This application note demonstrates how LC-HRMS coupled with UNIFI Scientific Information System can be used for qualitative and quantitative analysis of synthetic peptide impurities, from confirmation of impurities to relative % abundance calculation in a compliance-ready environment.
Peptide-based therapeutics are alpha amino acid polymers containing less than one hundred amino acids,1 represent a new class of highly evolving drugs in the market generating billions of dollars of revenue each year.2 These peptide drugs exhibit relatively low toxicity, high biological activity, and potential applications for many medical challenges compared to the most conventional drug products.3-4 A majority of these therapeutic peptides are produced by chemical synthesis, however, they can be also rDNA-derived or natural source-based.5 Each production method is equally challenged by impurities; for synthetic peptides, these can be processrelated or product-related that are likely to be biologically active.6 With the emergence of Reference Listed Drugs (RLD) versus generic peptide drugs, strict regulatory measures became a necessity for synthetic peptide drugs to ensure their efficacy and safety. The International Council for Harmonization (ICH) has quality guidelines applicable to impurity identification and specification and these have been adopted by regulatory agencies in the US, Europe, and Japan.6-9 Enforcing these quality guidelines in design, development and production processes of synthetic peptides has presented an opportunity for more sensitive analysis methods to be a part of the analytical toolbox.
The conventional synthetic peptide impurity profiling methods are mostly LC-optical-based assays that rely on chromatographic separation of impurities followed by molecular weight confirmation of each peak.10 Recent studies have shown that incorporating high-resolution mass spectrometry (HRMS) into the analytical workflow can obtain accurate mass-based identification of peptide API and impurities which also benefit from availability of fragment ions for peptide sequence verification.11 The most challenging task in an HRMS workflow is data interpretation to confirm and assign the impurities. In this application note, we introduce the UNIFI Scientific Information System with compliance-ready workflows for this purpose.
