In this study, we show the Progenesis QI workflow for metabolite identification using, as an example, a study on the effect of different bottling conditions on the nutritional composition of Italian wines.
Progenesis QI effectively streamlines and simplifies complicated metabolomics workflows and makes metabolite identification faster, easier, and more robust. User-definable search parameters dramatically decrease the number of false positive and false negative results in the identification workflow, improving the confidence of identification.
Progenesis QI Informatics simplifies the process of metabolite identification and biomarker discovery. Potential biomarkers can be searched in both publicly available and in-house databases for accurate mass, retention time, collision cross section, and fragmentation information. Such an approach is fast and it increases the confidence of metabolite identification in metabolomics experiments.
Metabolomics experiments offer a promising strategy for biomarker discovery. In a metabolomics workflow, however, the major bottleneck still remains metabolite identification. Currently, there are four levels of annotation for metabolite identification: 1) Confidently identified compound (two orthogonal properties based in authentic chemical standard analysis under the same condition); 2) Putative identified compounds (one or two orthogonal properties based in public database); 3) Putative identified compound class; and 4) Unknown compound.1 A typical database search that relies only on one property (i.e., accurate mass) usually leads to an extensive number of false positive and negative identifications. To increase the confidence of identification, a search engine should be able to use of in-house databases containing orthogonal molecular descriptors for each metabolite.2
Progenesis QI Informatics is a novel software platform that is able to perform alignment, peak-picking, and mining of metabolomics data to quantify and then identify significant molecular alterations between groups of samples. The software uses a search engine (MetaScope) for metabolite identification, with user-definable search parameters to probe both in-house and publicly available databases. With an easy-to-use interface, the user can combine information for metabolite identification, including accurate mass, retention time, collision cross section, and theoretical and/or experimental fragment ions. These physiochemical properties can increase the confidence of metabolite identification while concurrently decreasing the number of false positives.
In this study, we show the Progenesis QI workflow for metabolite identification using, as an example, a study on the effect of different bottling conditions on the nutritional composition of Italian wines.
|
System: |
ACQUITY UPLC |
||
|
Column: |
ACQUITY UPLC BEH HSS T3 1.8 μm 2.1 x 150 mm (p/n 186004120) |
||
|
Pre column: |
ACQUITY UPLC BEH HSS T3 VanGuard 1.8 μm, 2.1 x 5 mm (p/n186003976) |
||
|
Mobile phase A: |
Water + 0.1% formic acid |
||
|
Mobile phase B: |
Methanol + 0.1% formic acid |
||
|
Flow rate: |
0.28 mL/min |
||
|
Column temp.: |
40 °C |
||
|
Injection volume: |
10.0 μL |
||
|
Min |
A% |
B% |
Curve |
|---|---|---|---|
|
Initial |
100.0 |
0.0 |
Initial |
|
1.0 |
100.0 |
0.0 |
6.0 |
|
3.0 |
90.0 |
10.0 |
6.0 |
|
18.0 |
60.0 |
40.0 |
6.0 |
|
21.0 |
0.0 |
100.0 |
6.0 |
|
25.5 |
0.0 |
100.0 |
6.0 |
|
25.6 |
100.0 |
0.0 |
6.0 |
|
28.0 |
100.0 |
0.0 |
6.0 |
|
MS system: |
SYNAPT HDMS |
|
Mode of operation: |
Tof MSE |
|
Ionization: |
ESI +/- |
|
Capillary voltage: |
2.5 kV (+) and 2.5 kV (-) |
|
Cone voltage: |
25 V |
|
Transfer CE: |
Ramp 15 to 45 V |
|
Source temp.: |
150.0 ˚C |
|
Desolvation gas flow: |
1000 L/h |
|
Desolvation temp.: |
500.0 °C |
|
Cone gas: |
50 L/h |
|
MS gas: |
Nitrogen |
|
Acquisition range: |
50 to 1200 |
Progenesis QI Informatics
Mezzacorona winery (Trentino, Italy) provided the wines, which were bottled in typical 750-mL wine bottles with the filling industrial machine of the winery. The sample set included two types of wines bottled with nitrogen addition (N2) and without nitrogen addition (O2). Under nitrogen atmosphere, every wine was uncorked, 2 mL were transferred into a 5-mL amber vial, 2 mL Milli-Q water was added, and finally each sample was filtrated with 0.2-μm PTFE filters into a 2-mL Waters LCMS Certified Amber Glass Vial prior to LC-MS analysis.3
Wine is one of the most complex foods as far as its metabolomic profile is concerned, since grapes, yeasts, bacteria, fungi, exogenous antioxidants, fining agents, and other oenological materials, packaging, and aging are involved in its preparation. This great number of different primary and secondary metabolites, most of which are unknowns, highly affects wine quality and the important role it plays in human diet, health, and enjoyment. Different bottling and storage conditions may affect the molecular composition of wines, and thus value and quality.
