For research use only. Not for use in diagnostic procedures.
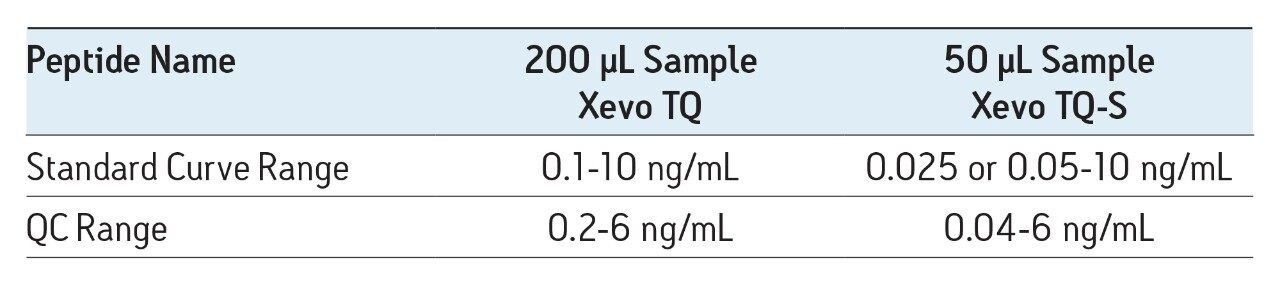
In this work, the mass spectrometry platform has been updated from the Xevo TQ MS to the Xevo TQ-S mass spectrometry system. This change facilitated both a 4X reduction in required sample size and a 4-5X increase in assay sensitivity.
A previous application note (720003682en) described in detail the development of a fast, flexible SPE-LC-MS/MS platform for the quantification of multiple amyloid beta (aβ) peptides from human or monkey CSF for use in a biomarker or preclinical discovery setting. In this work, the mass spectrometry platform has been updated from the Xevo TQ MS to the Xevo TQ-S mass spectrometry system. This change facilitated both a 4X reduction in required sample size and a 4-5X increase in assay sensitivity.
Historically, quantification of aβ peptides in biological fluids has relied mainly on the use of immunoassays, such as ELISA. These assays are time consuming and expensive to develop, labor intensive, are subject to cross reactivity, and an individual assay is required for each peptide. In order to meet the throughput requirements and constant flow of demands for new peptide methods in a discovery setting, there is a need for a highly specific yet flexible methodology based on an LC-MS/MS platform. In this work, this platform is coupled with selective sample preparation for the simultaneous quan-titation of multiple aβ peptides. This work focuses on methods for the 1-38, 1-40, and 1-42 aβ peptides, in support of preclinical and biomarker discovery studies. Sequence, pI and molecular weight (MW) information for these peptides is shown in Figure 1.

The solid-phase extraction (SPE) sample preparation protocol used to enrich the amyloid beta fraction in CSF is the protocol previously described. However, the sample size required is now only 50 µL instead of 200 µL. The SPE method concentrates the sample to improve detection limits while eliminating matrix interferences and optimizing solubility of the aβ peptides in the mass spectrometer injection solvent.
As strategies emerge for disease modification in Alzheimer’s Disease (AD), the quanti-fication of other aβ species (in addition to aβ 38, 40, and 42) that may be linked to AD pathology may be required. The method described herein shows promise for adaptation to quantify those peptides as well.
|
Column: |
ACQUITY UPLC BEH C18 300Å, 2.1 x 150 mm, 1.7 μm, Peptide Separation Technology |
|
Part Number: |
186003687 |
|
Column temp.: |
50 °C |
|
Sample temp.: |
15 °C |
|
Injection volume: |
10.0 μL |
|
Injection mode: |
Partial Loop |
|
Flow rate: |
0.2 mL/min. |
|
Mobile phase A: |
0.3% NH4OH in H2O |
|
Mobile phase B: |
90/10 ACN/mobile phase A |
|
Strong needle wash: |
60:40 ACN:IPA + 10% conc. NH4OH (600 μL) |
|
Weak needle wash: |
90:10 0.3% NH4OH in H2O:ACN (400 μL) |
|
Time (min) |
%A |
%B |
Curve |
|---|---|---|---|
|
0.0 |
90 |
10 |
6 |
|
1.0 |
90 |
10 |
6 |
|
6.5 |
55 |
45 |
6 |
|
6.7 |
55 |
45 |
6 |
|
70 |
90 |
10 |
6 |
|
Capillary Voltage: |
2.5 V |
|
Desolvation Temp: |
450 °C |
|
Cone Gas Flow: |
Not used |
|
Desolvation Gas Flow: |
800 L/Hr |
|
Collision Cell Pressure: |
2.6 x 10(-3) mbar |
|
MRM transition monitored, ESI+: |
See Table 1 |

50 μL human CSF or spiked artificial CSF + 5% rat plasma was diluted 1:1 with 5 M guanidine HCL and shaken at room temperature for 45 minutes. This was then diluted further with 50 μL 4% H3PO4 in H2O and mixed.
