This application note demonstrates the application of high-resolution mass spectrometry to generate exact mass MS/MS fragment analysis data in conjunction with the use of software to automatically identify product ion fragments.
Impurity profiling is a critical part of the drug development process. Structural elucidation of unknown substances such as synthetic impurities is a key factor to refining pharmaceutical drug potency and safety attributes.1 Success in solving these very complicated and challenging structural puzzles has been facilitated by evolutions in LC and MS instrumentation; however it still requires a high level of experience to solve these analytical problems.
Typically MS/MS data from a triple quadrupole mass spectrometer is interrogated for structural elucidation in impurity profiling, in order to provide fragmentation spectra. However, using MS/MS analysis alone – particularly nominal mass spectra – may not provide sufficient specificity to determine the analyte structure. Complementing MS/MS with time-of-flight (Tof) data acquisition provides elemental composition information. For small molecules, this significantly improves the elucidation process since it significantly reduces the number of possible elemental compositions.
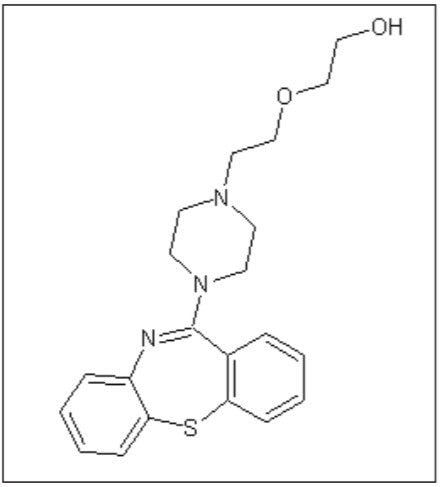
Quetiapine (Figure 1) is an atypical antipsychotic that has been shown to form more than 20 impurities/degradation products.2 In this application note, we demonstrate the application of high-resolution mass spectrometry to generate exact mass MS/MS fragment analysis data in conjunction with the use of software to automatically identify product ion fragments. This software applies a series of novel, chemically-intelligent algorithms to elucidate an unknown impurity peak of quetiapine that was previously isolated by mass-directed autopurification.3
|
LC system: |
ACQUITY UPLC |
|
Column: |
ACQUITY UPLC BEH C18 Column, 2.1 x 50 mm, 1.7 μm |
|
Column temp.: |
65 °C |
|
Flow rate: |
800 μL/min |
|
Mobile phase A: |
20 mM ammonium bicarbonate, pH 9.0 |
|
Mobile phase B: |
Acetonitrile |
|
Gradient: |
5 to 95% B for 3.0 min |
|
MS system: |
Waters SYNAPT MS System |
|
Ionization mode: |
ESI positive |
|
Capillary voltage: |
1.5 kV |
|
Cone voltage: |
15 V |
|
Desolvation temp.: |
350 °C |
|
Desolvation gas: |
650 L/Hr |
|
Source temp: |
120 °C |
|
Acquisition ranges: |
100 to 1000 m/z for MS 50 to 600 m/z for MS/MS |
|
Collision energies: |
Ramp from 15 to 30 |
|
Lock mass: |
300 pg/μL leucine/enkephalin flow at 50 μL/min |
MassLynx 4.1 Software with MassFragment
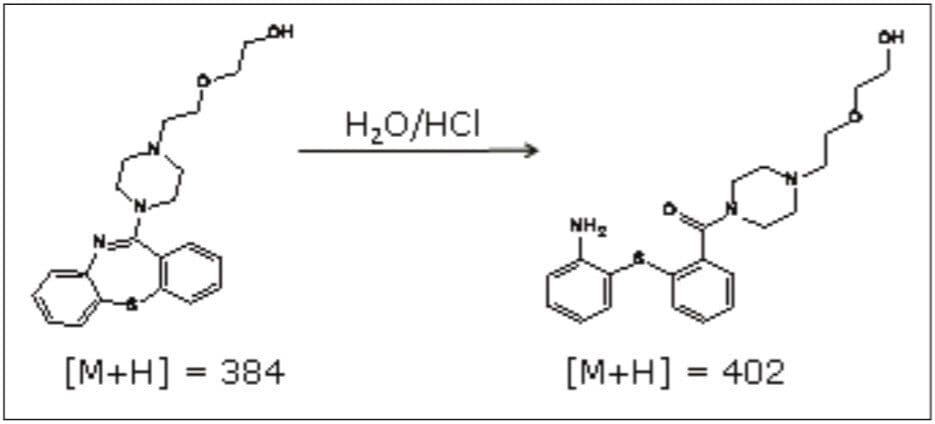
In a previous application note, forced degradation was employed to maximize the amount of an impurity formed during hydrolytic conditions to generate a sufficient mass of impurity for subsequent NMR analysis.2 The peak giving rise to the m/z 402 ion was isolated using preparative chromatography and mass-directed fraction collection. Although the desired substance was collected, the compound’s structure remained unknown.
