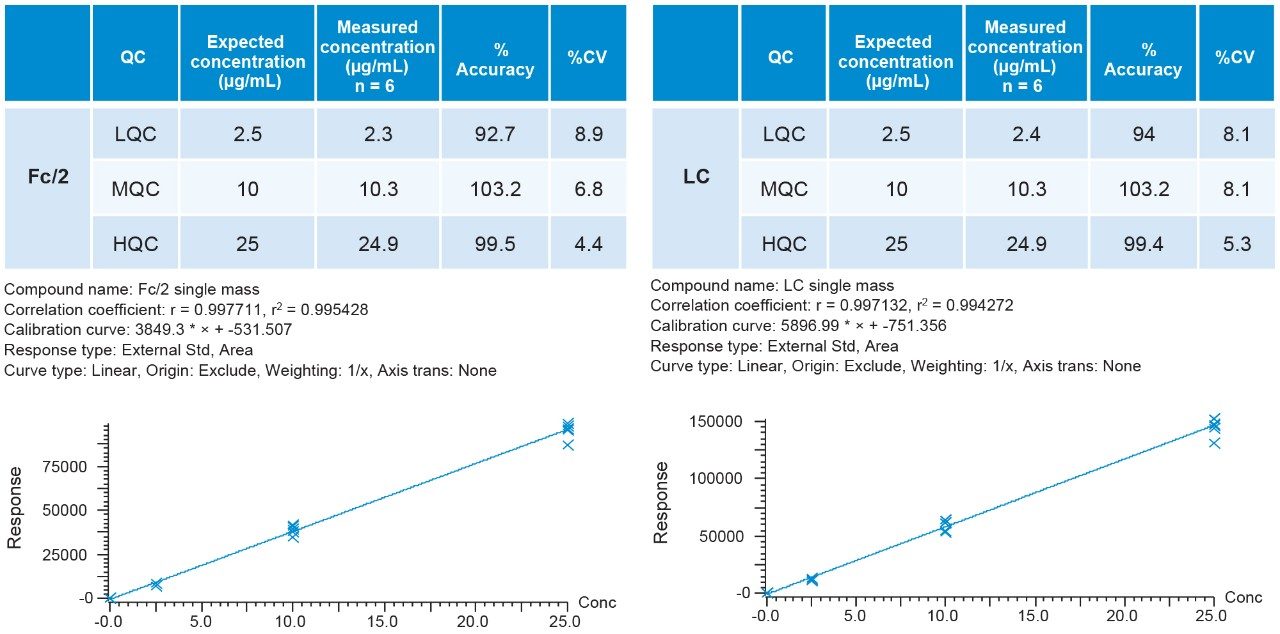
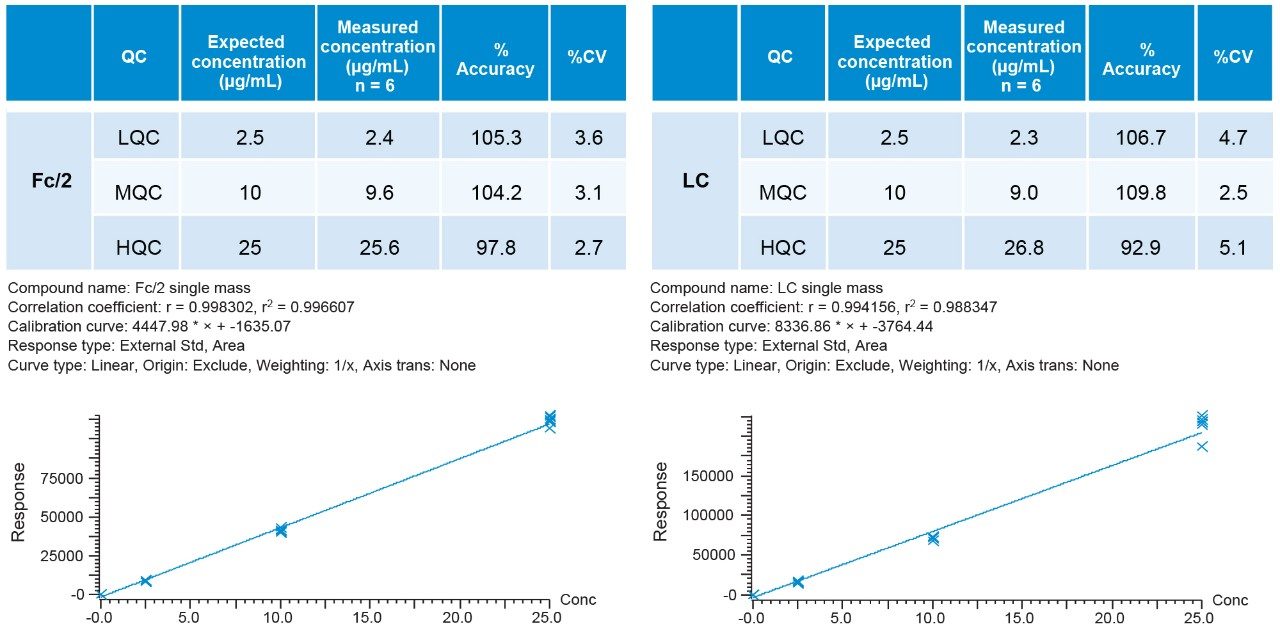
Standards were run in six replicates for the determination of the accuracy and precision. Peak integration and quantification were carried out by using TargetLynx (Figure 4). The precision of all measurements was below 10% for all modes of acquisition, well below the 15% criteria recommended by the FDA guidance.2 Displayed below is an example of %CV for the Fc/2 and LC in Tof-MS, Tof-MRM (Precursor-precursor) modes as well as their respective standard curves (Figures 5 and 6). This is intended as a rapid check for general performance as linear range and suitability testing for analytes of interest is recommended.