Note: For spiked samples, samples were allowed to equilibrate at room temperature for 30 min after spiking and prior to dilution with guanidine HCl.
Samples were extracted according to the protocol in Figure 2 below. All solutions are made up by volume. All steps applied to wells of μElution plate containing samples

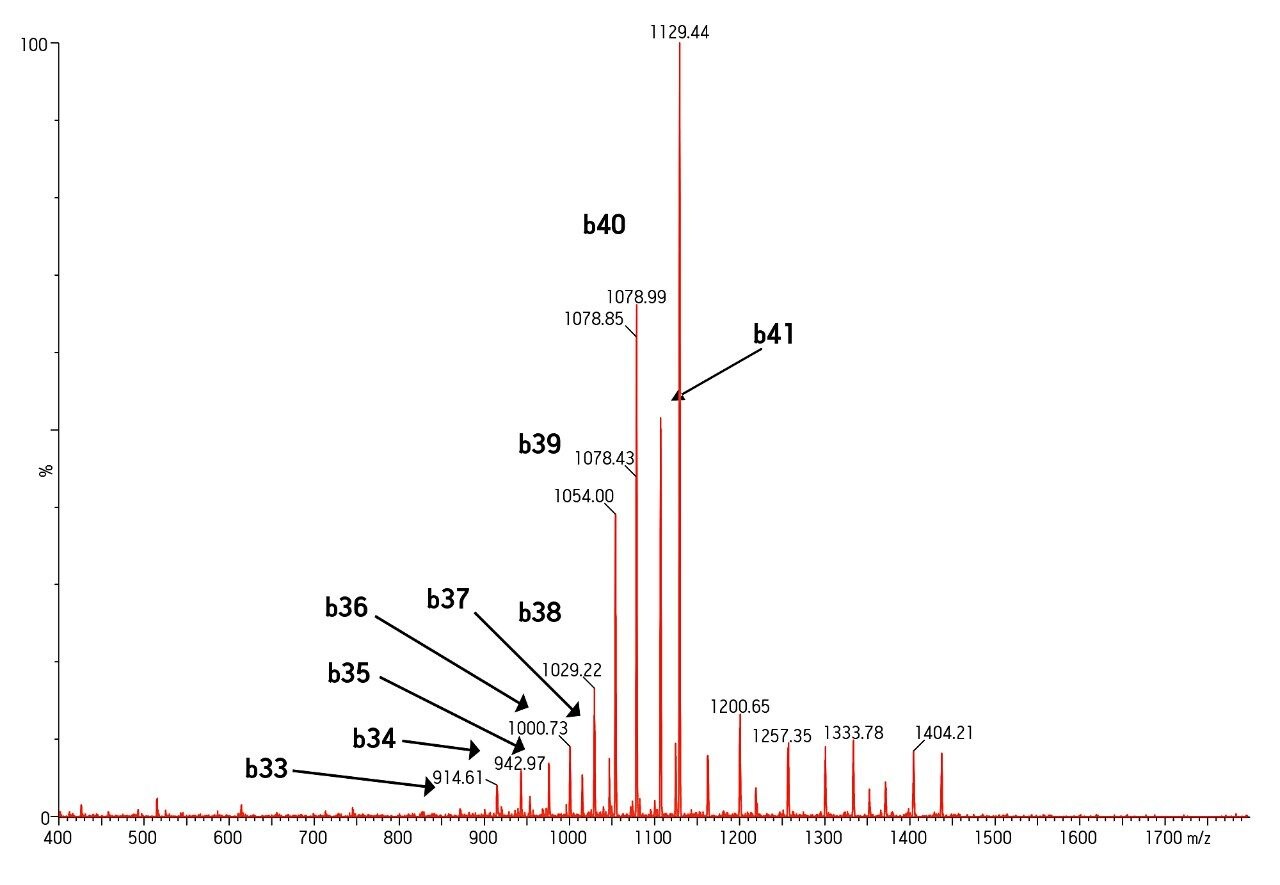
MS was performed in positive ion mode since CID of the 4+ precursor ion yielded several distinct product ions corresponding to specific b sequence ions (representative spectrum shown in Figure 3.)
The sensitivity increase provided by the Xevo TQ-S facilitated the use of 4X less sample whilst also improving detection limits by approximately 4-5X compared to the previous assay on the Xevo TQ. The lowest QC sample tested was 5X lower in concentration than the low QC when the standard Xevo TQ MS system was used. Earlier work by Rainville and Booth (application note 720003415en) describes the system improvement in more depth and demonstrates a similar sensitivity increase for the therapeutic peptide desmopressin.
Mass range of the instrument was also an important factor in obtaining specificity. The Xevo TQ-S MS has a mass range of 2048 on both quads, easily allowing us to choose a more specific 4+ rather than 5+ precursor and fragment pair.
