Size exclusion chromatography (SEC) is a common method used for investigating the extent of aggregation in protein therapeutics. In this application note, we demonstrate how to readily transfer a legacy SEC-HPLC method for monoclonal antibody (mAb) aggregation to the ACQUITY UPLC H-Class Bio System without any modification to existing method details.
Aggregation represents a serious concern for companies manufacturing large-molecule therapeutics. Numerous assays aid in establishing the extent of aggregation for a given product, one of which is size exclusion chromatography (SEC). SEC is a straightforward assay requiring minimal sample preparation that exploits the size, or more specifically the hydrodynamic radius, of a given molecule as the mechanism of separation. SEC is unique from many other large-molecule chromatographic approaches in the sense that separation occurs under isocratic conditions. Because separation is influenced exclusively by an analyte’s hydrodynamic radius and its ability to penetrate the pores of the stationary phase particle, there is no requirement for gradient conditions as adsorption is ideally non-existent and therefore does not influence migration times.
As observed with other large-molecule assays, significant benefits can be achieved in chromatographic quality when transitioning a method from HPLC to UPLC. The most obvious benefit is an increase in chromatographic resolution, driven principally by the increased chromatographic efficiency obtained through reduced column-particle sizes coupled with the use of low-dispersion instrumentation. This is particularly noticeable in SEC of large molecules as several higher-order aggregates can potentially be identified and quantified due to the increased resolution gained with SEC-UPLC. The added increase in sensitivity obtained through improved efficiency in SEC-UPLC also facilitates the identification of low abundant, higher-order aggregates that may have otherwise been undetected in SEC-HPLC.
Transferring SEC methods from HPLC to UPLC can be considered one of the more straightforward tasks as only a limited number of parameters need to be considered. With this in mind, there are significant opportunities for analysts to move legacy SEC-HPLC methods to UPLC technology.
To demonstrate the applicability of the ACQUITY UPLC H-Class Bio System for performing both SEC-HPLC and SEC-UPLC, we present here the transfer of an SEC-HPLC assay for monoclonal antibody aggregation from legacy HPLC instrumentation to the ACQUITY UPLC H-Class Bio. Following transfer of the HPLC method, we demonstrate a simplified approach to migrate the SEC-HPLC method to UPLC column chemistry.
Results of this study illustrate that the ACQUITY UPLC H-Class Bio System is capable of producing highly similar SEC-HPLC data compared to legacy HPLC instrumentation. Moreover, improvement in chromatographic resolution is obtained by moving from HPLC to UPLC. A reduction in run time is also observed without compromise to monoclonal antibody aggregate quantification. The experiment also illustrates the flexibility of using the ACQUITY UPLC H-Class Bio System for both SEC-HPLC and SEC-UPLC and the benefits of moving to smaller particle sizes for SEC-based assays.
|
LC system: |
ACQUITY UPLCH-Class Bio comprised of: ACQUITY UPLC H-Class Bio Quaternary Solvent Manager (QSM) ACQUITY UPLC H-Class Bio Sample Manager (SM) |
|
Extension loop: |
100 μL (p/n 430002625) |
|
Detector: |
ACQUITY UPLC Tunable UV Detector with Ti flow cell |
|
Absorption Wavelength: |
214 nm |
|
Column temp.: |
Ambient |
|
Mobile phase: |
20 mM Phosphate, 200 mM NaCl, pH 6.8 |
|
Sample: |
Waters BEH200 SEC Protein Standard Mix (p/n 186006518) |
|
HPLC column: |
Biosuite SEC Column, 250 Å, 10 μm, 7.5 mm x 300 mm (p/n 186002170) |
|
Injection vol.: |
20 μL |
|
Flow rate: |
0.400 mL min-1 |
|
Method length |
35 min |
|
UPLC column: |
ACQUITY UPLC Protein BEH SEC Column, 200 Å, 1.7 μm, 4.6 mm x 150 mm (P/N 186005225) |
|
Injection vol.: |
4 μL |
|
Flow rate: |
0.885 ml min-1 |
|
Method length: |
3 min |
To determine the ability of the ACQUITY UPLC H-Class Bio to perform legacy SEC-HPLC assays, we established a benchmark SEC-HPLC method using a quaternary pump HPLC instrument coupled with the BioSuite SEC 250 Å 10-μm Column (7.5 mm x 300 mm).
