Antibody drug conjugates (ADCs) are a sub-class of biotherapeutics that consist of monoclonal antibodies (mAbs) and cytotoxic drugs linked to mAbs by chemical linkers. In this application note, lysine-conjugated ADCs were used to demonstrate the utilization of the Accurate Mass Screening workflow to monitor the relative abundance of ADC conjugated peptides across samples and to complete relative conjugation site occupancy ratio calculations automatically.
The Waters UNIFI Scientific Information System provides the Accurate Mass Screening workflow to streamline and facilitate the challenging quantitative analysis of antibody drug conjugate (ADC) peptides. This workflow utilizes the UNIFI scientific library as the means to share the peptide and peptide-drug conjugate target component information characterized by the Peptide Mapping workflow within the Biopharmaceutical Platform Solution. This component information is then used to create the target lists and Custom Field to automatically quantify these key attributes of ADCs.
Antibody drug conjugates (ADCs) are a sub-class of biotherapeutics that consist of monoclonal antibodies (mAbs) and cytotoxic drugs linked to mAbs by chemical linkers. ADCs are a complex mixture of conjugates that differ in the number of drugs attached as well as the location of the drug linkage. Previous studies demonstrated that the location, distribution, and site occupancy of the drug molecules play a critical role in the Complementarity Determining Region (CDR) binding efficiency, efficacy, toxicity, stability, and pharmacokinetic profile of ADCs. It is crucial to establish efficient and robust analytical capabilities to characterize the heterogeneity of drug-linker content and modification sites as well as to demonstrate lot-to-lot consistency, stability, and process comparability. Structural characterization, along with qualitative and quantitative analysis of ADCs, is challenging due to the heterogeneous nature of the conjugates and the low abundancies of individually modified species. The quantitation of conjugated peptides and determination of site occupancy ratio has traditionally been done by LC-UV methods, which is limited by the spectroscopic nature of the drug, low sensitivity, and insufficient selectivity. Compared to UV-based methods, MS-based methods can provide improved sensitivity and selectivity. However, MS-based quantitation has its own challenges that often involve monitoring a large number of MS peaks per sample for complex ADC peptides. Currently, efficient quantitation of ADC peptides and direct generation of the key attributes used for ADC comparability studies are impeded by a lack of sophisticated software tools. Waters Accurate Mass Screening (MSE) workflow within the Screening Platform Solution with UNIFI addresses these challenges and provides targeted component quantitation across samples, Custom Field for key ADC attributes (such as relative conjugation site occupancy ratio), and generation of reports in a streamlined and automated fashion.
|
System: |
ACQUITY UPLC H-Class Bio System |
|
Detector: |
ACQUITY UPLC Tunable UV (TUV) Detector |
|
Column: |
ACQUITY UPLC BEH300 C18 Column, 300 Å, 1.7 μm, 2.1 mm x 100 mm (P/N 186003686) |
|
Mass spectrometer: |
Xevo G2-XS QTof |
|
Acquisition mode: |
MSE |
|
UNIFI 1.8 Scientific Information System |
|
|---|---|
|
Analysis type: |
Accurate Mass Screening (MSE) workflow within the Screening Platform Solution with UNIFI, Peptide Mapping (MSE) workflow within the Biopharmaceutical Platform Solution. |
Antibody drug-conjugated samples were denatured, alkylated, and digested by Asp-N endoproteinase. Leucine enkephalin (LeuEnk) was added to each sample at a final concentration of 50 fmol/ul as an internal standard.
In this application note, lysine-conjugated ADCs were used to demonstrate the utilization of the Accurate Mass Screening workflow to monitor the relative abundance of ADC conjugated peptides across samples and to complete relative conjugation site occupancy ratio calculations automatically. The reported method is universal and can be applied to ADC therapeutics generated by different conjugation chemistries. Multiple features in UNIFI facilitate the quantitation analysis. First, the library of target components, including accurate mass values and optional retention times, is created. This was achieved by processing representative samples using the Peptide Mapping (MSE) analysis that identifies and labels the various peptide and peptide-drug conjugates. These are then selected and the relevant information entered into the UNIFI scientific library. The target components are therefore stored and easily retrievable to construct analysis methods. Custom Field enables the automated calculation of important product attributes of the ADCs. This is completed by using a Custom Field when setting up the processing method. In addition, UNIFI features a review panel that provides the ability to easily organize and save data review workflow steps, including user-configured plots and tables for consistent data display and straightforward communication. All of the relevant information including the use of result filters is available to the user and can be copied into reporting templates to visualize the results and generate analysis-specific reports.
