This application note discusses key elements of the LC-MS intact mass workflow and applies them in the comparative analysis of two batches of a therapeutic monoclonal antibody. It details how BiopharmaLynx software can used to demonstrate batch-to-batch differences in the glycovariant profile of a recombinant therapeutic monoclonal antibody. BiopharmaLynx automates processing and bioannotation of data, and provides tools for sample comparison, data export, and report generation, resulting in improved workflow efficiencies.
Intact protein LC-MS analysis of a biotherapeutic provides a holistic view with a much simpler set of data than “divide and conquer” methods such as peptide mapping. The major tradeoffs for intact protein MS analysis are that the types of detectable modifications become more limited as protein mass increases and that modification sites cannot be deduced from the intact mass alone.
Assessing clone-to-clone or batch-to-batch variation for glycosylated therapeutic proteins is a common and routine task that often requires significant resources for manual data processing and results integration. When high throughput or rapid sample turnaround is desired, analytical approaches that provide rapid global information about a molecule are preferred over slower approaches that generate more detailed information.
For modifications such as oxidized methionines, there is roughly a 16 Da increase over the unmodified protein mass. These modifications can be readily detected at lower levels in the TOF mass spectra of smaller proteins (30 kD and below, e.g. antibody light chains), but require significant stoichiometry to be detected on a 50 kD protein (e.g. antibody heavy chains), and are not readily detectable on large proteins such as a 150 kD intact antibody. For this reason, intact antibody analysis is almost always used in combination with reduced antibody (LC/HC) analysis and peptide mapping studies for full characterizing batches of a therapeutic antibody.
In general, there are two classes of protein variants that are typically scrutinized using mass analysis of an intact antibody: glycan structure heterogeneity where variation of core glycans on each heavy chain is extended by a series of 146 to 291 Da carbohydrate units and, the potentially inefficient proteolytic processing of heavy chain carboxy-terminal lysine residues (+128 Da).
Assessing this heterogeneity using intact antibody mass profiles can be useful for selecting a clonal expression cell line with desirable product attributes and monitoring the effects of process changes on a biotherapeutic. It is also a useful approach for demonstrating the consistency and comparability of individual batches of drug product.
Comparative LC-MS analysis of intact antibodies can be accomplished using rapid LC-MS methods.1-3 Using proper methodology, this type of analysis is robust, and pharmaceutical companies have acquired intact mass data on literally thousands of proteins using OpenLynx enabled open access LC-MS analysis stations.4-6 The resulting deconvoluted intact mass information found in an OpenLynx report is usually sufficient for routine mass confirmation of a recombinant protein, but often proves limiting for in-depth characterization or comparative profiling studies of biotherapeutics.
BiopharmaLynx intact protein analysis workflows are designed to expand functionality for automated batch processing of data by providing additional processing capabilities and related deconvoluted masses to targeted proteins and their variants.

In addition, the graphical graphical and tabular tools necessary for comparisons of experimental samples against a “gold” reference standard are present in the application.
This application note discusses key elements of the intact mass workflow, and applies them in the comparative analysis of two batches of a therapeutic monoclonal antibody. Overall, the use of such automated data processing and annotation workflows should allow biopharmaceutical organizations to screen more samples in less time, and make better decisions faster for the development of their biotherapeutic molecules.
The LC-MS configuration included an additional post-column desalting valve (system controlled) as detailed in the Care and Use document for the MassPrep Intact Protein Desalting Kit.
The data were acquired using BiopharmaLynx v.1.1, a MassLynx Software application manager that allows laboratories to streamline and automate the processing, bioannotation, and comparison of peptide map and intact protein LC-MS data sets.
|
LC system: |
Waters nanoACQUITY UPLC System |
|
Column: |
MassPREP Micro desalting 2.1 x 5 mm |
|
Column temp: |
65 °C |
|
Flow rate: |
20 μL/min |
|
Mobile phase A: |
Water with 0.1% formic acid |
|
Mobile phase B: |
ACN with 0.1% formic acid |
|
Gradient: |
10% B for 5 min, 10% B to 90% B over 5 min |
|
MS system: |
SYNAPT High Definition Mass Spectrometry (HDMS) System |
|
Ionization mode: |
ESI+ |
|
Capillary voltage: |
2.0 kV |
|
Cone voltage: |
65 V |
|
Desolvation temp: |
250 °C |
|
Desolvation gas: |
100 L/hr |
|
Source temp: |
105 °C |
|
Acquisition range: |
1000-4600 m/z |
|
Calibration range: |
1000-4600, CsI, external |
|
Sample: |
0.5 mg/mL of a monoclonal IgG1 in 25 mM ammonium bicarbonate |
|
Sample volume: |
3 μL |
Screening of intact antibody variants was accomplished using a simple desalting LC-MS approach. We have previously described1-3 rapid, reversed-phase desalting methods that use short four-minute cycle times for an intact antibody to facilitate post-column salt diversion during sample loading. Such desalting strategies are advantageous when fast sample turnaround or higher sample throughput is desired.