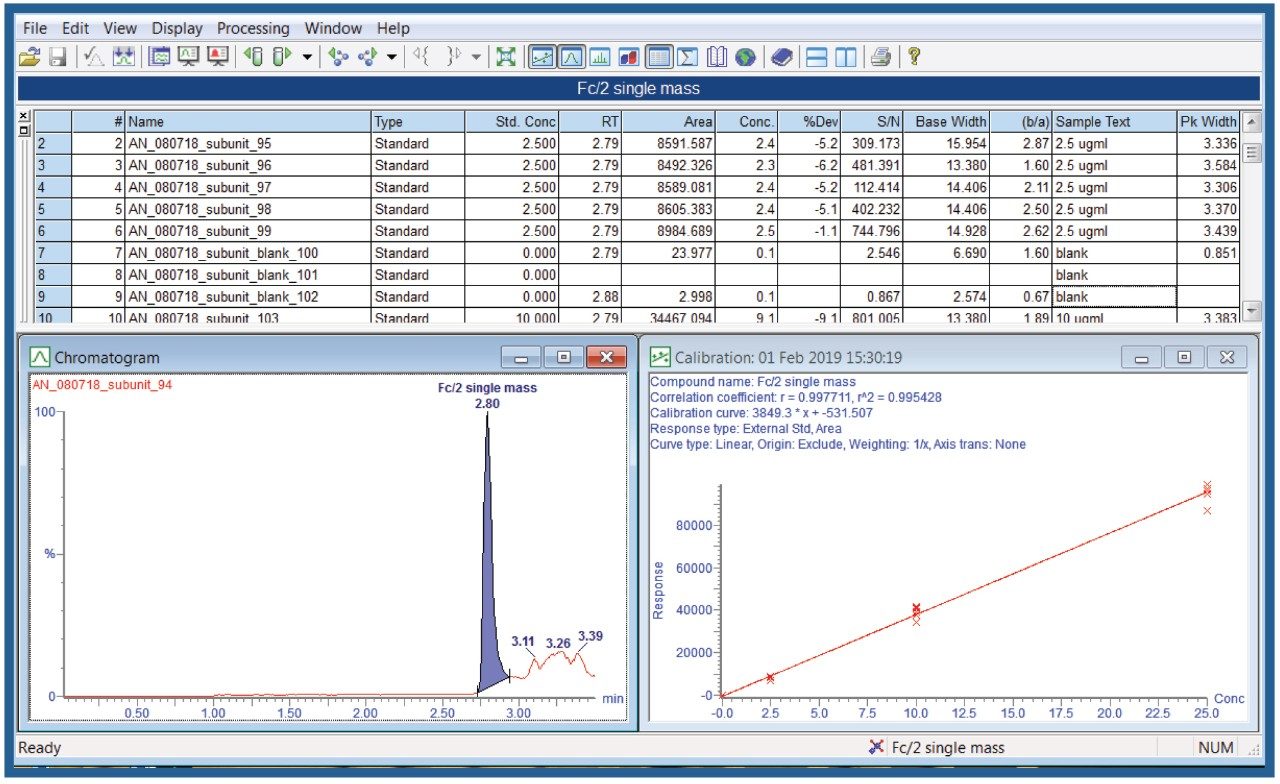
The retention time of the subunits was monitored over an extended period, and reproducibility was determined to be within ±0.1 min. This retention time reproducibility is well within the accepted limits for bioanalytical method validation guidance. Additionally, base width, FWHM (full width at half maximum), and peak asymmetry (b/a) were calculated and were shown to be reproducible on different days of the SC.3 These data are presented in Table 1. Significant increases in retention time reproducibility (>0.1 min) or deviations from typical criteria shown in Table 1 (>30%) could be signs of poor LC performance (e.g., autosampler, pump, injector), higher column backpressure, insufficient conditioning, or even issues related to the sample.

We report a routine system check to assess LC-MS system performance prior to intact protein quantification (intact or subunit analysis). This procedure assures optimal and robust performance. The method can also be used for troubleshooting a system that is presenting abnormal behaviors. Key metrics were measured including retention time, linearity, and chromatographic peak characteristics of subunits. It should be noted that this procedure and standards were not tested to a level expected for use as a system suitability test (SST).