Instrumentation capable of producing high mass accuracy information is an essential tool in the area of impurity determination. Modern Q-Tof technology makes the acquisition of sub-5-ppm mass accuracy data readily achievable. This, coupled with fast data acquisition speeds that allow the detection of the narrows peaks produced by UPLC chromatography, make it an ideal tool for impurity identification.
The sample was analyzed on the SYNAPT MS System to obtain accurate mass MS data and hence elemental composition determination. The impurity had an observed accurate mass of m/z 402.1838. This is an addition of 18.0092 amu to that of quetiapine (m/z 384.1746).
The high mass accuracy of the SYNAPT MS System produced a short list of possible elemental compositions. The potential elemental composition list was further reduced by a series of filters. These filters included the ability to limit molecular formulas based on:
The Elemental Composition browser returned three possible molecular formulas (Figure 2).
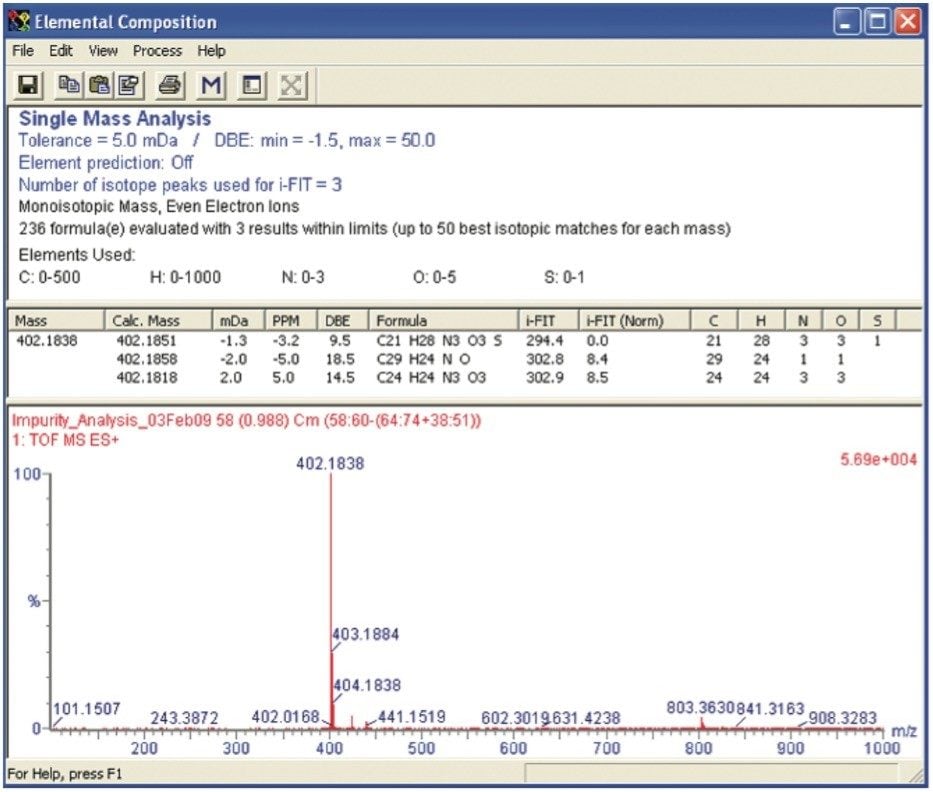
The Elemental Composition calculator was set to order the list of proposed elemental compositions according to the fit of the experimental data to the theoretical isotope distribution using i-FIT. The i-FIT criterion calculated a molecular formula of C21H28N3O3S as the best fit. The double bond equivalency (DBE) reported a value of 9.5 and a mass error of -1.3 mDa.
