This is an Application Brief and does not contain a detailed Experimental section.
This application brief demonstrates how 2D technology can be used to couple incompatible separation and detection methods by interfacing an IEX method using a salt gradient with RPLC-MS.
Multiple heart-cut 2D-LC enables IEX-RPLC-MS analysis of monoclonal antibody charge variants using compliant informatics.
Many biopharmaceutical separations require the use of mobile phases containing salts or other components that are not compatible with MS. Often these separations rely on fraction collection and offline desalting prior to MS characterization. Ion exchange chromatography (IEX), which is commonly used to analyze charge heterogeneity, is an example of an assay that is not directly compatible with MS when a traditional salt gradient is used for elution. A Waters ACQUITY UPLC System with 2D technology offers a solution for coupling incompatible methods and providing an interface for MS analysis.
In this study, a multiple heart-cut approach is used to selectively transfer monoclonal antibody charge variants separated by a salt gradient in the first-dimension to a second dimension column for intact mass analysis. By incorporating 2D technology, a traditional IEX method using a salt gradient can be coupled to MS, which eliminates the need for offline fractionation and additional sample handling. Formulated infliximab at 20 mg/mL is used as a model protein to demonstrate mass confirmation of major lysine variants.
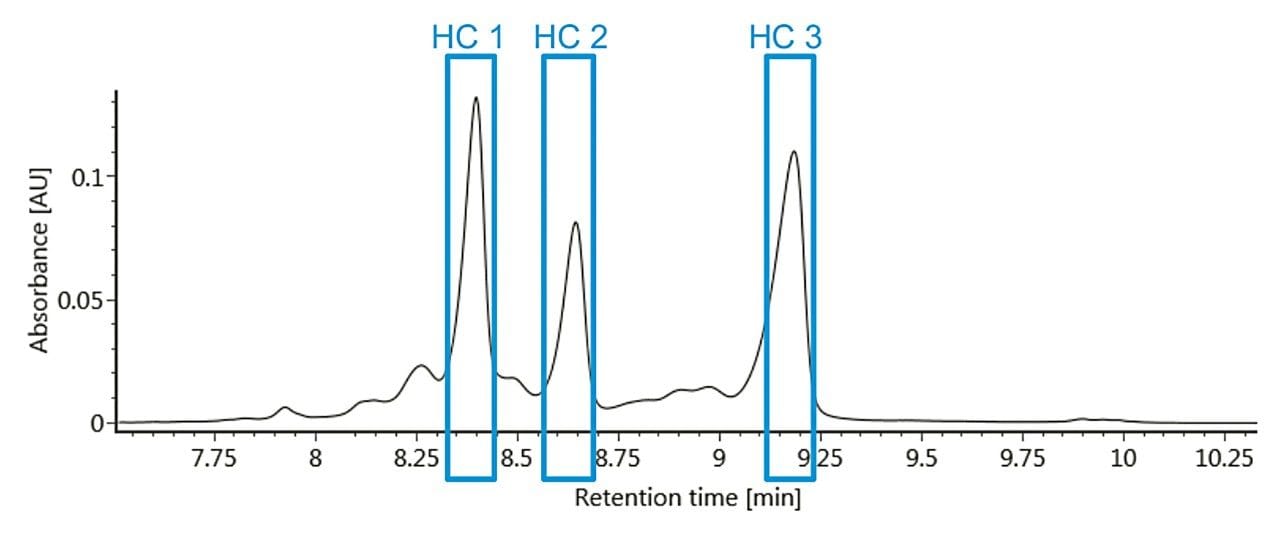
Auto•Blend Plus technology was used to develop a gradient method using an MES buffer system and a BioResolve SCX mAb Column (3 μm, 4.6 x 50 mm).¹ By preparing mobile phase stock solutions, pH and ionic strength were rapidly screened to determine optimal separation conditions (pH 6.60, 10–55 mM salt over five minutes). It is of critical importance to develop a method with good retention time repeatability so that the desired fractions can be transferred reproducibly to the second dimension for analysis. A representative chromatogram of the first-dimension separation can be used to determine the heart-cut window for each of the lysine variants to be transferred to the seconddimension for subsequent analysis (Figure 1).