Tof-MS method setup
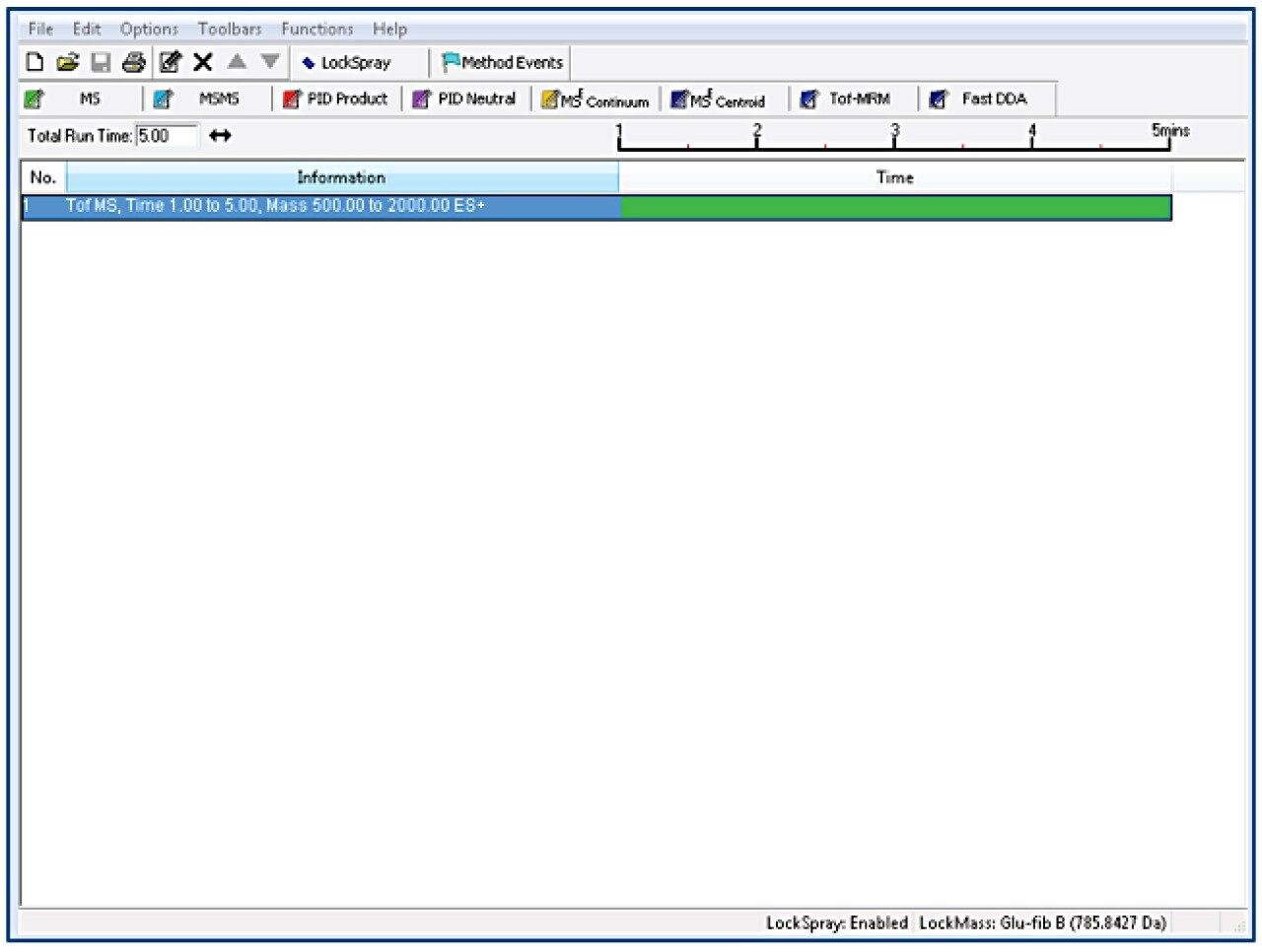
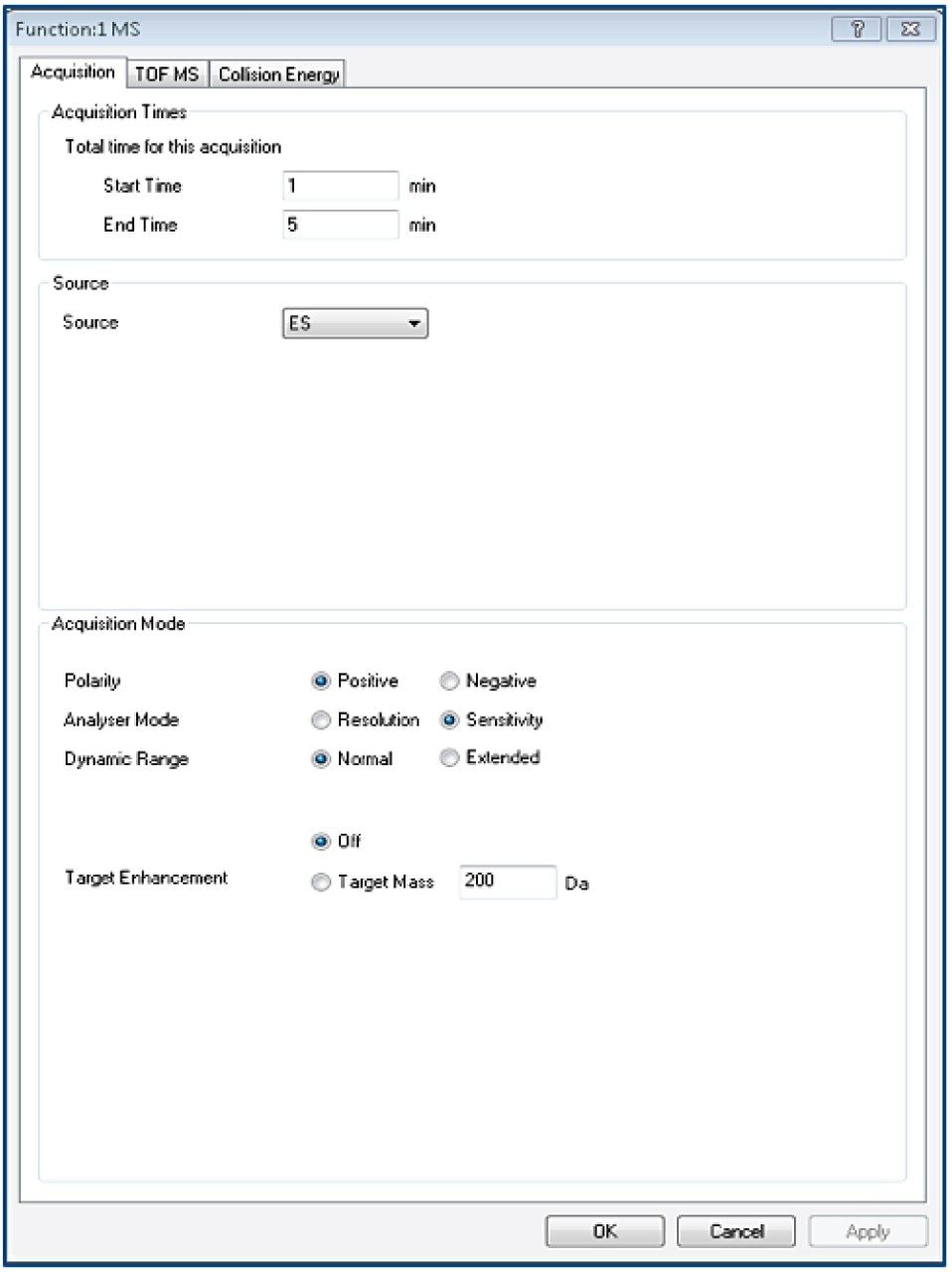