Two samples were used to evaluate the separation. The first sample was the Waters SEC200 protein standard mix, used to determine the total inclusion and exclusion volume of each column. The second sample was a therapeutic monoclonal antibody, infliximab, which was selected to measure aggregation, if any, and therefore a useful sample for evaluating the accuracy of method transfer between SEC run on the HPLC instrument and the ACQUITY UPLC H-Class Bio System. With respect to HPLC method parameters, separation in SEC is isocratic and, as such, the basic requirement of the method is that it run long enough to deliver a minimum of one column volume. For the selected HPLC column, a volume of approximately 13 mL was required and therefore a run time of 35 min was selected based on a delivered flow rate of 0.4 mL min.-1 A standard mobile phase of 20 mM phosphate buffer, 200 mM NaCl at pH 6.8 was selected for use and initial analyses.
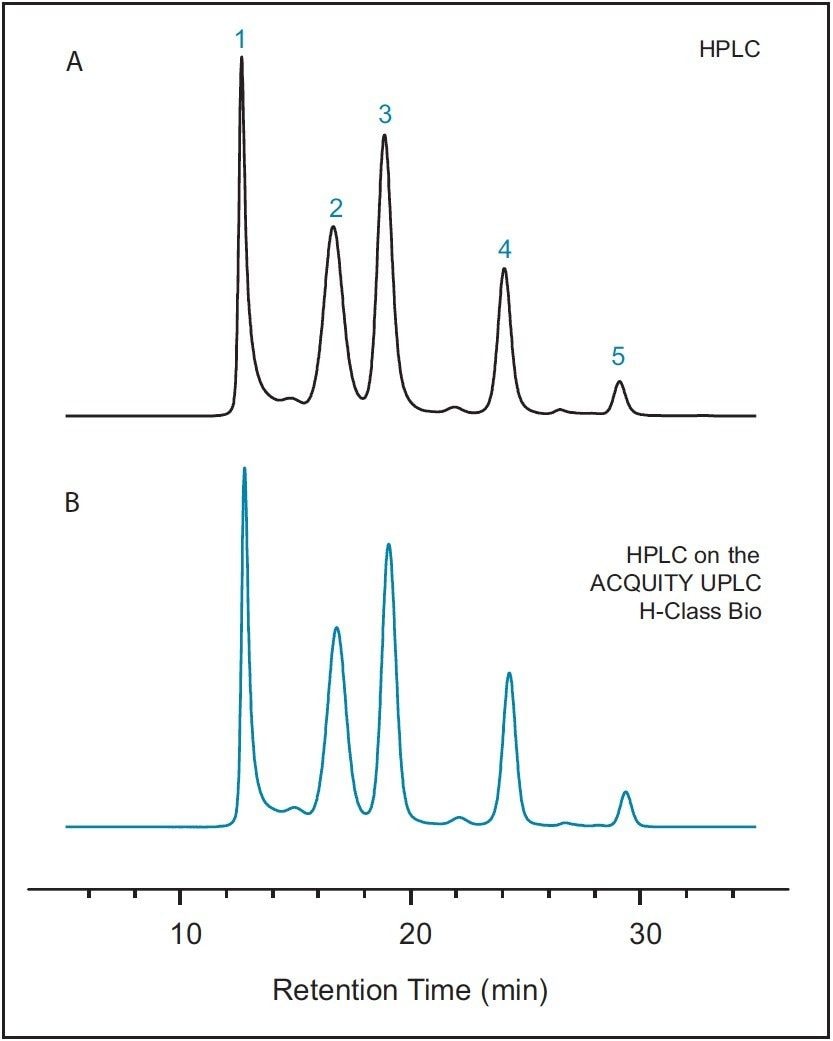
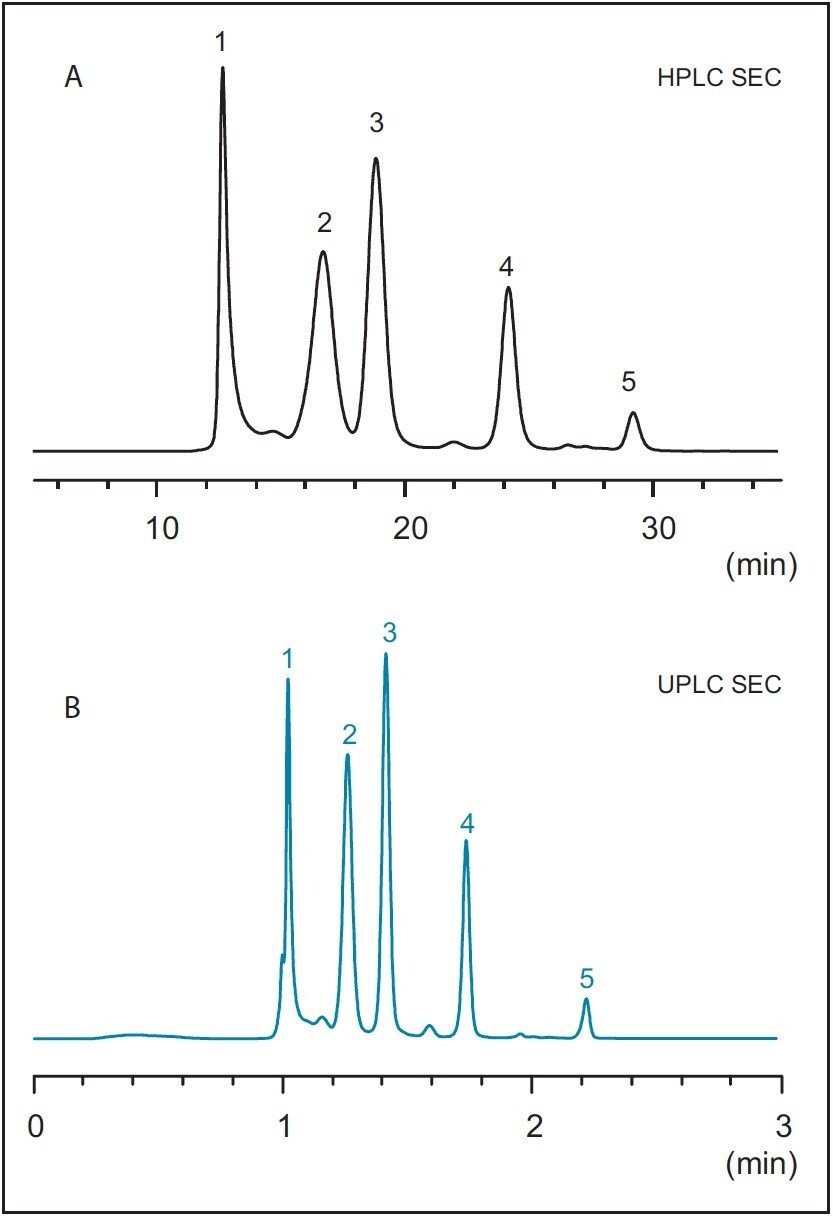
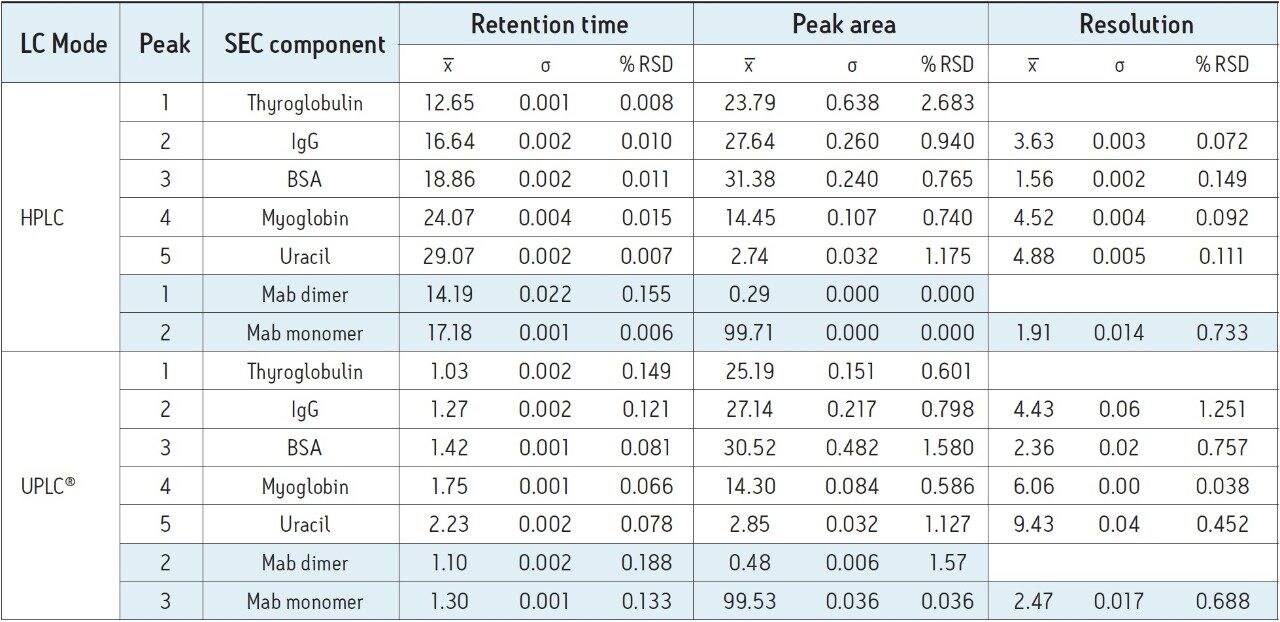
To first establish a benchmark chromatogram, the SEC200 protein mix standard was separated using the quaternary pump HPLC instrument with the SEC-HPLC column. Results of this separation are presented in Figure 1A. The SEC-HPLC column was then transferred to the ACQUITY UPLC H-Class Bio System without any changes to the method parameters. The same protein mix standard was separated, with results presented in Figure 1B. As can be clearly seen from these figures, very little difference is observed in the chromatography performed on each instrument. Details of the retention times and relative peak areas provided in Table 1 further indicate the accuracy of method transfer across analytical instruments, as evidenced by minimal differences between data recorded from each instrument.