Residual salts on the bound protein were washed away at low organic modifier; rapid gradients were generated to elute proteins as concentrated peaks for mass analysis; and additional sawtooth gradients were applied to ensure that the column was in pristine condition for the subsequent injection. In this work, a variant of this methodology was optimized for the nanoACQUITY UPLC System, and applied for the analysis of two commercial monoclonal antibody preparations.
Automating the data processing and interpretation of two intact antibody desalting runs in BiopharmaLynx was accomplished using spectral summation over the chromatographic elution profile of the antibody, subjecting the summed spectrum to MaxEnt1 spectral deconvolution, and searching the resulting deconvoluted masses against the antibody sequence and a series of common biantennary glycan modifications. The following sections will describe this processing workflow in greater detail.
Our standard approach for processing very large proteins, such as intact antibodies, is to deconvolute a selected high signal-to-noise m/z range within the summed mass spectrum. This “heartcut” m/z region (typically comprising at least six charge states) contains the best population of peaks with good signal-to-noise that are not artificially broadened by the presence of adducts, (which is an issue at higher m/z). These higher m/z peaks, (fewer charges) represent more folded structures that can preferentially maintain neutral and salt adduct associations during the electrospray process.
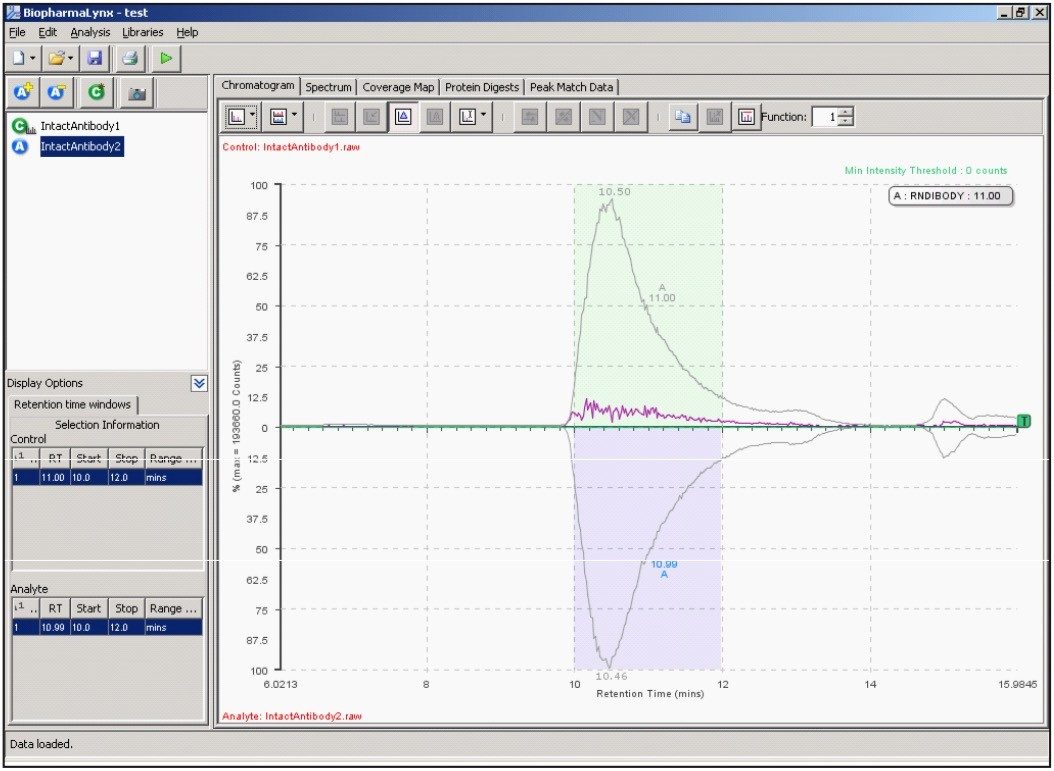
Our method (Figure 1) specifies the spectral deconvolution of peaks between 2500 and 3000 m/z in the raw data summed between 10 and 12 minutes to produce a MaxEnt1 zero-charge-state deconvolution spectrum in the range of 147,000 to 150,000 Da. This selected retention window and mass range are highlighted by shaded regions on the mirrored differential plots of the total ion chromatogram (Figure 2) and summed raw mass spectra (Figure 3A). Enlarging several charge-states within this region (Figure 3B) shows the complex nature of each charge-state arising from the various glycovariants of the antibody, while the differential plot shows consistent relative intensity differences of individual glycovariants across charge-states. From this repeating pattern of differential intensity, it would be expected that the final deconvoluted result would reflect a similar pattern of relative glycovariant intensity differences between the two samples.