Eledoisin, a vasodilator, is a synthetic therapeutic peptide with 11 amino acid residues used as a case study to demonstrate LC-UV-HRMS workflow with UNIFI Scientific Information System. The compliance-ready UNIFI workflows enable data acquisition, data processing to confirm impurities, and calculate the % relative amount, and reporting. In addition, UNIFI Software has custom library features to archive API and impurities. Once the library is built, it can be used for high throughput targeted impurity profiling for the same drug product. The overall strategy used for eledoisin impurity profiling can be applied for other peptide therapeutics during the production process and is ideal for regulated laboratory environments.
The synthetic peptide Eledoisin (pEPSKDAFIGLM-NH2) was purchased from New England Peptides Ins (Gardner, MS) and calcitonin (salmon) (CSNLTCVLGKLSQELHKLQTYPRTNTGSGTP-NH2) was purchased from Bachem (Torrance, CA). A 2 μg/μL stock solution in water was further diluted to a final concentration of 0.2 μg/μL for analysis.
|
LC system: |
ACQUITY UPLC H-Class Bio |
|
Detector: |
ACQUITY UPLC Tunable Ultraviolet (TUV) |
|
Wave length: |
214 nm |
|
Column: |
ACQUITY UPLC Peptide CSH C18, 130 Å, 1.7 μm, 2.1 mm × 100 mm |
|
Column temp.: |
65 °C |
|
Mobile phase A: |
0.1% Formic acid in H2O |
|
Mobile phase B: |
0.1% Formic acid in acetonitrile |
|
Gradient: |
Eledoisin: 16–24% B over 30 min, |
|
Calcitonin: |
14–34% B over 20 min |
|
MS instrumentation: |
Vion IMS QTof Mass Spectrometer |
|
Capillary voltage: |
2.8 kV |
|
Cone voltage: |
50 °C |
|
Source offset: |
50 °C |
|
Source temp.: |
80 °C |
|
Desolvation temp.: |
300 °C |
|
Cone gas: |
20 L/hr |
|
Desolvation gas: |
500 L/hr |
UNIFI 1.9.2 Scientific Information System
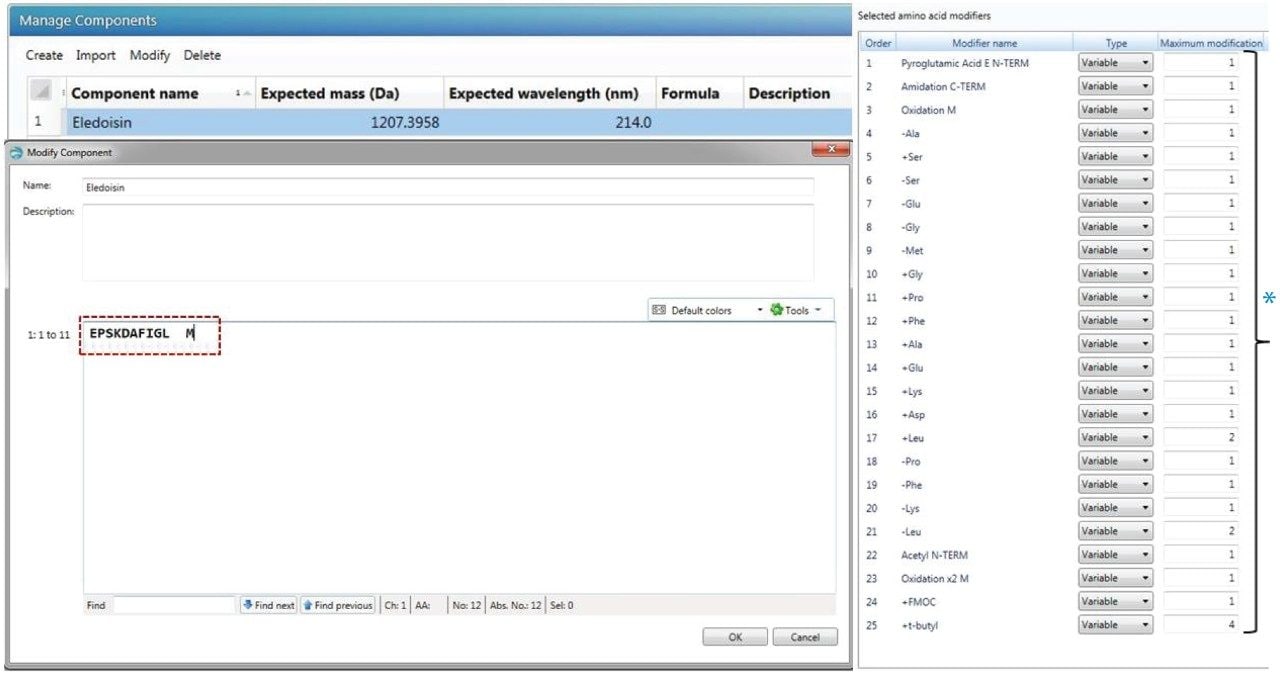
Here, we demonstrate how UNIFI Scientific Information System, a compliance-ready software, comprised of built-in workflow methods with data acquisition, processing, and reporting can be utilized in synthetic peptide characterization and impurity profiling. This work has been done in parallel to our previous study using MassLynx and ProMass Software.12 For UNIFI-based data processing, the primary amino acid sequence of the synthetic peptide can be imported from the scientific library or created by the user. The challenge is to identify amino acid modifications causing the impurities. The UNIFI modification database is comprised of custom chemical modifications that can be easily incorporated into the workflow method for locating the impurities based on MS and MS/MS profiles. Figure 1 (*) shows the modifiers of synthetic impurities used in Eledoisin impurity analysis such as: pyroglutamic acid modification (Figure 1, line 1), insertion and deletion of amino acids (line 4–21), addition of Fmoc, and t-butyl groups due to incomplete deprotection (line 24, 25), and peptide degradation-related products (line 3,23).