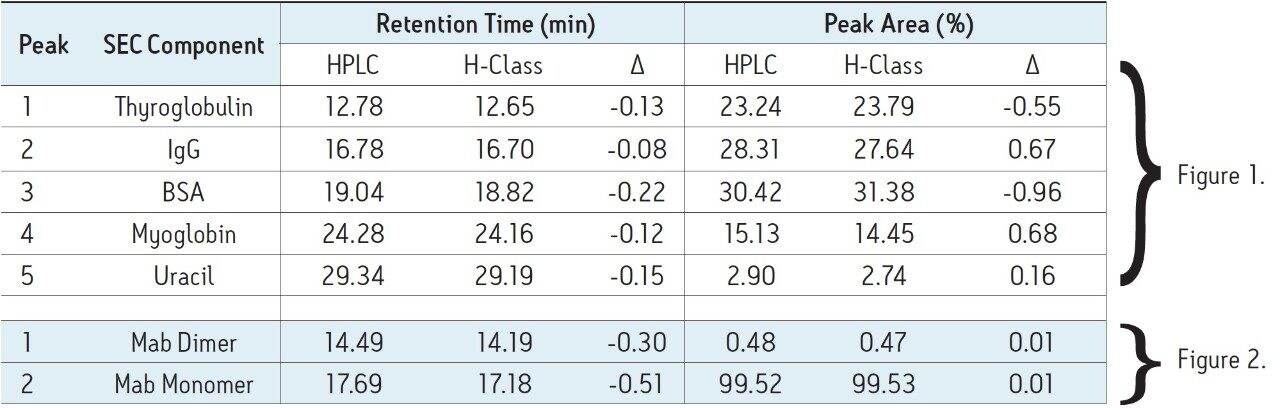
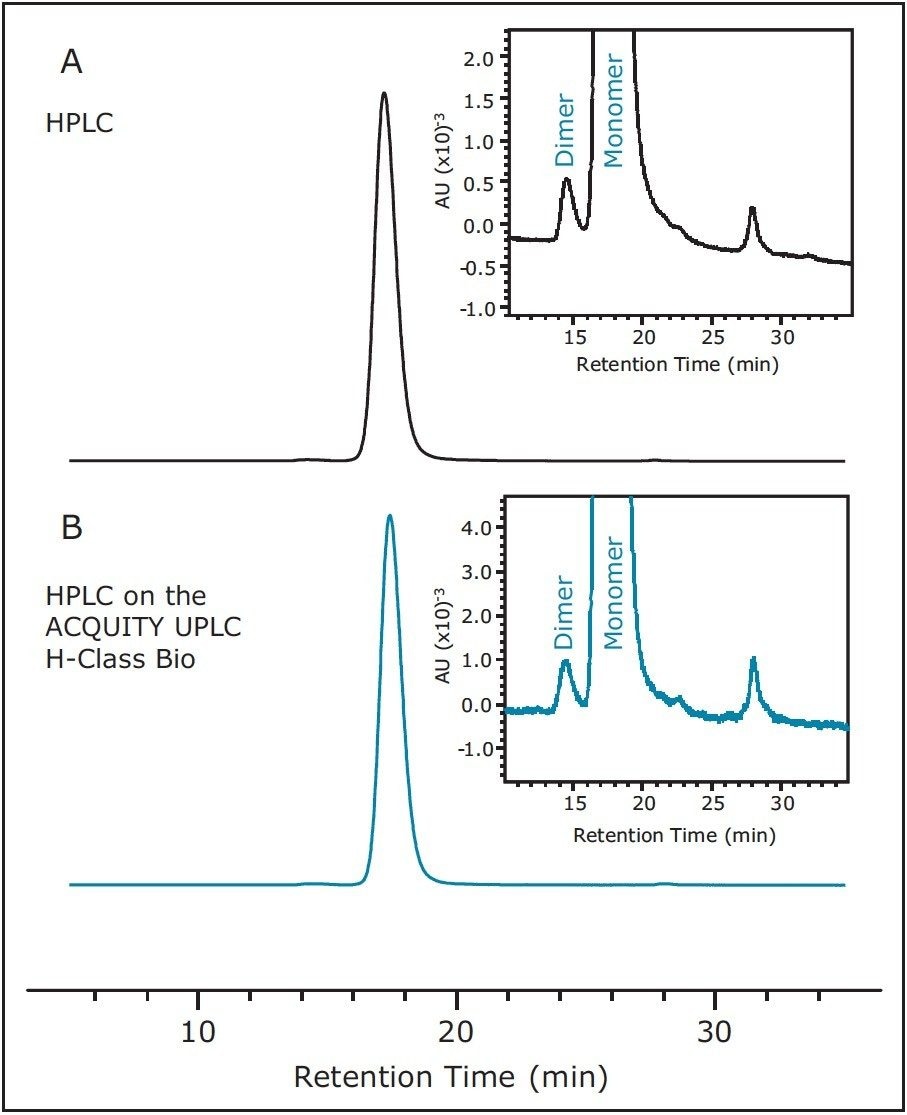
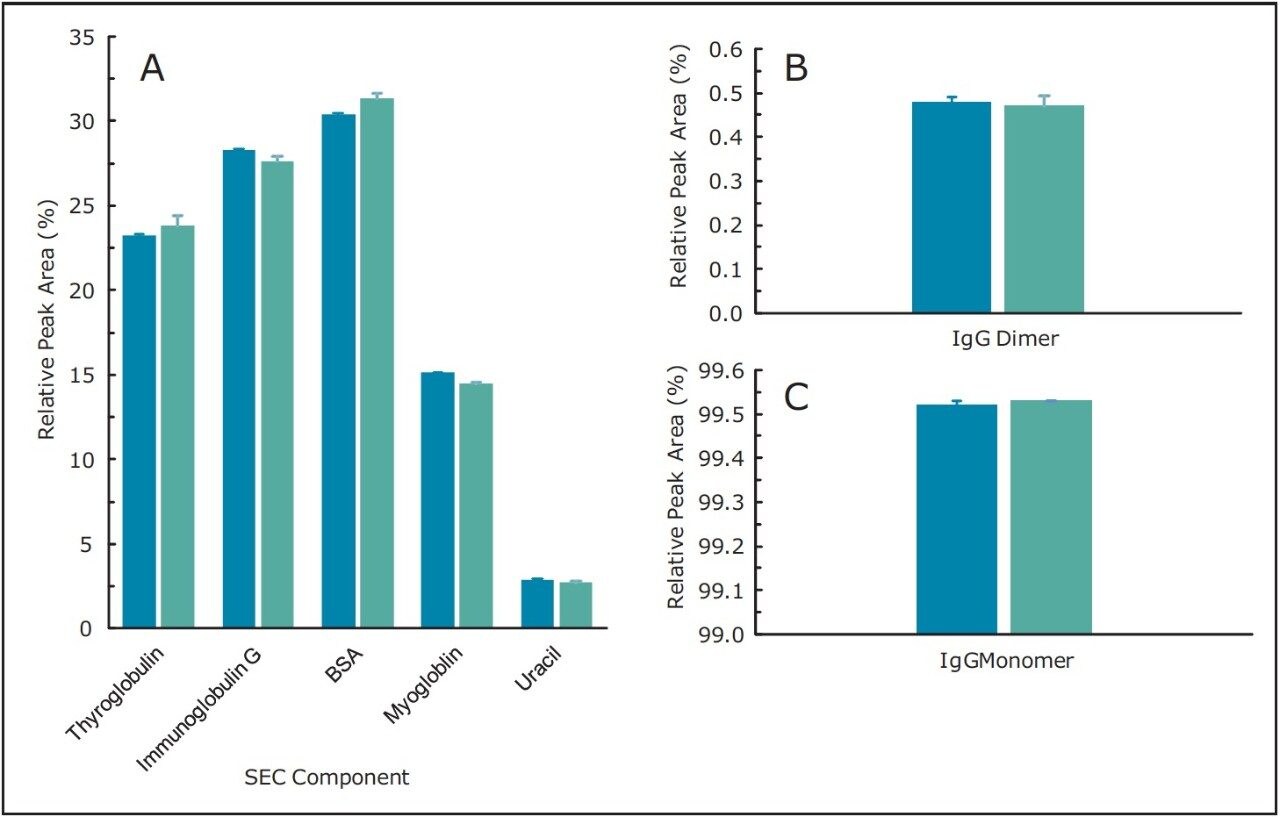
With conditions indicating a successful transfer of the SEC method to the ACQUITY UPLC H-Class Bio System, the monoclonal antibody infliximab was separated on both the quaternary pump HPLC instrument and the ACQUITY UPLC H-Class Bio System to determine the relative abundance of monomer and aggregate species within the sample. Similar to the protein standard results across instruments, very little difference was observed in chromatography (Figure 2) as well as relative peak areas in both the SEC200 protein standard mix and infliximab samples (Figure 3). Retention times, relative peak areas, and the resolution between the IgG monomer and dimer, also presented in Table 1, provide convincing evidence that SEC-HPLC assays can successfully be transferred to the ACQUITY UPLC H-Class Bio System without any compromise to the legacy analytical method criteria.