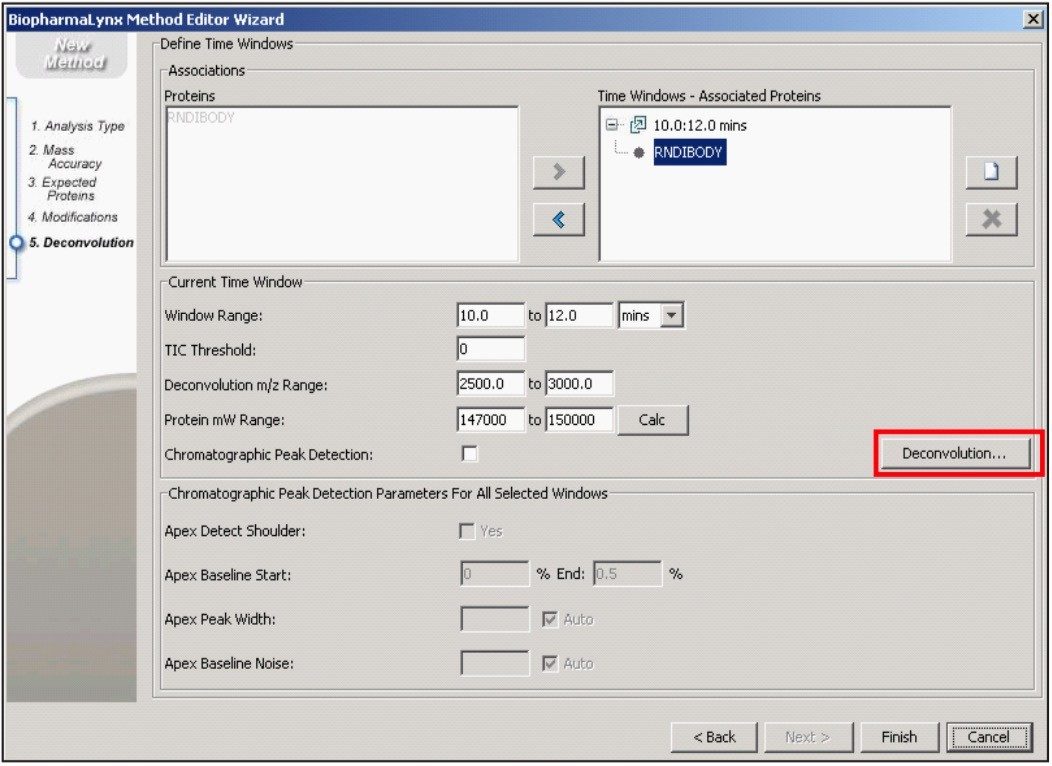
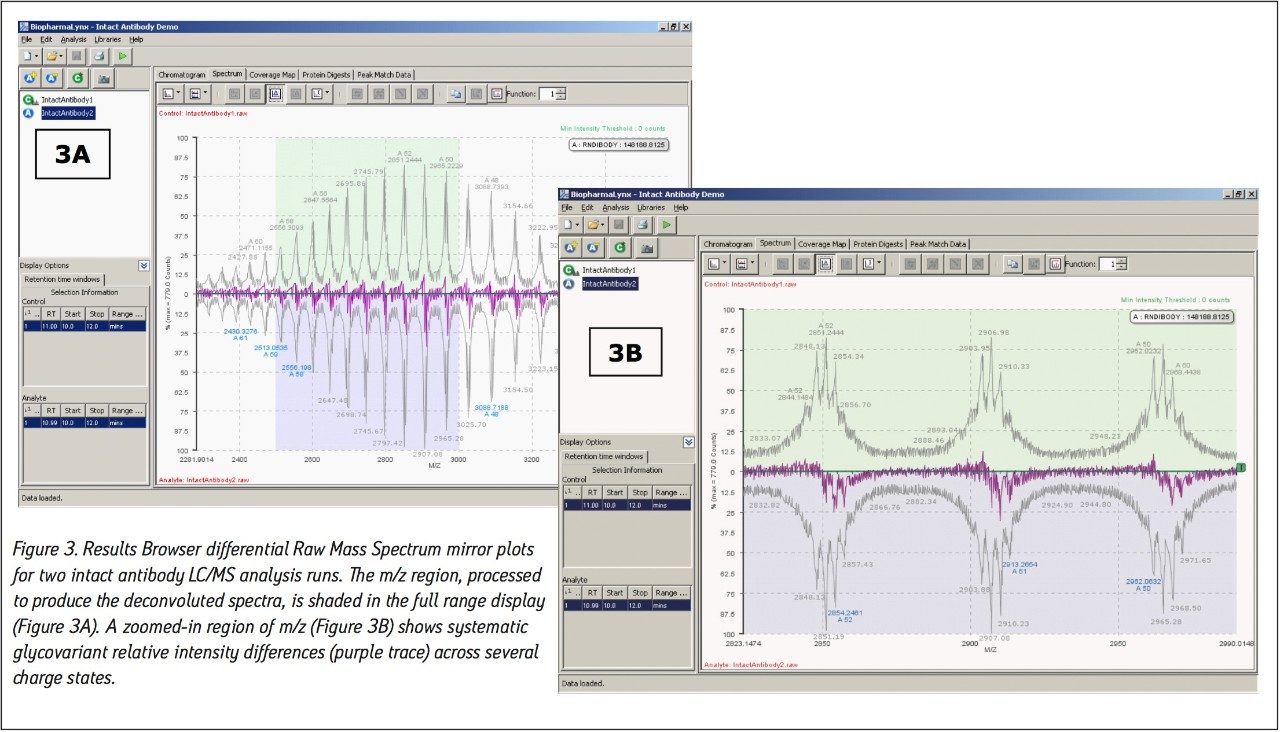
Additional settings for MaxEnt1 deconvolution of the antibody data are accessible through the advanced deconvolution settings button (Figure 1, red box highlight). These settings (Figure 4) show that the summed spectrum was background subtracted prior to applying 15 cycles of MaxEnt1 spectral deconvolution, and that the MaxEnt1 deconvoluted spectrum was background subtracted prior to measuring the height centroid of each peak. It is these centroid mass-intensity values that populate the results table and that are used for annotating protein variant structures.