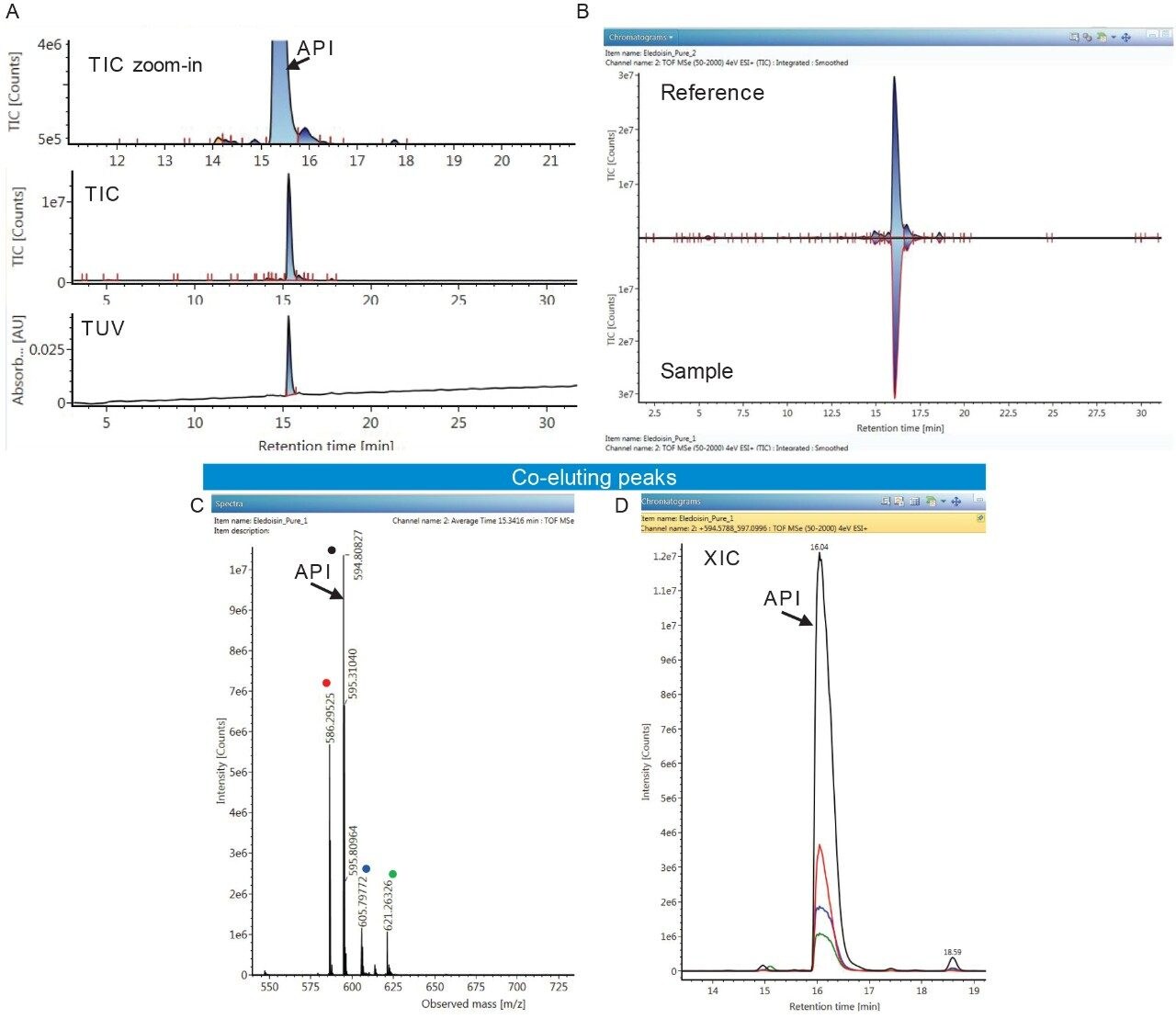
The goal of the analysis is to confirm the API sequence (e.g. Eledoisin) and identify major impurities. In brief, the peaks were separated using a 16–24% acetonitrile in 0.1% formic acid gradient over 30 minutes using an ACQUITY UPLC CSH C18 Column. Even at optimal chromatographic performance, obtaining baseline resolved peaks for optical detection of low abundance impurities is a challenge. Adding HRMS to the analysis resolves this by improving the detection and identification of low level impurities based on the accurate mass and MS/MS information. The TUV (214 nm) in Figure 1A indicates that only a few low abundance peaks are present in the sample. The comparison of sample chromatogram to the reference using UNIFI mirror plot feature in Figure 1B further confirms this observation. However, the summed mass spectrum for the API peak contains multiple monoisotopic m/z’s. The most abundant co-eluting ions in Figure 1C are +2 charged and based on the Eledoisin sequence the MS spectrum confirms that only m/z 594.8083 belongs to the API. The extracted ion chromatograms (XIC) of selected impurity peaks co-eluting with Eledoisin API are given in Figure 1D.