LC-MSE peptide mapping data were acquired using the ACQUITY UPLC H-Class Bio System (with both UV and MS detection) and Xevo G2-XS QTof MS. MSE data acquisition enables sufficient points across the chromatographic elution peaks and provides MS (Low CE) and MS-MS (High CE) spectra in alternate scans. The data was processed using the Peptide Mapping MSE workflow. This software identifies the potential peptide and peptide-drug conjugate components through de-novo digestion of a target protein and searches the accurate mass MS and MS-MS data considering adducts and modifications, including the linker and drug. The identified components can then be saved into a user-defined UNIFI scientific library. The Peptide Mapping workflow presents identified components and quality attributes that indicate the relative confidence of the identification. User-defined filters can be created and saved such that these quality attributes can be applied to the components that will be stored within the library and used as the basis for the creation of the target list in the Accurate Mass Screening method.

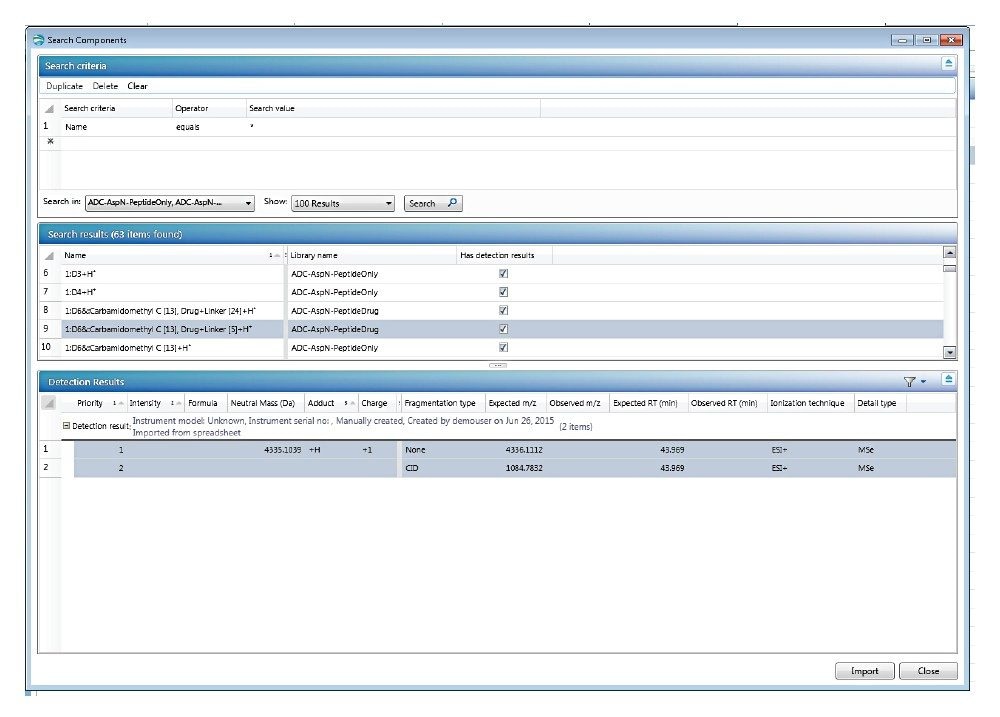
The UNIFI scientific library is a central library function within the software that contains commercial and user-created libraries. These libraries can be used to store component-centric chemical and physical information as well as detection results. This information can be used to facilitate the creation of analysis methods. Figure 2 shows an example of an ADC peptide library and detection results, including information such as the expected mass-to-charge ratio (m/z), expected retention time, and expected fragment ion m/z. Information can be imported through an excel spreadsheet, input into the library manually, or sent from the results of certain UNIFI analysis types.