BiopharmaLynx contains the latest generation MaxEnt1 spectral deconvolution algorithm. While a full discussion of this improved algorithm will appear in a separate technical note, it is important to focus on several settings (Figure 4) selected for processing of the antibody LC-MS data. First, the “automated peak width” function is not selected. This new capability automatically populates the MaxEnt1 spectral peak width setting based on the entered instrument resolution and average output mass range. This width value can be modeled effectively for smaller proteins (under 40 kD) that do not have significant charge-state peak broadening due to adducts or micro-heterogeneity. However, the width value cannot be modeled effectively for larger proteins due to the greater occurrence and magnitude of these effects. In both the automated and manual peak width modes, the improved MaxEnt1 algorithm now better models TOF instrument data by recognizing that peak width on TOF instruments systematically varies with m/z.
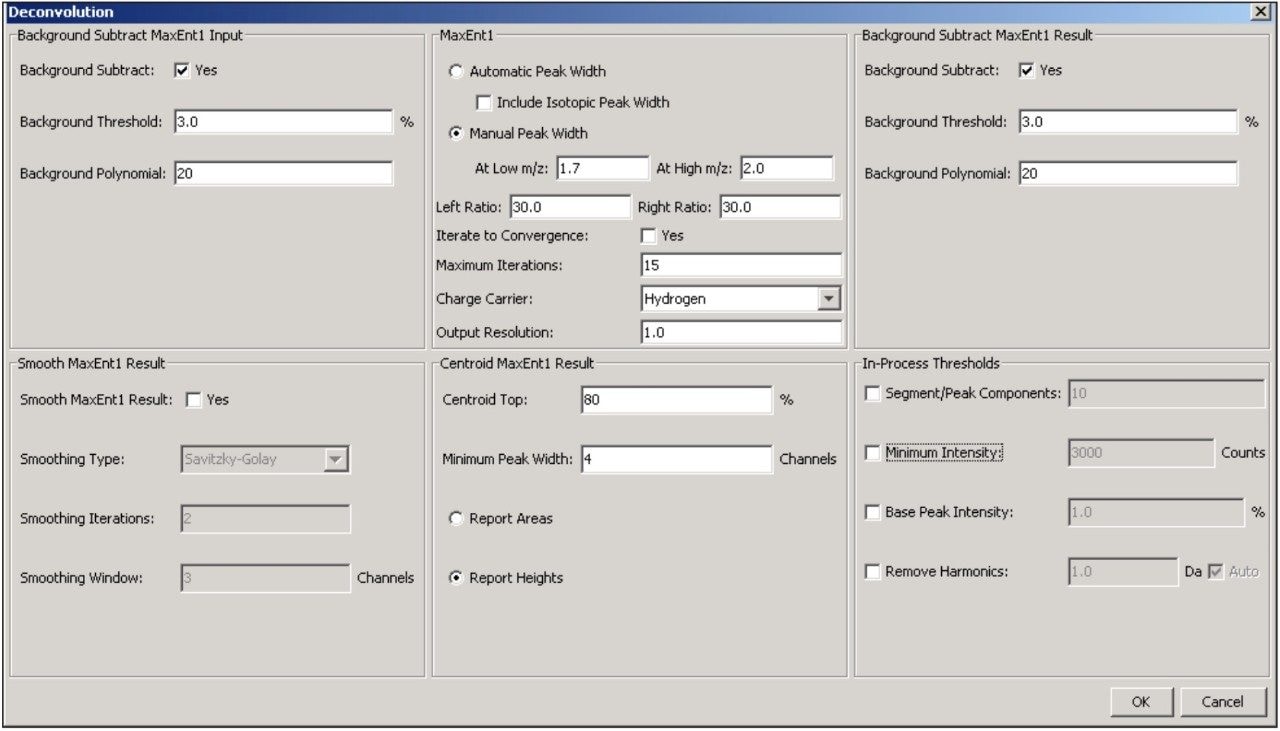
Thus, two spectral peak width settings (typically measured for the best resolved or most intense variant at half peak height) that correspond to the highest (low m/z) and lowest (high m/z) charge-states used for deconvolution can be entered in the manual mode. For the two antibody data sets, these values were determined as 1.7 and 2.0, respectively. Selecting a common value for both entries would mirror what is implemented in the current MassLynx Software v.4.1 implementation of MaxEnt1, when the “Uniform Gaussian” peak width model is used.