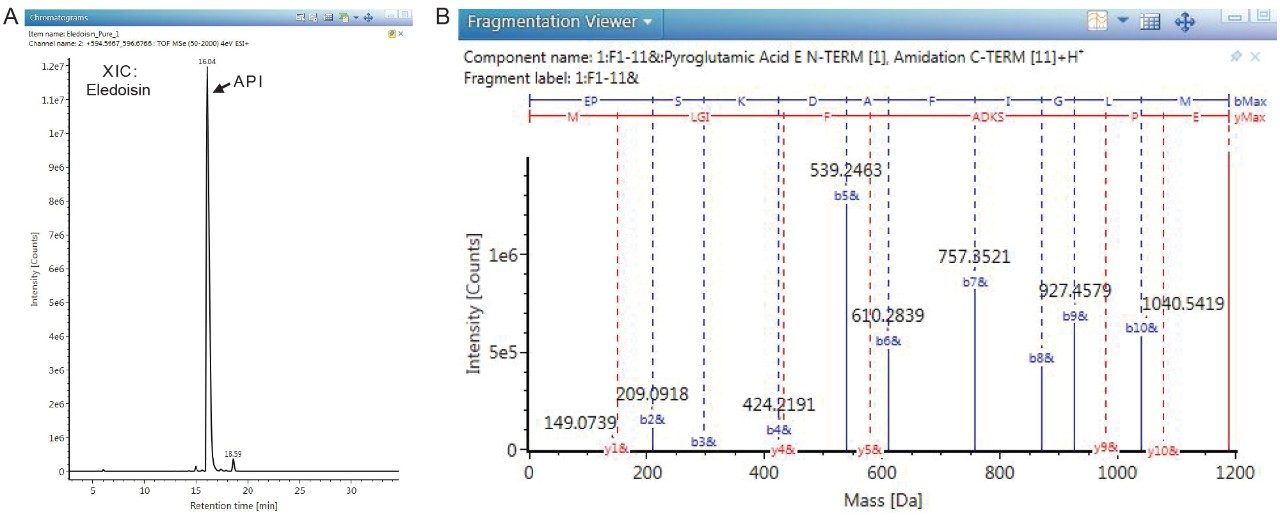
The deconvoluted mass spectra were used to identify these co-eluting impurities. The UNIFI peptide mapping workflow identified these peaks to be impurities containing insertions and deletions of amino acids, oxidation, and amino acid protection groups. The MS/MS fragment ions generated using data independent acquisition (MSE) was used to further confirm the sequence of the Eledoisin API and impurities. Below is the XIC (Figure 3A) and MS/MS spectrum for Eledoisin API peak confirming the amino acid sequence and respective modifications of its most abundant form: N-terminal pyroglutamic modification and C-terminal amidation (Figure 3B). The key features in the UNIFI workflow include automated data processing, chromatographic peak and fragment ion annotation, and customizable scientific library. The components of interest identified in the peptide mapping workflow can be imported into a dedicated custom impurity library to be used in high throughput analysis later in the production process using the UNIFI screening workflow.