Significant benefits can be obtained by transferring legacy SEC-HPLC methods to SEC-UPLC columns, which provide improved chromatographic resolution through reduced column particle size. To illustrate this benefit, the SEC-HPLC method used for method transfer between the quaternary pump HPLC and the ACQUITY UPLC H-Class Bio System was adapted to be run on SEC-UPLC using the Waters ACQUITY UPLC Protein BEH SEC Column (200 Å, 1.7 μm, 2.1 mm x 150 mm).
To demonstrate the improved resolution as a result of reduced particle size on SEC, we separated the SEC200 protein standard mix using the 10-μm Biosuite SEC Column and the 1.7-μm ACQUITY UPLC Protein SEC column and calculated the differences in chromatographic performance. Both HPLC and UPLC separations were performed using the Waters ACQUITY UPLC H-Class Bio System. To accommodate for differences in particle size and column dimensions between the HPLC and UPLC columns, the flow rate was adjusted based on the following formula:
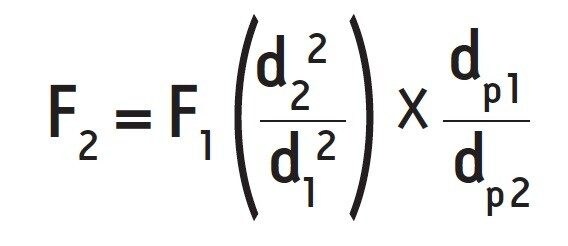
Where F refers to flow rate, d refers to column internal diameter, and dp refers to particle diameter. Based on the two columns used and an HPLC flow rate of 0.4 mL min,-1 a new flow rate of 0.885 mL min-1 was calculated. A new run time of 3 minutes was also determined for the UPLC separation, given the reduced volume of the UPLC column and the increased flow rate.
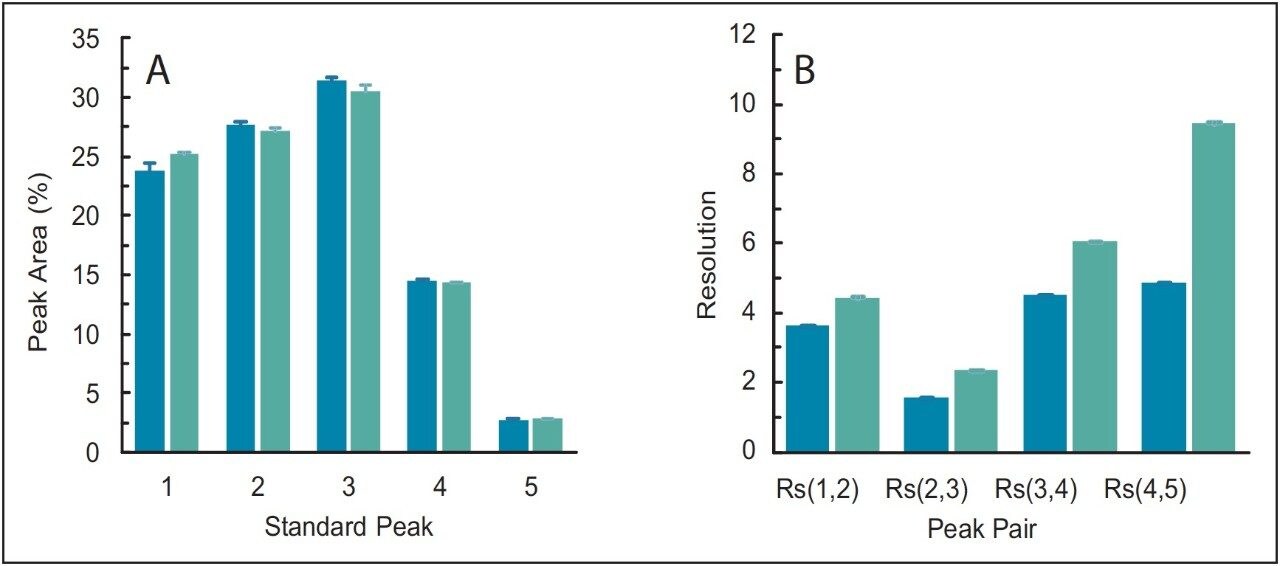
To determine the improvement in UPLC-based SEC, the SEC200 protein standard mix was separated on the BEH SEC 200 Å 1.7-μm column using the updated run time and flow rate. Most noticeable from the separation was the significantly reduced run time from 35 min in HPLC to just 3 minutes in UPLC (Figure 4). This was not at the cost of chromatographic performance, where relative peak areas are equivalent between HPLC and UPLC (Figure 5A) but a significant improvement in resolution between all peak pairs is observed in UPLC (Figure 5B). Resolution between all peak pairs are tabulated for reference (Table 2).