This method applies 15 iterations of MaxEnt1 processing to the data rather than taking the deconvolution process to full convergence of the mass spectral and model (“mock”)-spectral data sets. This choice is made in recognition that the peak models used by MaxEnt, even with variable spectral peak width, cannot perfectly match data for large proteins (>100 kD). There is unresolved micro-heterogeneity within each charge-state, and adducts preferentially interact with lower charge-state/structures of a protein. Both effects produce additional variance in peak width and peak asymmetry that are not directly predictable. This unaccounted-for spectral signal can contribute to additional peak structure in the deconvoluted spectrum, which is not evidenced within the raw spectral data, and which typically begins to appear in later MaxEnt1 iterations. The improved TOF model has reduced the occurrence and magnitude of such peaks, but they may still be generated during large protein deconvolution studies.
Selecting the number of iterations provides proper resolution and relative quantitation of protein variants, while avoiding these potential artifacts, is an important part of developing a robust data processing workflow for large proteins. The proper balance can be achieved for biotherapeutic antibody analysis, as indicated by the recent published work of Gadgil and colleagues.8
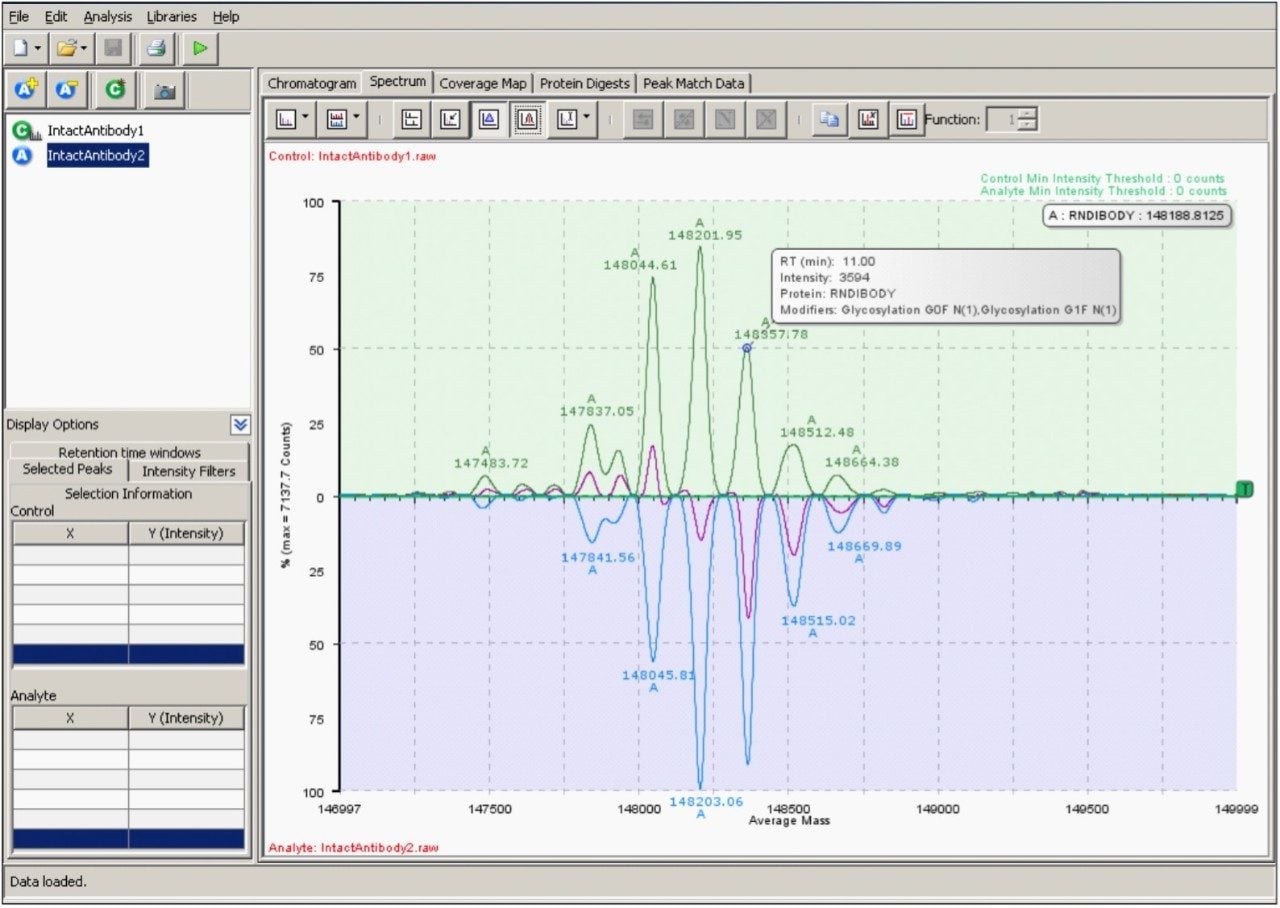
The resulting deconvoluted spectrum produced by MaxEnt1 processing of the two data sets (Figure 5) displays systematic differences between antibody samples. Consistent with the raw spectra differential display (Figure 3B), the lower sample is enriched with higher mass glycovariants and deficient in some lower mass glycovariants. As a simple validation exercise, true differences between deconvoluted protein spectra should be observable in the underlying raw charge-state data, although these differences can be subtle. Additional information about a selected peak in the deconvoluted data (corresponding to the G0F/G1F glycoform) was obtained by mouse-over at the peak apex. The centroided mass-intensity information derived from this deconvoluted spectrum was used to produce tabular results and processed spectra/chromatogram views (not shown) of the data.