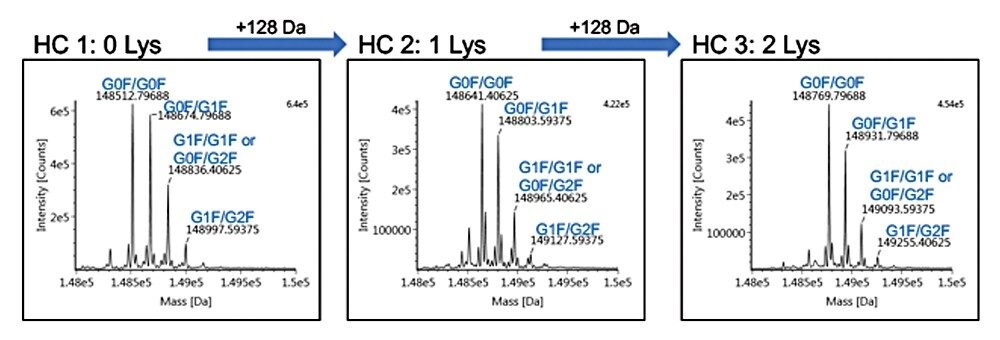
After establishing retention time repeatability and determining the elution window for each of the lysine variants, a multiple heart-cut method was created where a single injection was used to selectively transfer infliximab variants to individual trap columns. After incorporating a five-minute wash step (reversed-phase starting conditions) for each of the trap columns, an ACQUITY UPLC Protein BEH C4 Column (300 Å, 1.7 μm, 2.1 x 50 mm) was used in the second-dimension for intact mass analysis. Deconvoluted data with assigned peaks are shown in Figure 2 for each lysine variant.
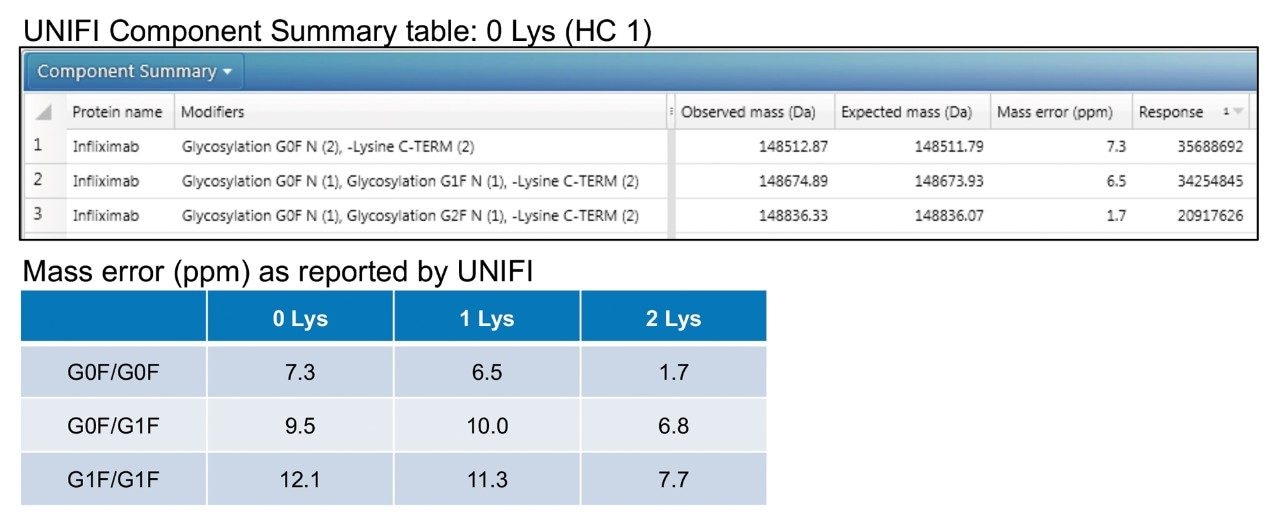
The intact protein workflow within UNIFI Software was used to process data according to a defined MS-RT window. By providing sequence information for infliximab, a component table summarizing results from each heart-cut was generated (Figure 3). This information was then collated to report mass error for the three most abundant glycoforms for each of the lysine variants (Figure 3). The calculated mass error is ~10 ppm for all species, which agrees with results historically reported for the instrumentation used.
By incorporating 2D technology into biopharmaceutical workflows, traditional one dimensional methods that do not use MS-friendly mobile phase can be interfaced with MS, bypassing the need for fractionation and offline sample preparation. A multiple heartcut IEX-RPLC-MS assay was used to analyze infliximab lysine variants by coupling an IEX method with a salt gradient in the first-dimension to a RPLC-MS method in the second-dimension. This same multiple heart-cut approach can be extended to additional biopharmaceutical applications that require multiple peaks to be transferred to the second dimension. 2D-LC-MS, in combination with system capabilities for compliant data collection, processing, and reporting, offers improved productivity and added confidence in results for biopharmaceutical laboratories working in development and manufacturing.
720006755, February 2020