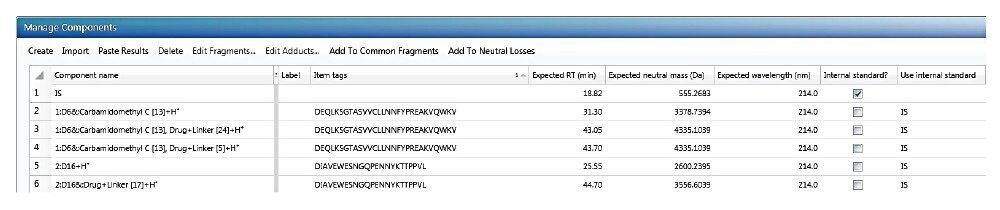
a. Manage Components table
First, the Manage Components table is created. Screening components can be imported directly from the scientific library. Figure 3 shows an example Manage Components table. Please note that in order to recognize the conjugated peptide with its unconjugated peptide counterpart, the peptide sequence must be included as an “item tag” for each component. This item tag is stored in the library along with the peptide sequence from the result fields in the Peptide Mapping MSE analysis. The internal standard can be added to the component table as shown in Figure 3. The expected retention time (RT) field is not required; however it can increase screening accuracy if the value is known from a previous analysis. The screening time window can be defined by the “target by retention time” setting in the analysis method. The expected neutral mass (Da) value for each component is required for the screening workflow. Mass-to-charge (m/z) values of different charge states or peptide adduct ions (m/z) are automatically calculated from the neutral mass during data processing.
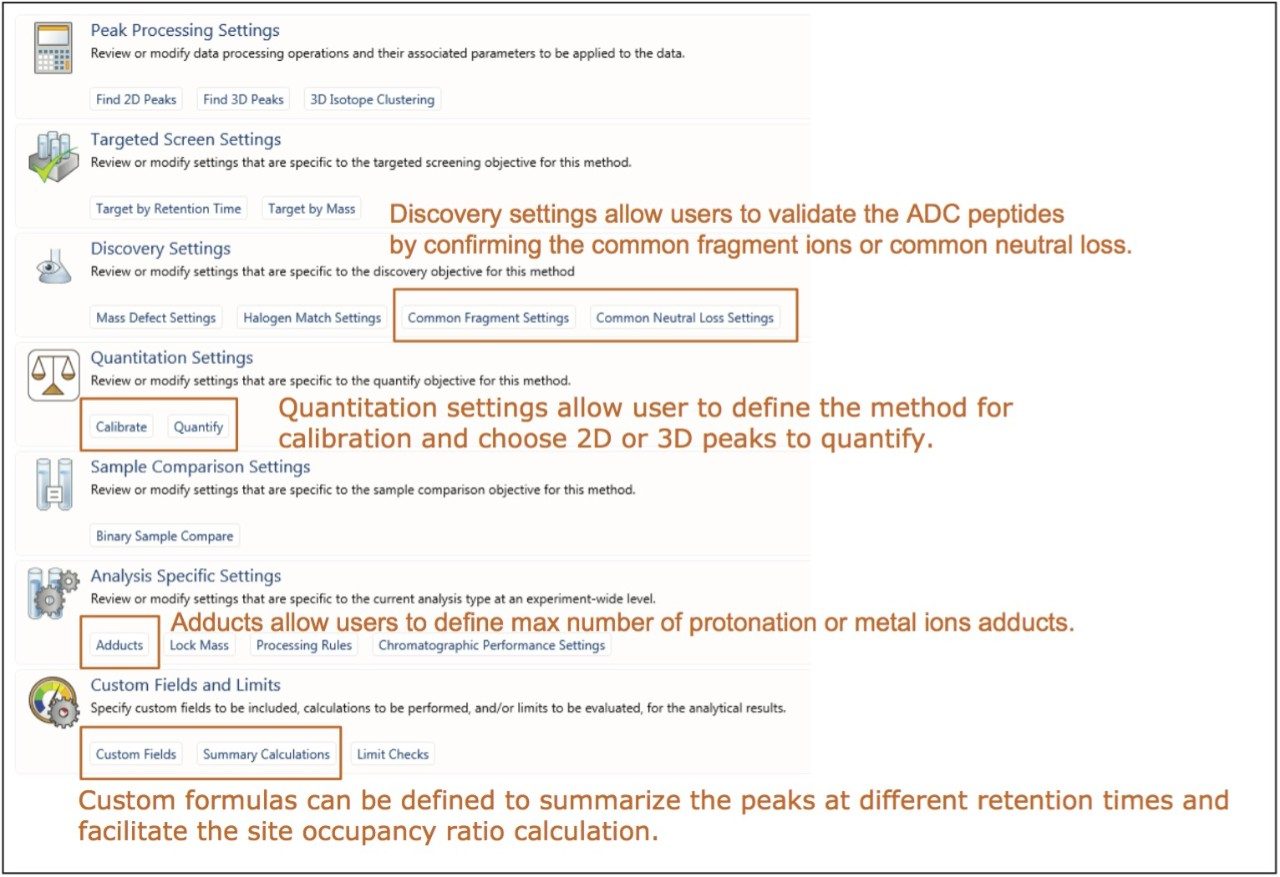
b. Processing parameters
Figure 4 shows the “processing sections” of the accurate mass screening workflow. The quantitative analysis of ADC peptides is facilitated through multiple features of UNIFI, some of which are highlighted and described in detail in Figure 4. The users have the flexibility to define the channel (UV or MS) or ions used for quantitation (most intense monoisotopic or sum of all adducts and isotopes). MS components can be quantified using the 2D peak (2D extracted ion chromatograms (XIC) area or height) or 3D peak volume. In addition, all the quantitation settings can be set on a per-component basis to allow flexibility in the application of different settings for separate target groups.