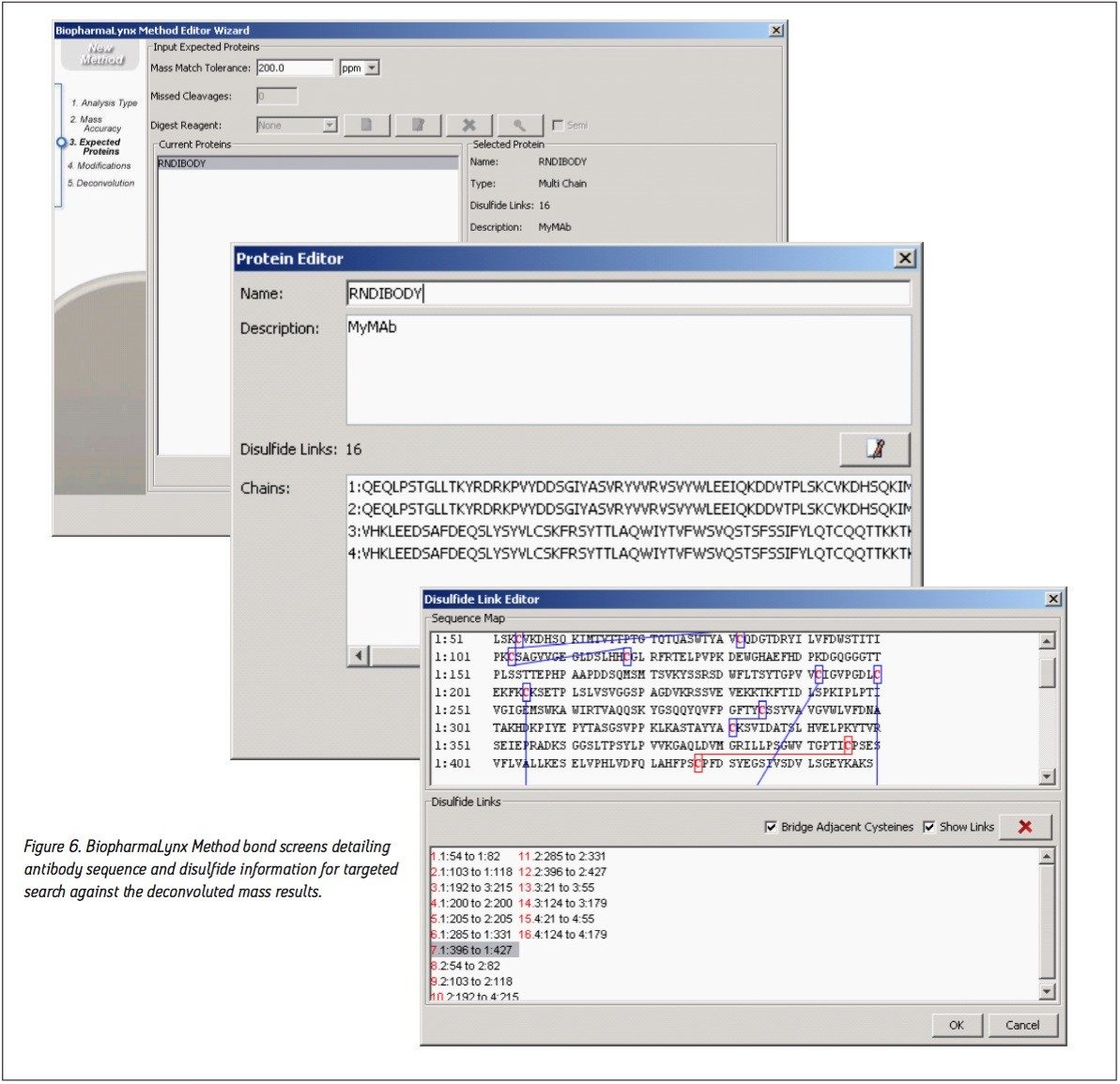
The bioinformatic assignment of antibody masses to protein structures was also accomplished as part of the processing method. The recombinant IgG1 antibody sequence was defined in the method (Figure 6) as a single protein with four protein chains (two heavy, two light) containing 16 intermolecular and intramolecular disulfide bonds. The sequence has been randomized to obscure the identity of the biotherapeutic molecule used in this comparative analysis. Defining the correct number of disulfide linkages is important, as each disulfide will increment the predicted protein mass downwards by 2 Da, or roughly a -32 Da mass difference for the intact antibody.