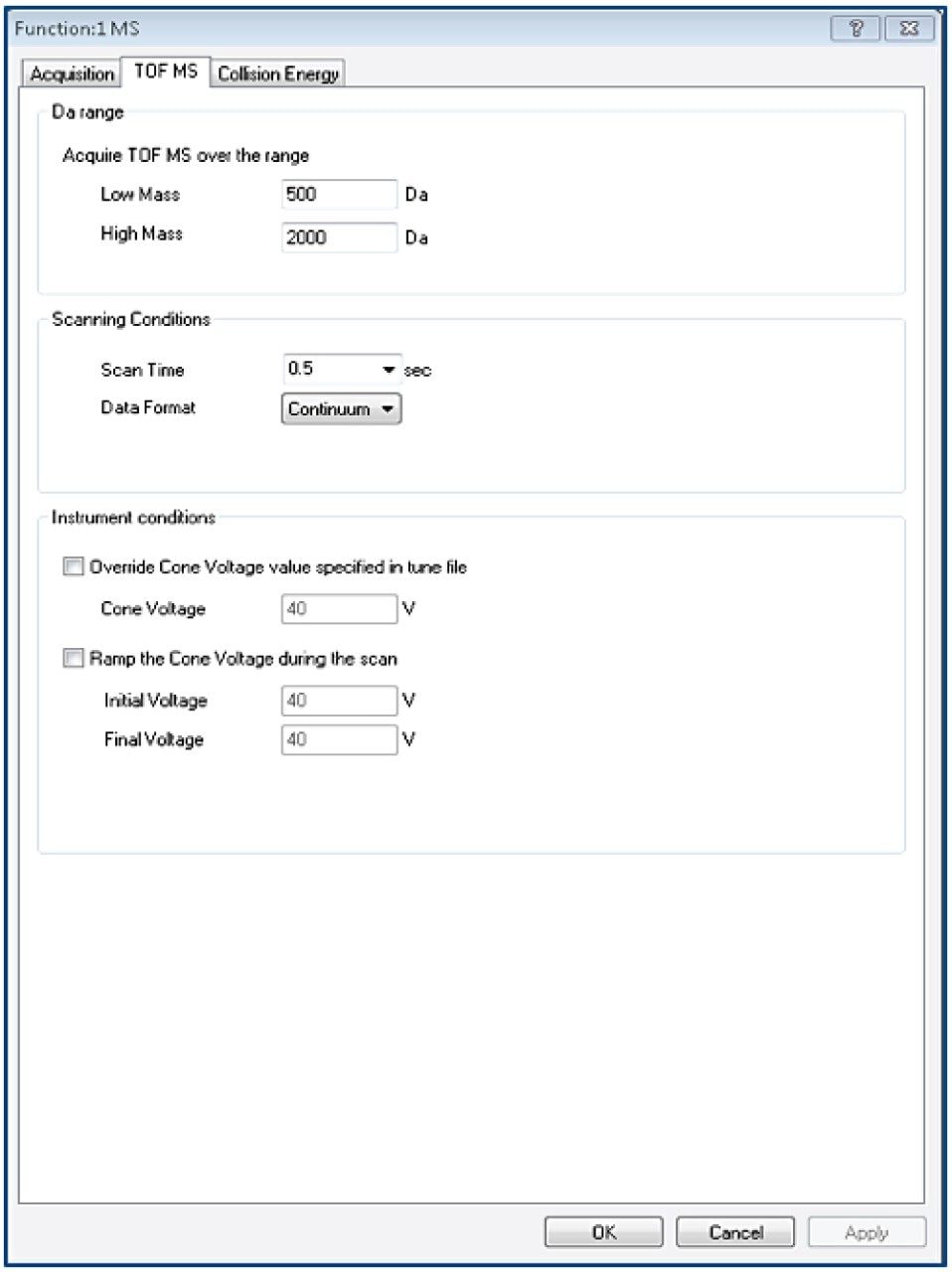
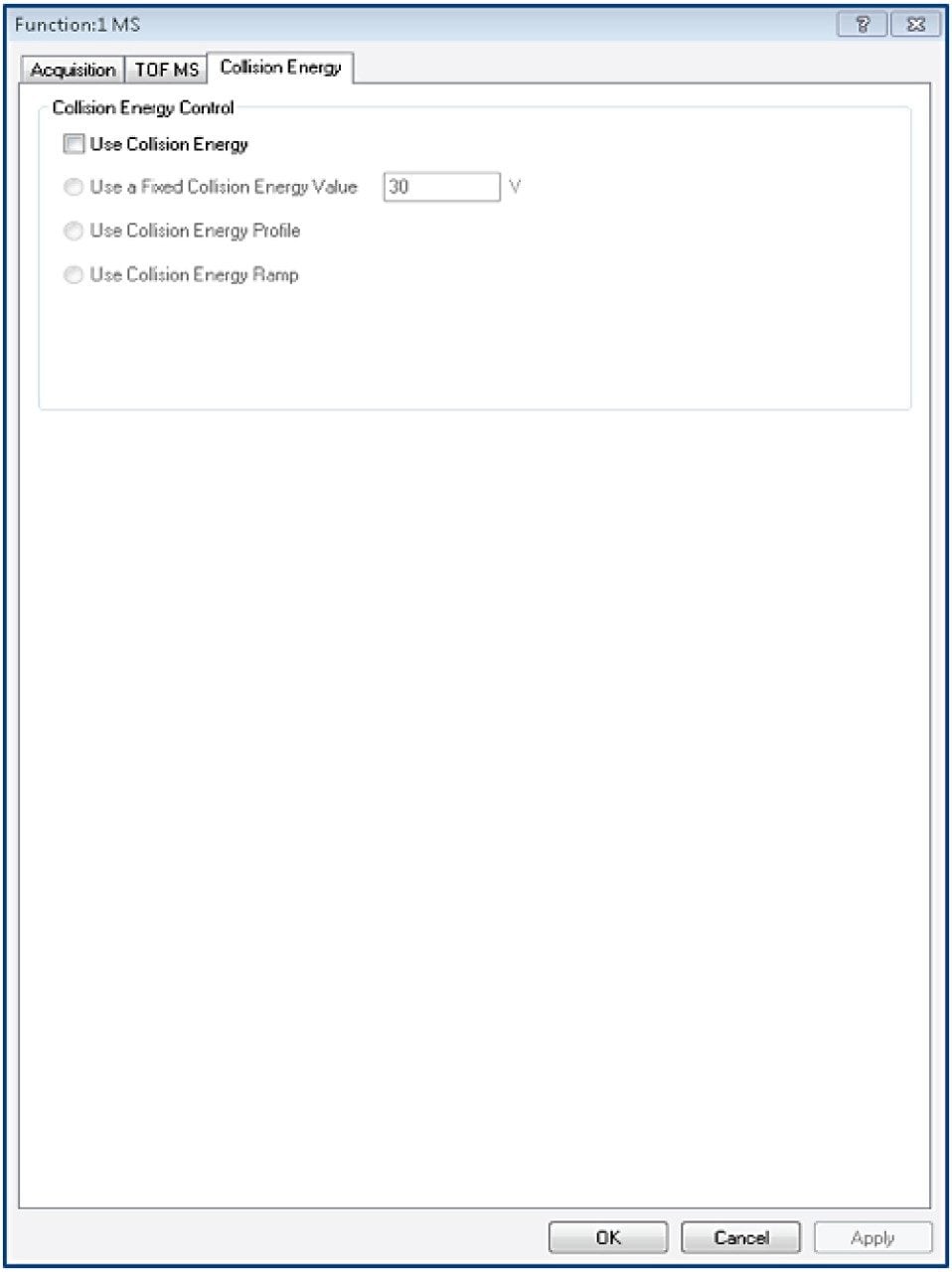