Custom Field is designed to perform the specific calculations for target analysis. The Custom Field can be stored in the analysis method with the results directly available for viewing, plotting, and reporting. Automated calculations within the analysis method eliminate the tedious extra steps for manual data manipulation and re-processing. Custom Field and analytical methods are protected by a secure framework via configurable user access controls based upon roles that facilitate their use in routine analysis, particularly in regulated laboratories.
Custom Field and the formulae to support the calculation of ADC attributes in a quantitative analysis are shown in Table 1. The Custom Field is formulated generically so that it can be applied to other types of ADC peptide analyses. The text marker “drug” is used to identify the conjugated peptide components in the screening results. As long as the component name of the conjugated peptides contains “drug” in the Manage Components table, the Custom Field will find these components and apply the determined values. “Calculate amount” is a standard field generated by UNIFI processing that is obtained by normalizing the intensity of each identified quantitative response against the spiked-in internal standard response. The peptide sequence values are included in the “item tag” field, so that the peptide components with the same sequence can be summed using the “Peptide Tot AMT” Custom Field. Using these four Custom Fields, users can effectively monitor the amount of conjugated peptides and relative site occupancy ratios across sample sets.
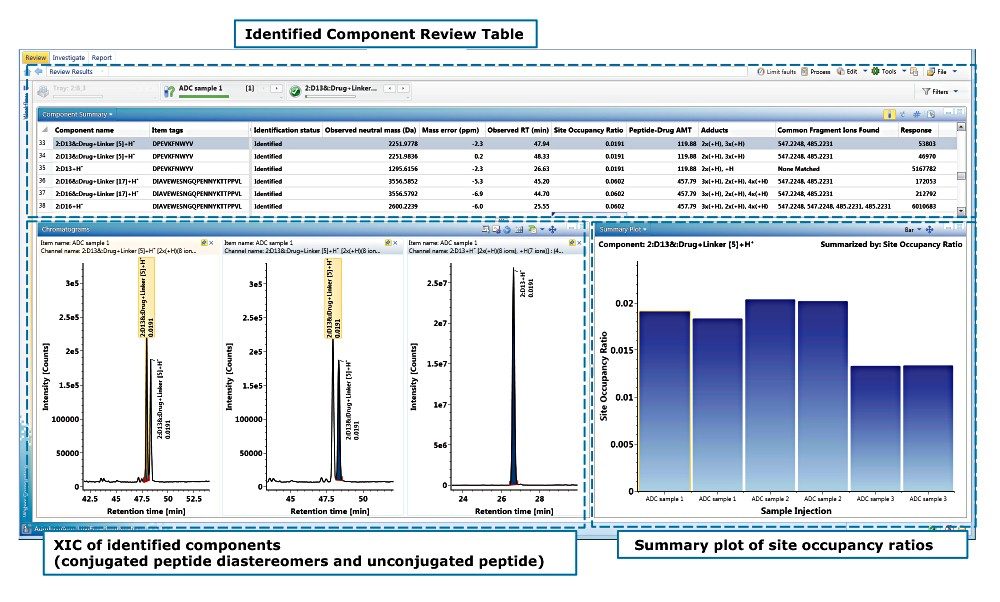
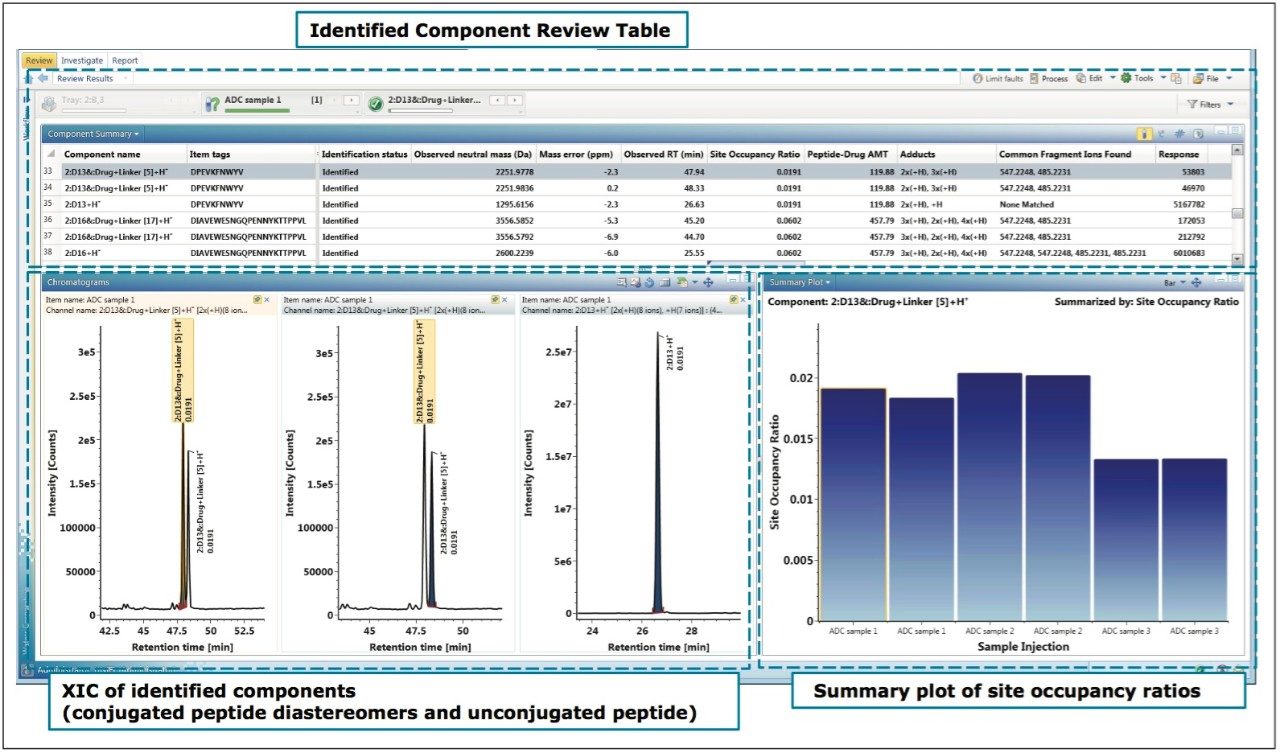
The detection and calculation results are displayed in the UNIFI review tab as shown in Figure 5. It consists of three areas:
This application note describes how to perform ADC peptide quantitation using the Accurate Mass Screening workflow in UNIFI and further demonstrates the capability of comparing the key product attributes of ADCs across samples through this automated workflow. This eliminates tedious and labor-intensive manual data processing, which reduces associated human errors. Furthermore, the defined workflow in UNIFI can be saved and transferred to promote high quality data generation, consistent processing, and reporting for complex ADC samples. Combined with the UNIFI Peptide Mapping MSE workflow, this will enable scientists to overcome the analytical challenges for both structural characterization as well as qualitative and quantitative analysis of complex ADCs. Collectively the Accurate Mass Screening workflow (quantitative tool) combined with UNIFI scientific library and Custom Field is a great addition to the traditional Peptide Mapping MSE workflow (qualitative tool) within the same UNIFI architecture to facilitate the challenges of ADC biotherapeutics research and development.
720005603, February 2016