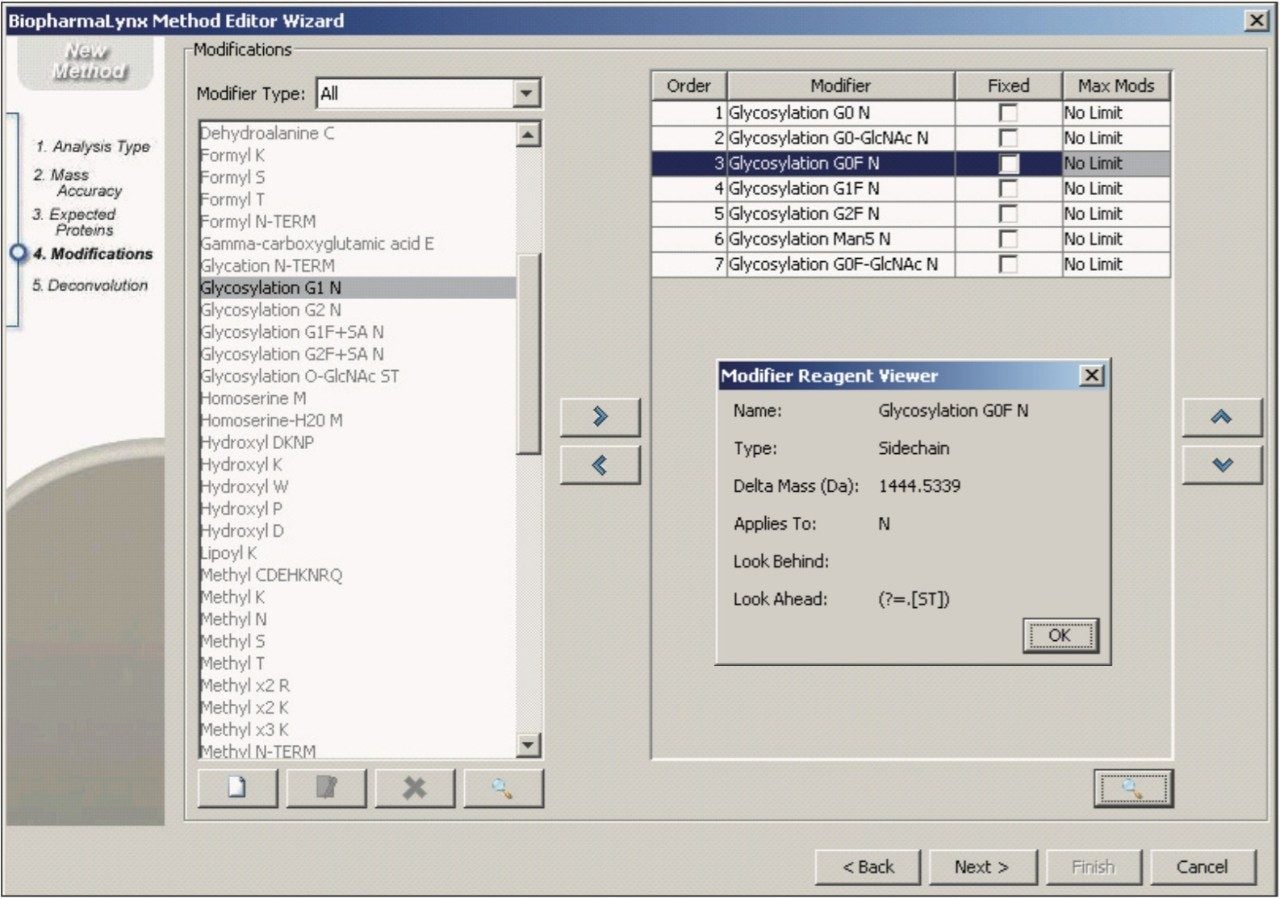
Modifications searched in the method (Figure 7) included a set of seven common biantennary glycan structures targeted to canonical NX(S/T) N-glycosylation sites (e.g. G0F in Figure 7, inset).
The peak match data table (Figure 8) links the processed data to the proteins and modifications assigned through a targeted bioinformatic search. Shown are the top 10 most intense components identified in the control sample, sorted from the most to least intense.
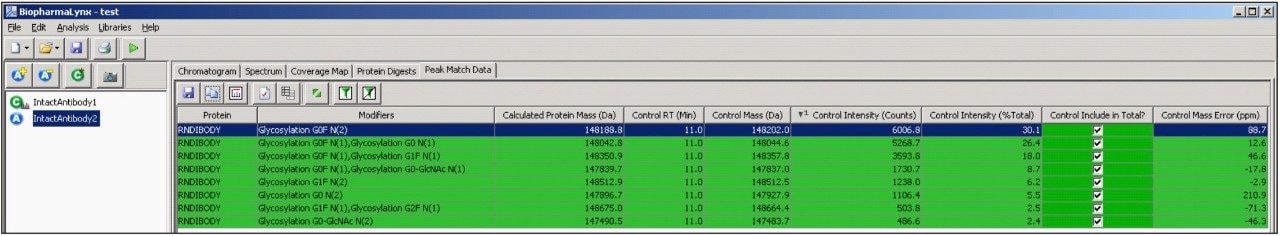
The table also contains information about the detection of these components in the analyte, and mass/intensity differences between the two samples (not shown for figure legibility). All 10 components were assigned to antibody glycovariants structures, and the green shaded row indicates that the components were all detected within the deconvolution results generated for both samples.