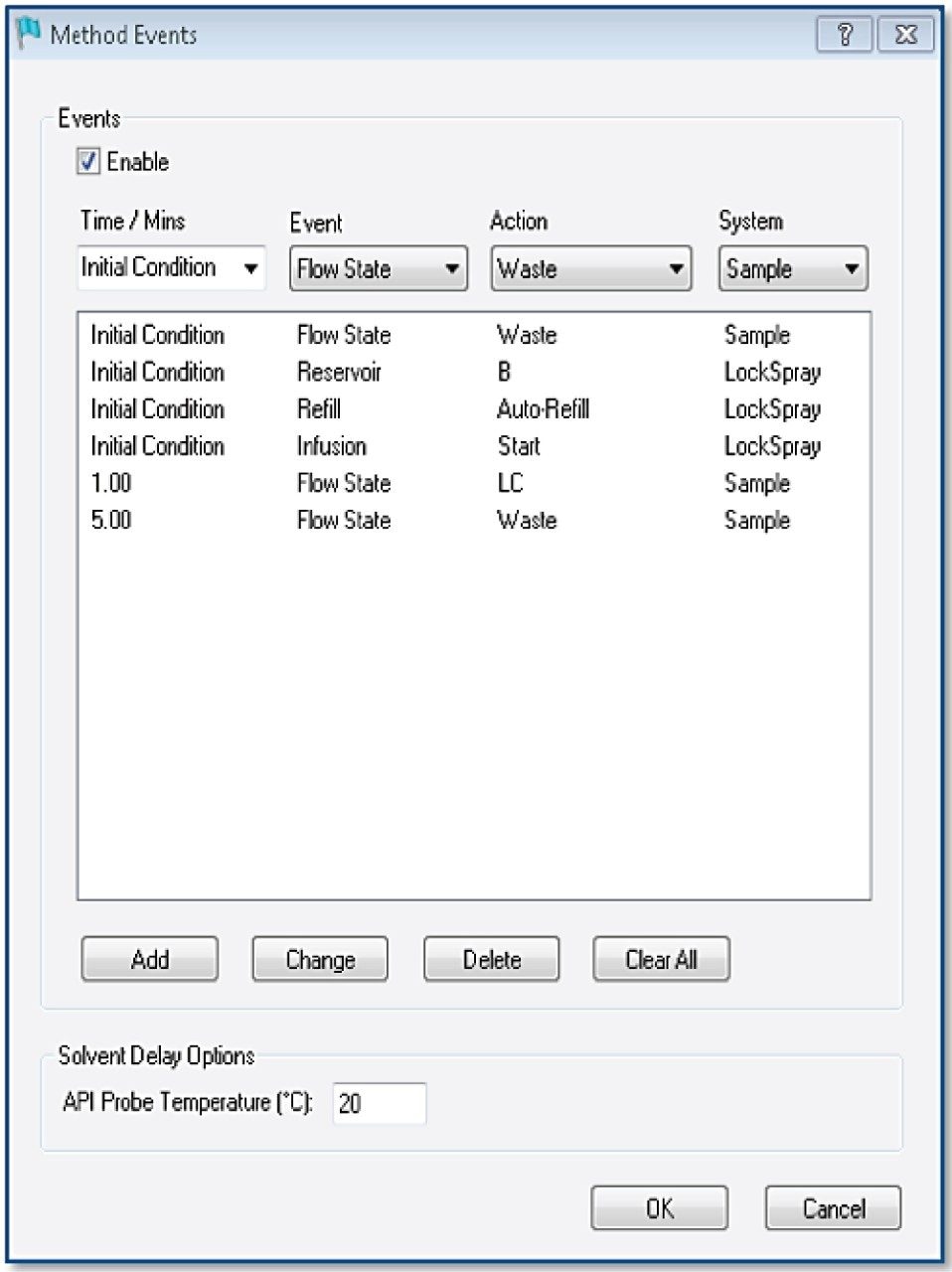
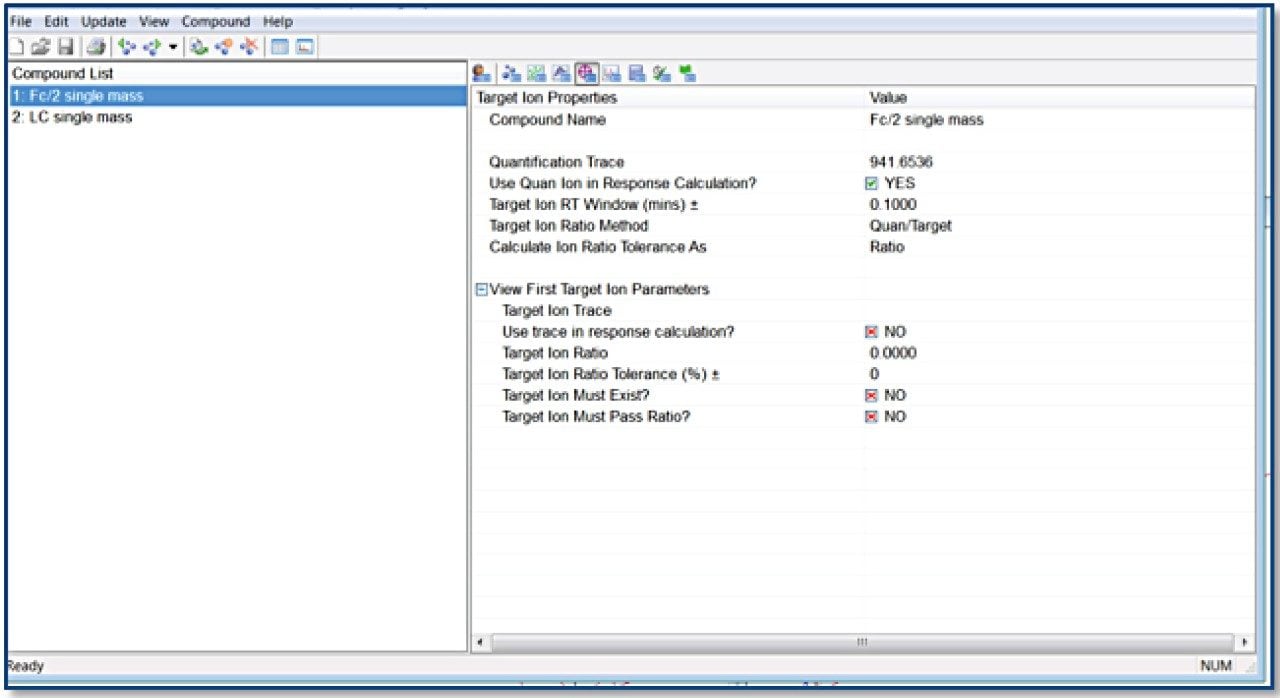
TargetLynx method based on the Tof-MRM (Precursor-precursor) acquisition is displayed. Detailed parameters are shown only for Fc/2.