Possible structures that fit the elemental compositions based on the mass accuracy data for 402.1838 include:
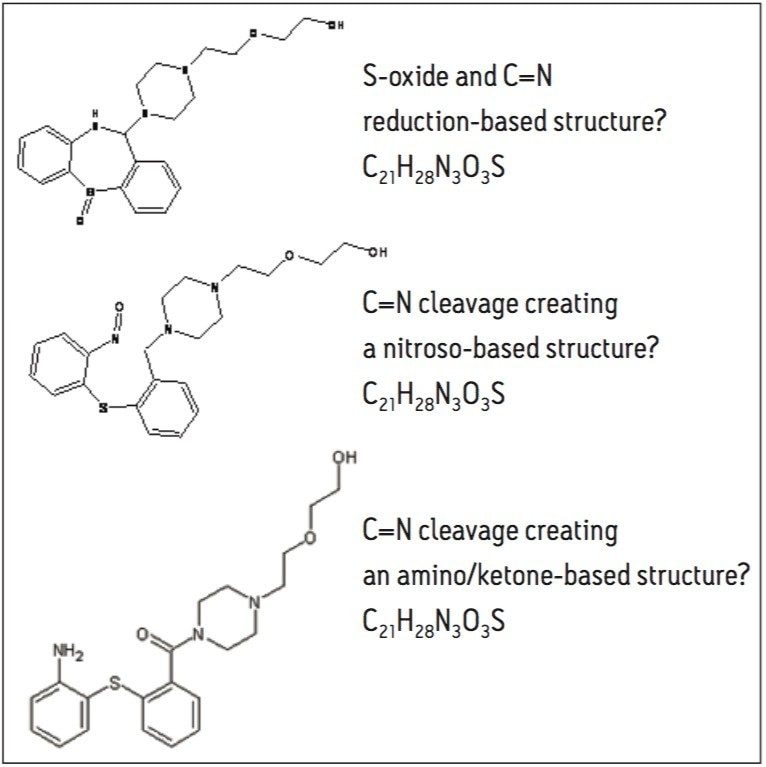
Examining the forced degradation reaction that enriched the production of the impurity suggests the most probable structure is as shown in Figure 4. Other chemical structures could be proposed based on the accurate mass information of the base m/z alone. Knowing the reaction that produced the addition of 18 amu aids the elucidation decision process.
A careful examination of the fragmentation patterns obtained during related impurity analysis allows the impurities’ structural information to be associated with that of the active pharmaceutical ingredient (API). The key to informative MS/MS data is the quality of the fragmentation spectra. The quality of MS/MS spectra assessed in small-molecule analysis is based on the number of fragments and spatial location of the fragments, thus providing added structural informative value for elucidation determination.
The SYNAPT MS Systems performs both MSE and traditional MS/MS experiments. MSE experiments allow for simultaneous high and low collision energy data to be collected with one injection. Traditional MS/MS experiments provide additional information in the form of product ion scans, precursor ion scans, and common neutral loss scans. An accurate mass MS/MS product ion scan of 402.1838 was performed to evaluate the fragmentation pattern of the isolated impurity to further support the proposed structure shown in Figure 3c (Figure 5). The fragments produced are most likely associated to cleavage of the seven-member ring that differentiates the three unknown impurities’ proposed structures.
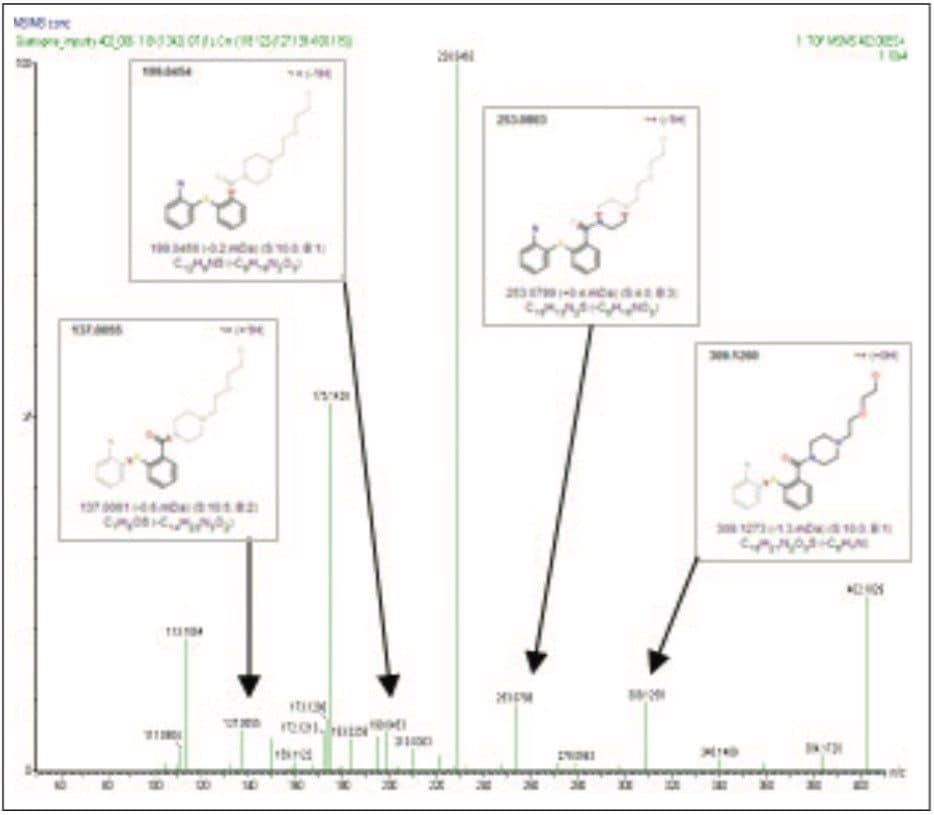
The key structural fragments that added greater confidence in differentiating amine/ketone-based structures to that of the alternatively-proposed structures were the m/z 137.0063, m/z 199.0454, m/z 228.0464, m/z 253.0803, and m/z 309.1260 ions.