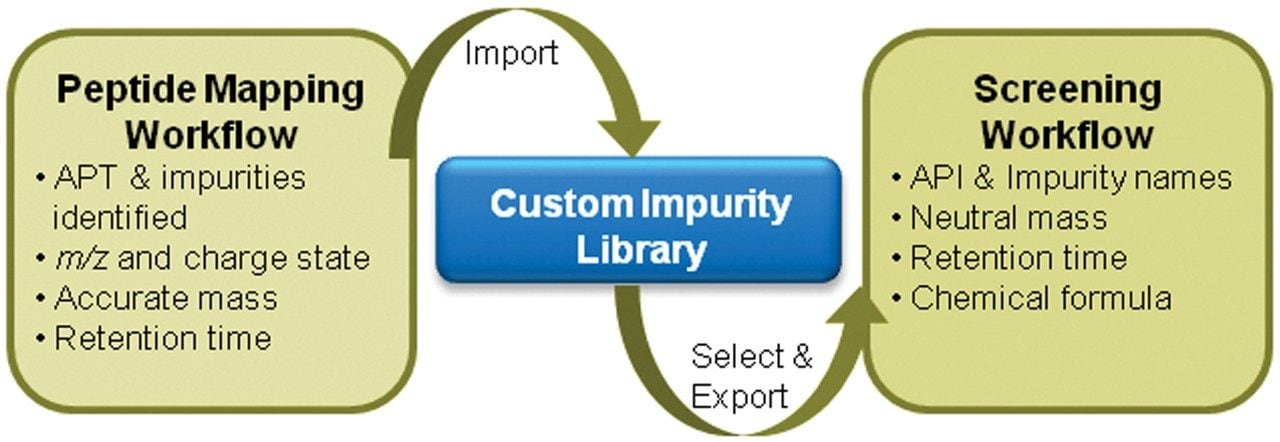
Not all impurities are biologically active or harmful; however, identifying undesirable synthetic byproducts is critical in maintaining the quality of the final therapeutic product. Profiling impurities at drug development and production using a streamlined workflow can significantly reduce impurity levels and analysis time spent on assessing the product at the end. The key feature is being able to use the custom impurity library to facilitate impurity screening. Impurities identified in the characterization stage by peptide mapping workflow can be added to the library or created as a new entry based on prior knowledge. Those imported from the peptide mapping data processing workflow contains information about the m/z, charge state, retention time, neutral mass, formula, and fragment ion spectra associated with each component peak. When batch selected, these components can be retrieved and imported back into the screening workflow with their formula, neutral mass, and retention time information for impurity profiling. The schematic in Figure 4 displays this process.
Using Eledoisin as an example, we demonstrate the use of screening workflow to profile impurities related to oxidation, deletion of Gly/Ala/Pro/Met, and insertion of Lys/Ser/Fmoc. It is noteworthy that these modifications were imported from the impurity library built for Eledoisin.
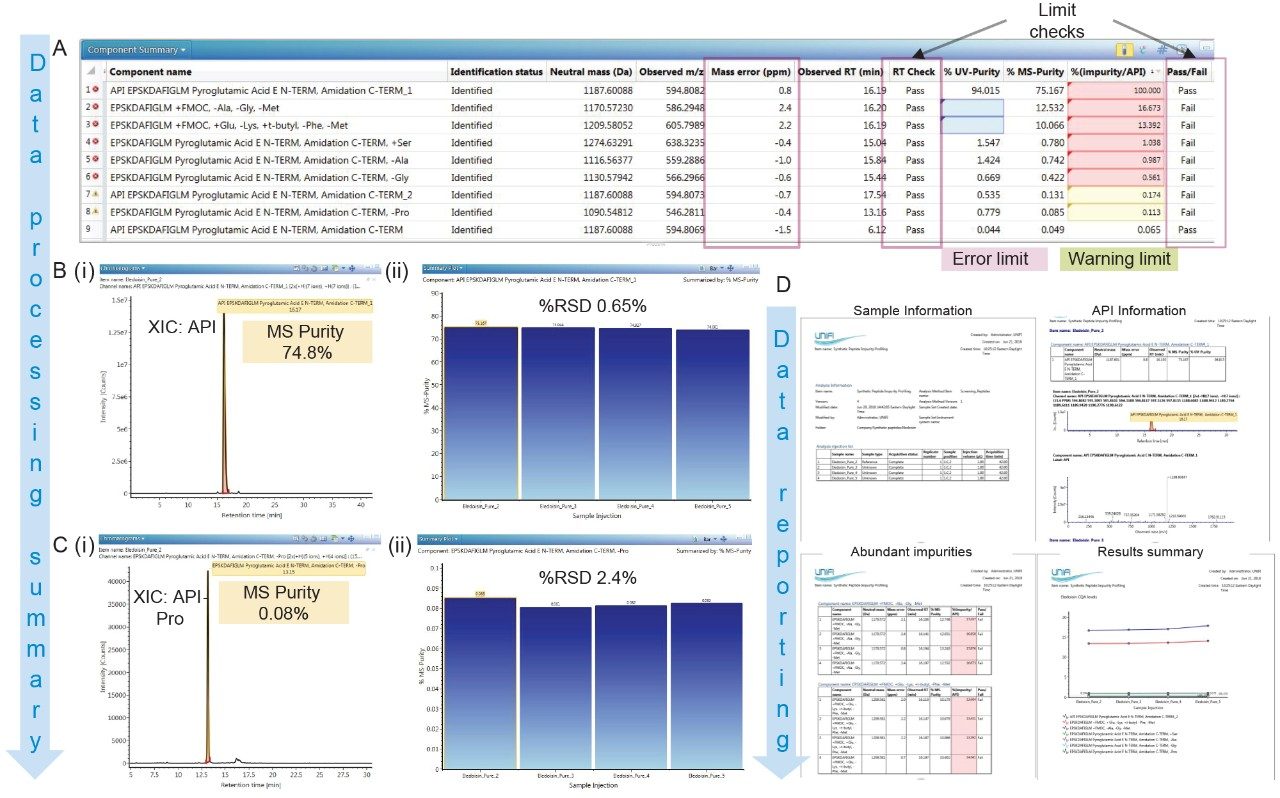