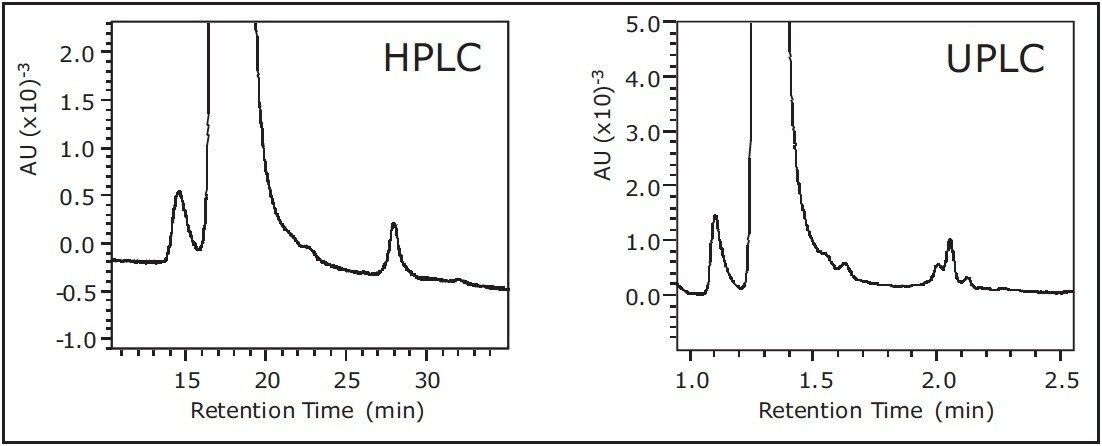
To determine what effect, if any, occurred on quantification in SEC, we separated the monoclonal antibody infliximab using the SEC-HPLC column and the SEC-UPLC column. In each SEC experiment, peaks corresponding to the monomer and dimer were integrated. As expected, a noticeable improvement in resolution was observed when separating infliximab in UPLC (Figure 6). Most importantly, the improvement in resolution did not affect the amount of aggregation quantified, where near identical relative peak areas for infliximab monomer and dimer were calculated on HPLC and UPLC. All chromatographic data has been reported in Table 2.
Size exclusion chromatography is a common method used for investigating the extent of aggregation in protein therapeutics. In this application note, we have demonstrated that legacy SEC-HPLC methods can be easily transferred to the ACQUITY UPLC H-Class Bio System without any modification to existing method details. With adjustments to flow rate and run time based on changes in particle size and column dimensions, SEC-UPLC can easily be performed with the same ACQUITY UPLC H-Class Bio System without any consequence to protein quantification requirements.
This permits QC labs to align LC technology and associated methods with development labs, while continuing to support legacy large-molecule assays currently deployed in the QC environment.
720005057, May 2014