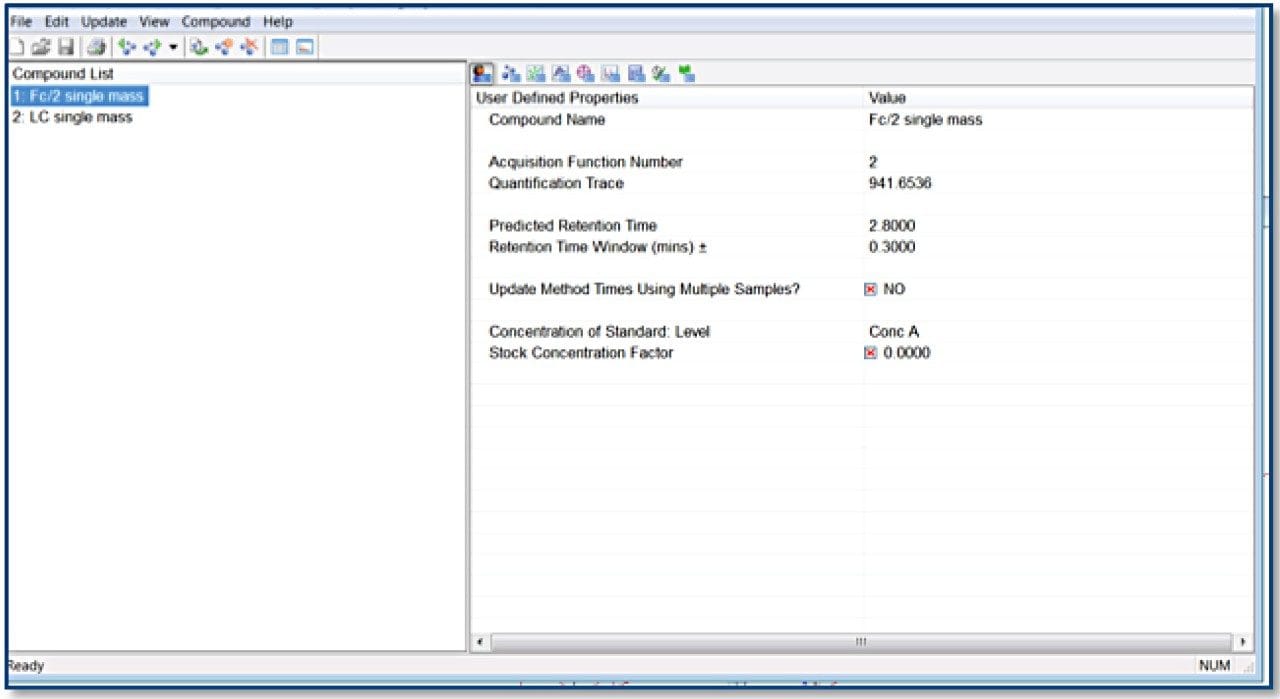
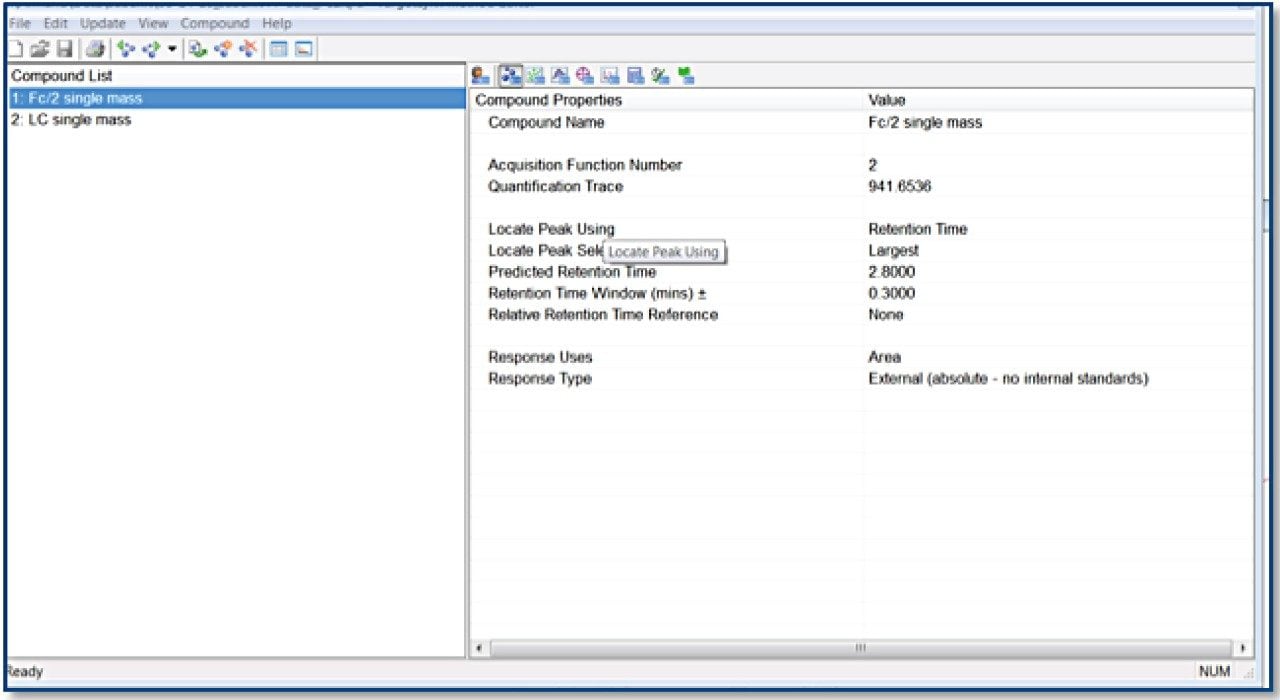
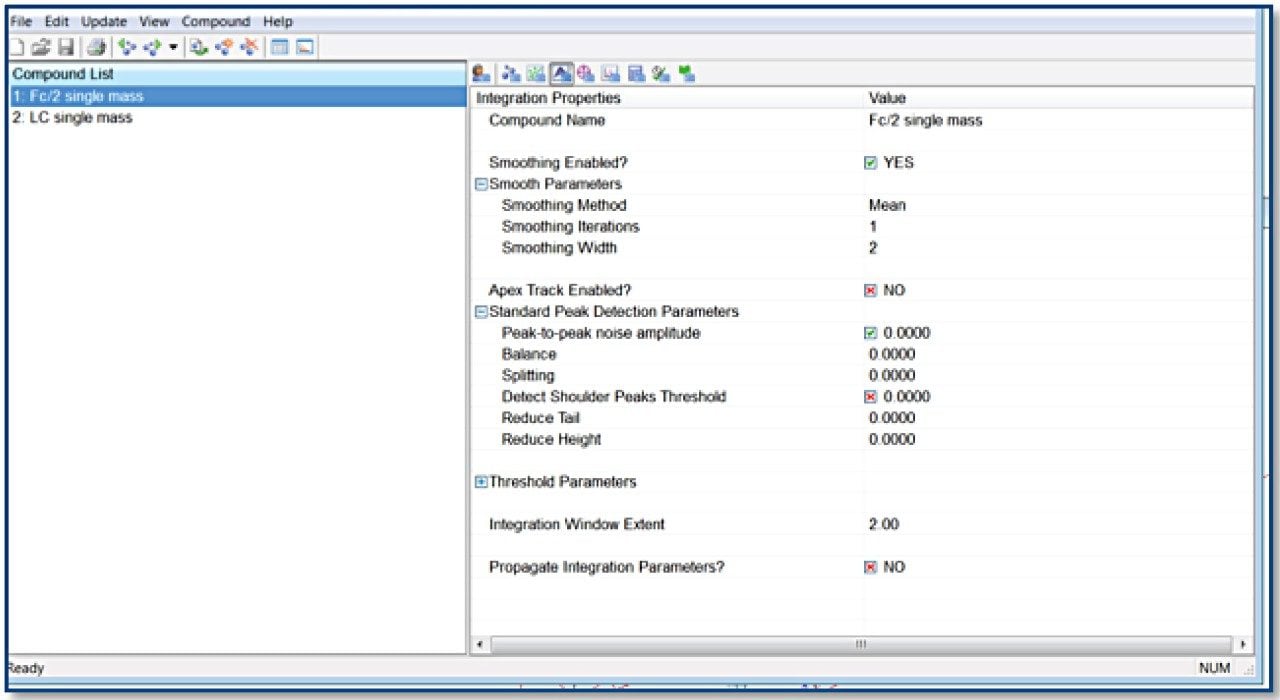
Reagents and materials
Use the following procedure to prepare calibration and lock mass solution:
—Prepare a 1 mg/ml stock solution of leucine enkephalin solution in water
—Take 100 μL of the stock solution and add to 500 mL of 50:50 acetonitrile:water + 0.1% formic acid
—Dissolve 0.1 mg of purchased [Glu1]-Fibrinopeptide B Standard (p/n: 700004729) in 200 mL of water:acetonitrile (75:25, v/v) to obtain a 500 ng/mL solution
—Dilute above solution 1:32 to obtain approximately a 100 fmol/μL solution
—Dissolve 1 mg of NaI powder in 1 mL of water:acetonitrile (50:50, v/v) to obtain a 1 mg/mL solution
—Weigh 0.1 mg of BSA and dissolve in 1 mL of 0.1% formic acid in water
720006614, August 2019