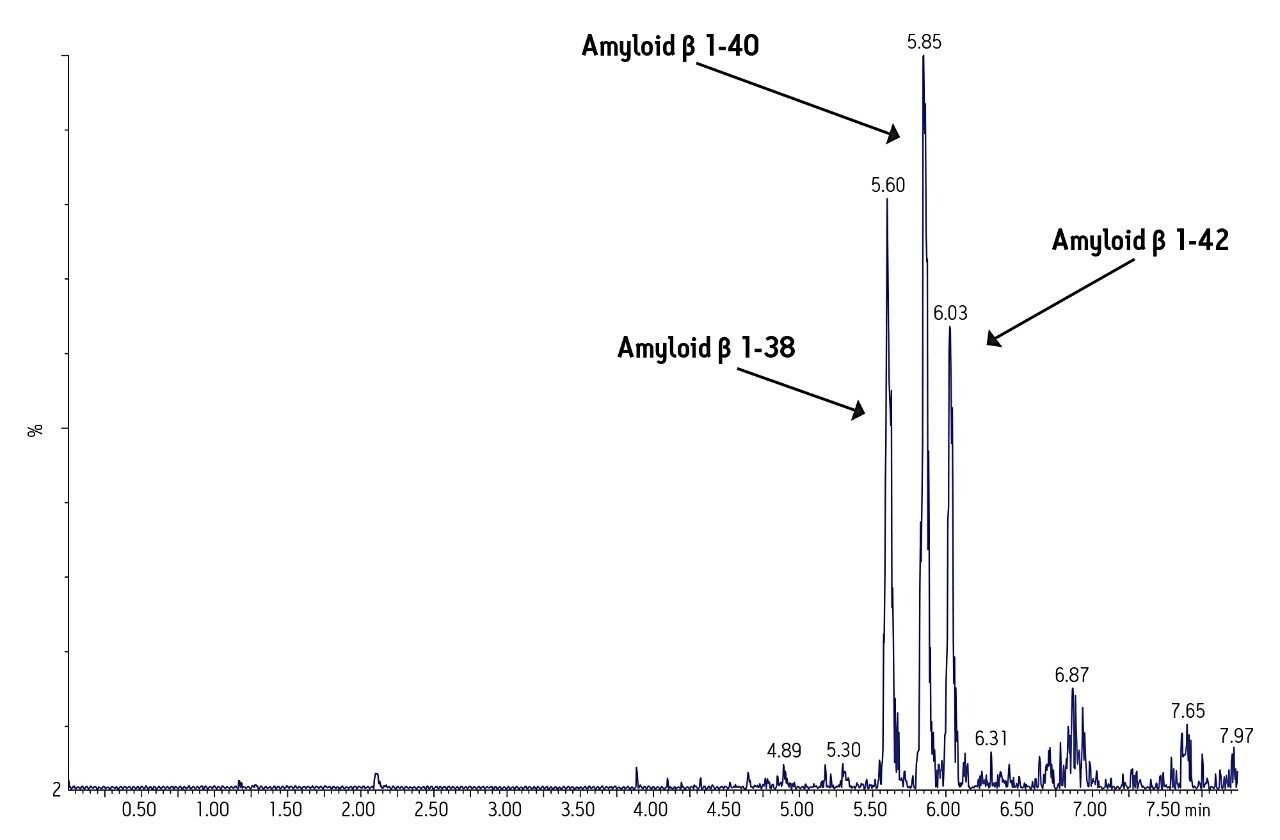
Separation of the three amyloid β peptides is shown in Figure 4. While the exact amount of NH4OH in the mobile phase was critical for negative ion sensitivity, the signal in ESI positive proved to be more robust to subtle changes in mobile phase composition, providing a minimum of >24 hour LC/autosampler stability. In contrast, 50% or more of the ESI negative signal was lost after 10-12 hours due to the natural change in NH4OH concentration (volatility) in the mobile phase. This further reinforced the robustness of an ESI positive MS method.
SPE was performed using Oasis MCX, a mixed-mode sorbent, to enhance selectivity of the extraction. The sorbent relies on both reversed-phase and ion-exchange retention mechanisms to selectively separate the aβ fraction from other high abundance polypeptides in complex CSF samples. The Oasis µElution plate (96-well format) provided sample concentration, eliminating the need for evaporation and reconstitution. This has the benefit of saving time and eliminating peptide losses due to adsorption to the walls of the collection plate during dry down.
During initial method development, a high degree of non-specific binding (NSB) was observed when artificial CSF was extracted. Thus, 5% rat plasma (having a different amyloid β sequence) was added to bind to surfaces, eliminating NSB.
The SPE method was one of the more critical aspects of the overall methodology. Very selective isolation of the amyloid fraction coupled with the resolution of analytical-scale flow UPLC, facilitates analysis of pre-clinical samples without the need for antibodies or time-consuming immuno-precipitation associated with ELISA methods. The increased sensitivity of the Xevo TQ-S enabled the sample volume to be reduced from 200 µL to 50 µL of CSF, making this method amenable to use in pre-clinical species.
720003860, February 2011