Structural elucidation requires an expert knowledge of chemistry and bond reactions. MassFragment is a software solution that is capable of automatically identifying product ion fragments using a series of chemically-intelligent algorithms. This approach is based on systematic bond disconnection of the precursor structure instead of the traditional rule-based approach. MassFragment was used to help visualize the correlation of the product ion data with possible structural assignments for the observed fragment ions (Figure 6).

MassFragment increases our confidence in the MS/MS fragment analysis of the proposed structure in Figure 3c. The proposed structures were drawn and saved as *.mol files. Each *.mol file was loaded separately into MassFragment to be structurally correlated with the spectra in Figure 5. The MassFragment results ruled out the proposed nitroso-based and S-oxide based structures.
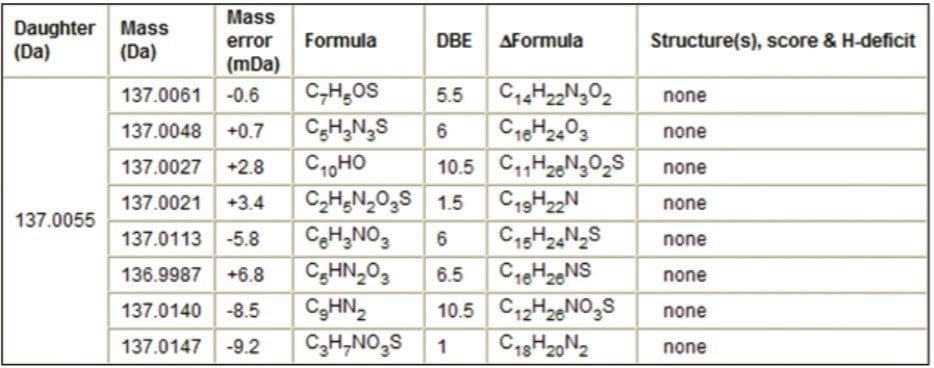
The fragment ion m/z 137.0063 was observed in the MS/MS spectra in Figure 5. The MassFragment reports for the S-oxide based structure and the amino/ketone-based structure observed the ion fragment m/z 137.0063 and displayed possible structures. MassFragment was not able to report a structure for this particular fragment ion when the nitroso-based precursor structure was loaded into MassFragment (Table 1).
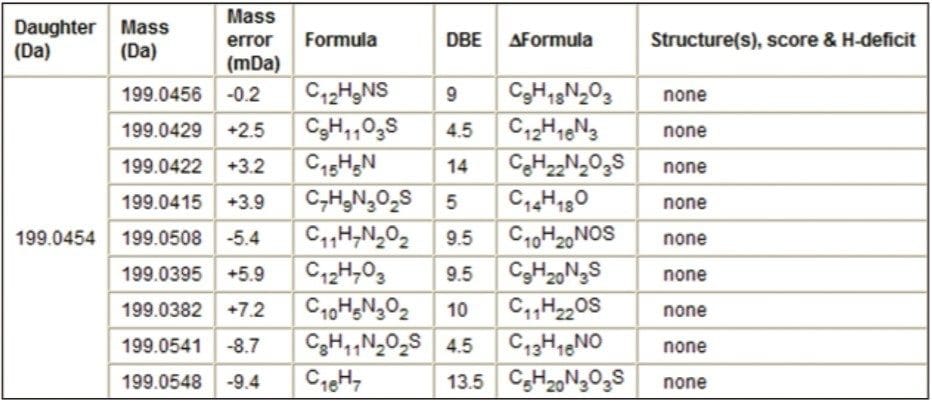
The fragment ion m/z 199.0454 was also observed in the MS/MS spectra. MassFragment reported a candidate structure for fragment ion m/z 199.0454 based on the amino/ketone-based structure (Figure 6). There were no reported structures for this fragment ion when the S-oxide based structure was loaded into MassFragment (Table 2).
The collected fraction was dried down and prepared in deuterated methanol for analysis by NMR. It was determined from the NMR data (not shown) that the impurity m/z 402 was consistent with the proposed structure.
For small molecule analysis, the advances in MS technologies with greater mass accuracy readily complement the advances in LC technology and sub-2-μm column particles capable of peak widths of 1 to 3 seconds. Laboratories performing impurity analysis can realize significant benefits by using chemically-intelligent software that improves how this LC/MSE data is analyzed.
Proper characterization of a compound provides greater insight to the reactive behavior of the compound that will lead to more informative formulation and packaging decisions.
720003079, June 2015