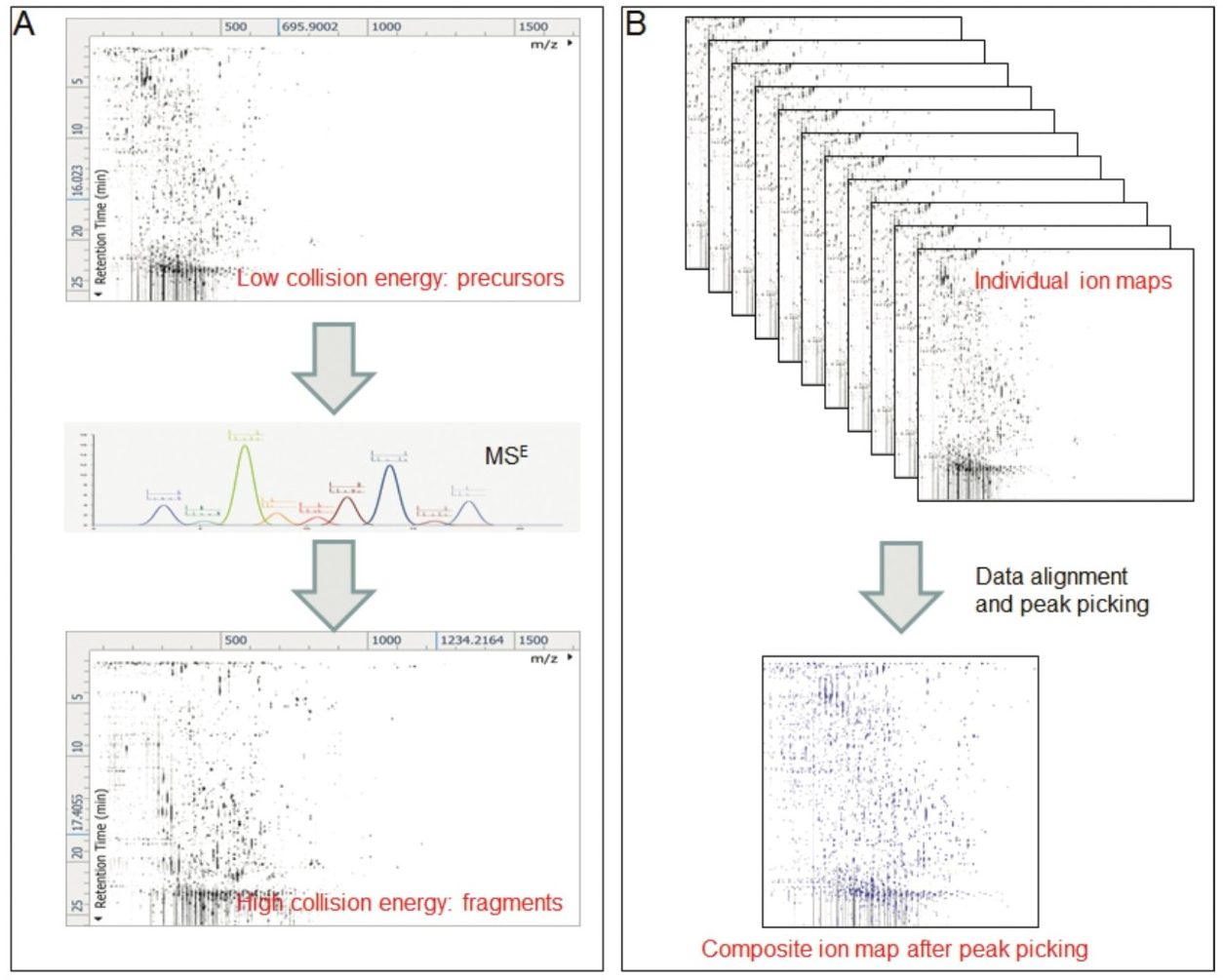
In this study, we used Progenesis QI to identify the metabolites that were altered between wines bottled under two different levels of oxygen: high level (O2) versus low level (N2). Data were acquired in LC-MSE mode (Figure 1A) and pre-processed using retention time alignment and peak picking (Figure 1B). A composite ion map was built, which contained more than 3,000 compounds after isotopic and adduct deconvolution (Figure 1B). Metabolites of interest were filtered according to the ANOVA P value <0.01 and fold change >2, which decreased the number of metabolites of interest (markers) to less than 200 (Figure 2A). This data reduction strategy allowed us to focus on the metabolites that clearly discriminate the two groups of samples as shown by principal component analysis (Figure 2B).