In addition to detecting these impurities in a large batch of synthetic Eledoisin samples, the screening workflow illustrated here was used to obtain the %purity of each peptide based on UV and MS responses and the %relative abundance to Eledoisin API MS response. The purity levels determined from MS and UV response are different due to co-eluting peptides only detected in the MS-based purity assessment. The reported optical purity for Eledoisin API was 94.7% while MS was 74.8% (the FDA drafted guidelines for synthetic peptides state that any impurity with 0.1% abundance13 or higher relative to the API should be identified; the sum of impurity levels of an eligible drug product must not exceed 0.5%13). By modifying the limit check parameters the users are able to deploy thresholds for potentially harmful impurities in the sample. Similarly, an example is given in Figure 5A, where %(impurity/API) levels are highlighted in red or yellow based on whether they exceed error limit (maximum threshold) or warning levels respectively. The user-defined error limit used in here is 0.5%. The pass/fail limit checks for % (impurity/API) and retention time are given by using a simple Boolean type custom formula in the method. In addition, the workflow provides sample statistics validating the quality of the analytical method and instrumentation. The examples given in Figure 5B and C show the XIC and %RSD for Eledoisin API and a selected low abundance impurity, API-Proline. Even at 0.08% level API-Pro has a %RSD as low as 2.4% (Figure 5C i, ii).
A report has been generated as shown in Figure 5D following the automated processing using a template to summarize the final results. The built in automated reporting capability in UNIFI allows the users to create a report based on an existing, modified or a user-defined template. Moreover, the data can be filtered and copied from the analysis method directly on to the template.