The control intensity (% Total) value is linked to the selection column to its right, and generates on-the-fly calculations of relative composition of the selected components within an individual sample.
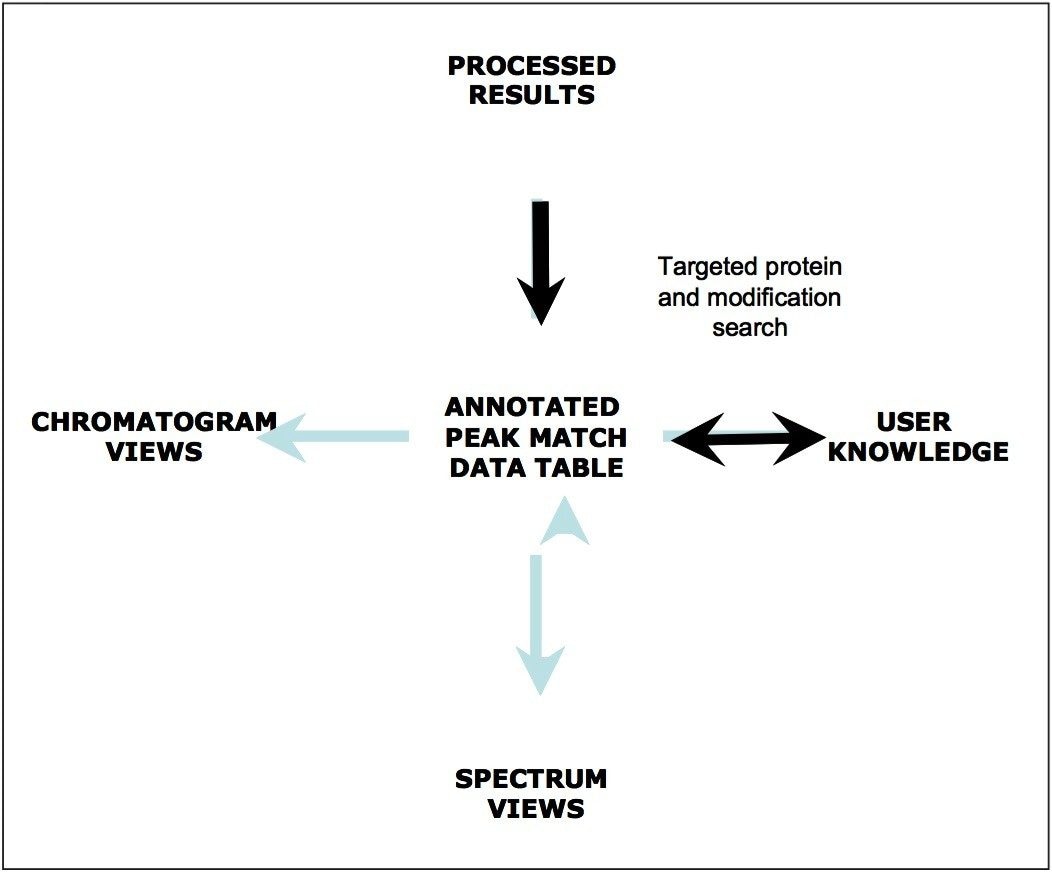
This peak-match data table represents a central location for processed data that supplies annotation information to the various tools that facilitate graphical result viewing and sample-sample comparisons. As depicted in Figure 9, a central BiopharmaLynx paradigm is that scientists are the ultimate arbiters for interpreting results generated by the program. This may include aligning and normalizing data, altering minimum intensity thresholds for peak detection, adjusting incorrect assignments produced by a search, choosing between alternative assignments fitting the protein search criteria, or using their knowledge to manually annotate components within a sample. Chromatographic and spectral displays are immediately updated with any changes to annotations, and user-comments associated with individual components can be included when constructing reporting templates.
This application note has detailed how the BiopharmaLynx Application Manager can demonstrate batch-to-batch differences in the glycovariant profile of a recombinant therapeutic monoclonal antibody. Important points in this note cover many of the best practices we employ to build processing methods capable of recognizing and comparing LC-TOF MS profiles of therapeutic antibody variants, and a rational framework for assessing that deconvolution conditions have been properly selected. Today, LC-MS data on intact antibodies can be routinely collected using fast and robust analytical methodologies. The productivity-limiting step for many analytical groups has become the processing and synthesis of useful information from the abundance of raw data generated by these studies. The BiopharmaLynx Application Manager has been designed to address this productivity limitation by automating processing and bioannotation of such data, while providing tools for sample comparison, data export, and report generation. Overall, these improved workflow efficiencies facilitate the rapid communication of results to other scientists and organizations, and help these groups make better decisions faster about the development of their biotherapeutics.
720002820, November 2008