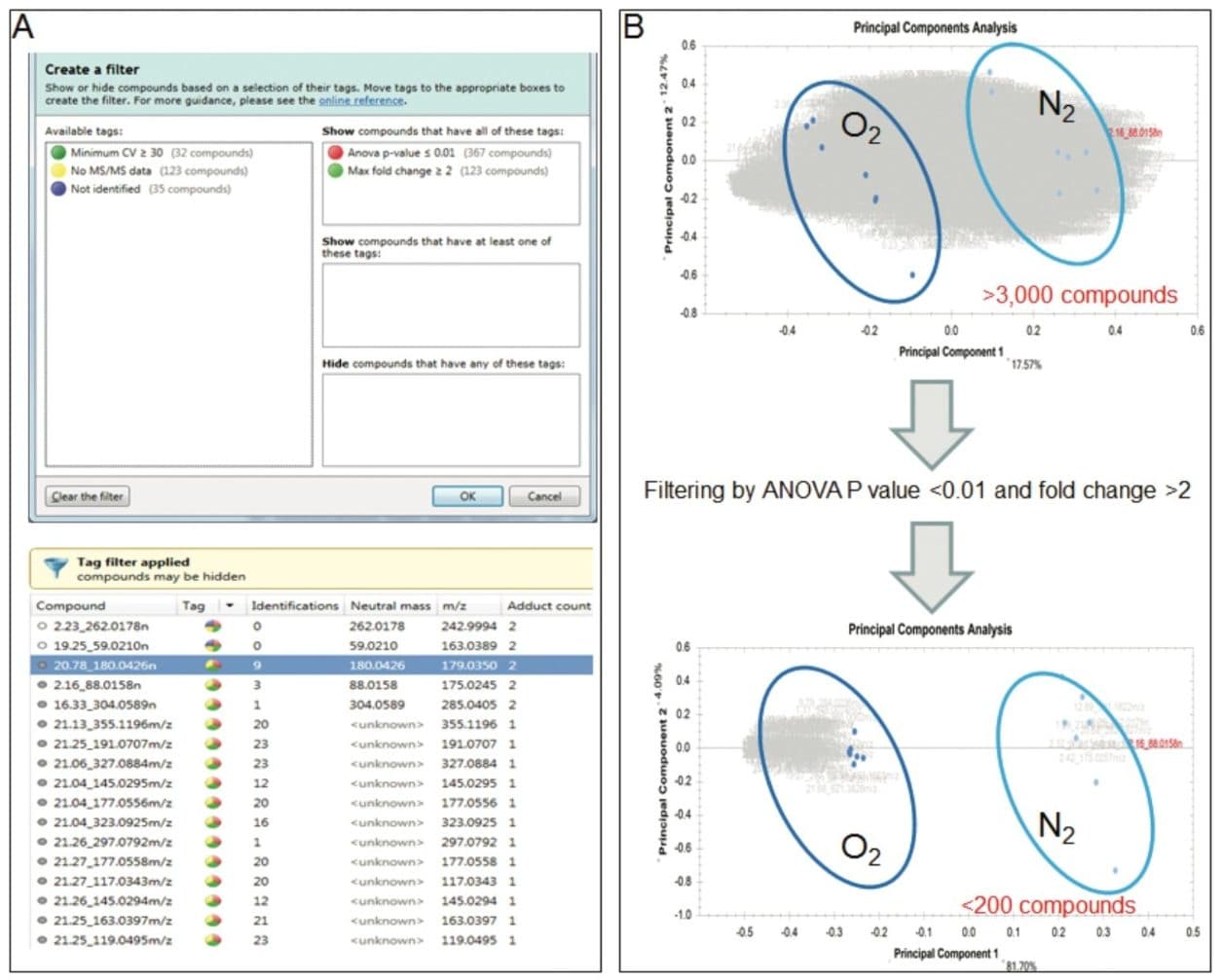
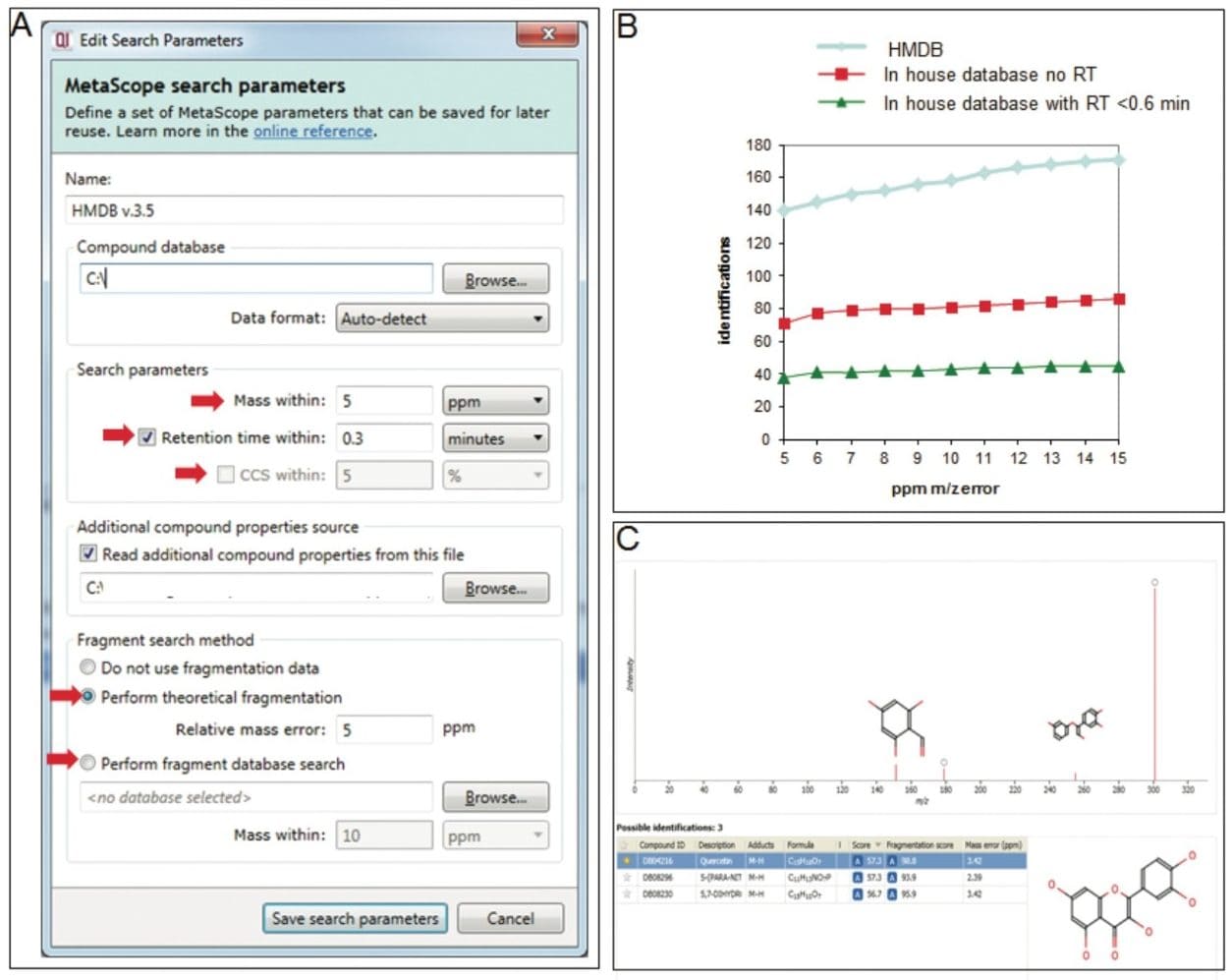
Initial identification of metabolites was performed using the Human Metabolome Database (HMDB), leading to multiple ambiguous identifications for each compound of interest (Figure 3A and 3B). To decrease the number of false positives, we used in-house metabolite databases, which contain accurate mass, retention time, and fragment information.2 (Figure 3A-C). We customized the search engine parameters for these orthogonal measures (Figure 3A), allowing a more balanced set of tolerance criteria, which significantly decreased the number of false positives and false negatives (Figure 3B). Experimental fragments were matched against those derived from theoretical fragmentation to further increase the confidence in metabolite identification (Figure 3C). The entire metabolomics workflow for data processing, mining, and identification was completed in just a few hours.
Progenesis QI effectively streamlines and simplifies complicated metabolomics workflows and makes metabolite identification faster, easier, and more robust. User-definable search parameters dramatically decrease the number of false positive and false negative results in the identification workflow, improving the confidence of identification.
720005044, April 2014