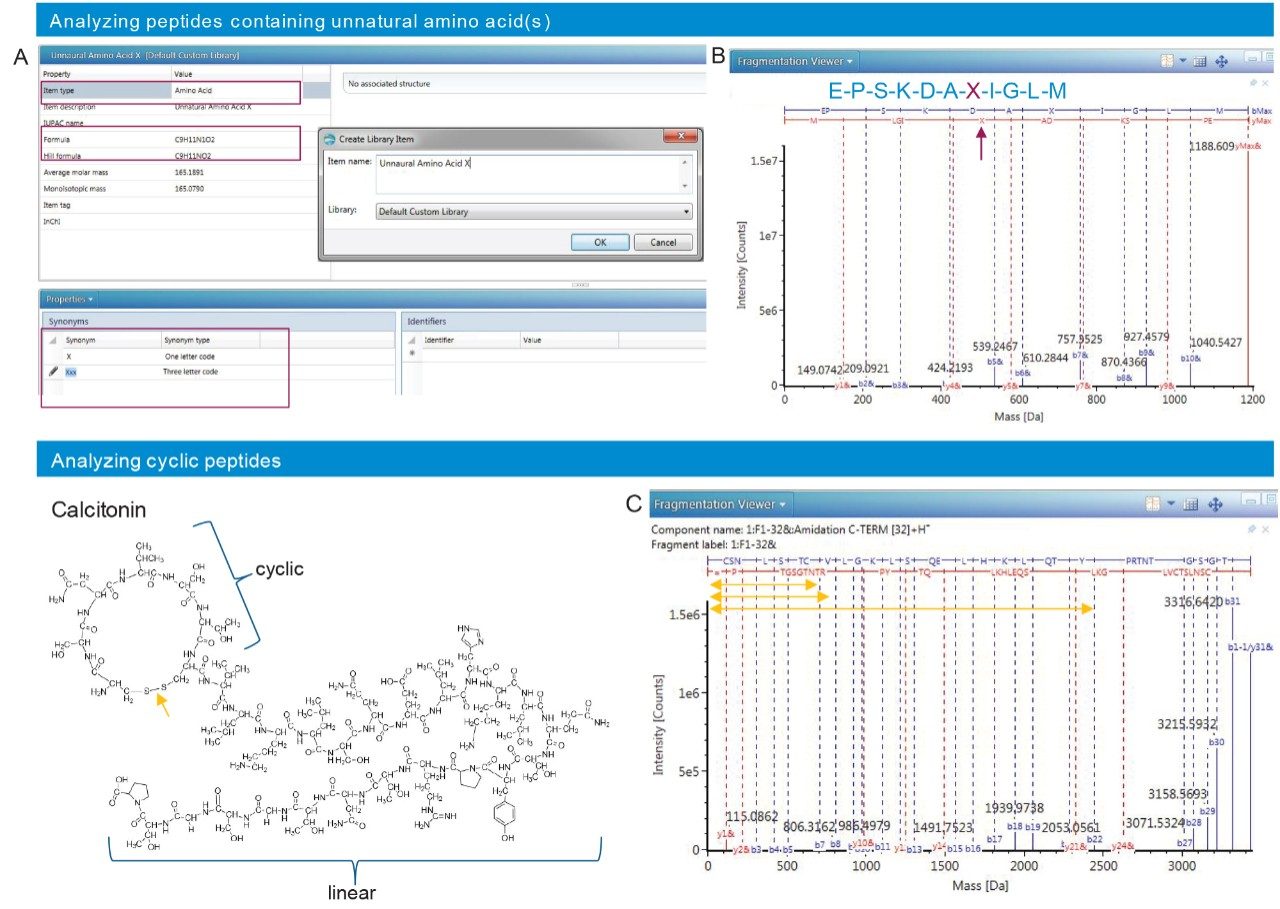
Many synthetic peptides contain unnatural amino acid(s) or have a cyclic/semi-cyclic structure. Data acquired for these types of samples are challenging to annotate since most informatics platforms are designed for analyzing natural or recombinant proteins and peptides. To demonstrate UNIFI-based data analyses in the above scenarios, we present the following examples: generating unnatural amino acids in UNIFI for data analysis (Figure 6A), identifying the fragments ions of a linear peptide containing an amino acid “X” with a formula C9H11O2N (Figure 6B) and fragmentation spectrum of semi-cyclic peptide Calcitonin that has a disulfide-bonded cyclic structure at its N-terminus (Figure 6C). These examples demonstrate the flexibility of data processing with UNIFI, showcasing that it can be used in impurity identification for a variety of synthetic peptides.
This application note demonstrates how LC-HRMS coupled with UNIFI Scientific Information System can be used for qualitative and quantitative analysis of synthetic peptide impurities, from confirmation of impurities to relative % abundance calculation in a compliance-ready environment. With automated data processing, impurity archiving and retrieving, custom calculations, limit check capability, and result statistics, this platform provides an ideal compliance-ready solution for profiling and reporting impurities in accordance with the ICH and FDA regulatory guidelines13 for peptide therapeutics.
720006367, August 2018